Abstract
Key message
In this paper, we determine the genetic architecture controlling leaf flecking in maize and investigate its relationship to disease resistance and the defense response.
Abstract
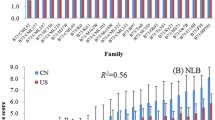
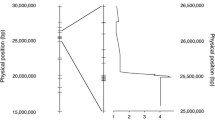
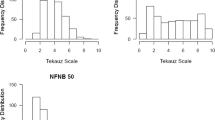
Flecking is defined as a mild, often environmentally dependent lesion phenotype observed on the leaves of several commonly used maize inbred lines. Anecdotal evidence suggests a link between flecking and enhanced broad-spectrum disease resistance. Neither the genetic basis underlying flecking nor its possible relationship to disease resistance has been systematically evaluated. The commonly used maize inbred Mo17 has a mild flecking phenotype. The IBM-advanced intercross mapping population, derived from a cross between Mo17 and another commonly used inbred B73, has been used for mapping a number of traits in maize including several related to disease resistance. In this study, flecking was assessed in the IBM population over 6 environments. Several quantitative trait loci for flecking were identified, with the strongest one located on chromosome 6. Low but moderately significant correlations were observed between stronger flecking and higher disease resistance with respect to two diseases, southern leaf blight and northern leaf blight and between stronger flecking and a stronger defense response.



Similar content being viewed by others
References
Balint-Kurti PJ, Carson ML (2006) Analysis of quantitative trait loci for resistance to southern leaf blight in juvenile maize. Phytopathology 96(3):221–225
Balint-Kurti PJ, Zwonitzer JC, Wisser RJ, Carson ML, Oropeza-Rosas MA, Holland JB, Szalma SJ (2007) Precise mapping of quantitative trait loci for resistance to southern leaf blight, caused by Cochliobolus heterostrophus race O, and flowering time using advanced intercross maize lines. Genetics 176:645–657
Balint-Kurti PJ, Wisser R, Zwonitzer JC (2008) Use of an advanced intercross line population for precise mapping of quantitative trait loci for gray leaf spot resistance in maize. Crop Sci 48:1696–1704
Balint-Kurti PJ, Yang J, Van Esbroeck G, Jung J, Smith ME (2010) Use of a maize advanced intercross line for mapping of qtl for northern leaf blight resistance and multiple disease resistance. Crop Sci 50:458–466
Brodersen P, Petersen M, Pike HM, Olszak B, Skov SR, Ødum N, Jørgensen LB, Brown RE, Mundy J (2002) Knockout of Arabidopsis ACCELERATED-CELL-DEATH11 encoding a sphingosine transfer protein causes activation of programmed cell death and defense. Genes Dev 16:490–502
Büschges R, Hollricher K, Panstruga R, Simons G, Wolter M, Frijters A, van Daelen R, van der Lee T, Diergaarde P, Groenendijk J, Töpsch S, Vos P, Salamini F, Schulze-Lefert P (1997) The Barley Mlo Gene: a novel control element of plant pathogen resistance. Cell 88:695–705
Chintamanani S, Hulbert SH, Johal GS, Balint-Kurti PJ (2010) Identification of a maize locus that modulates the hypersensitive defense response, using mutant-assisted gene identification and characterization. Genetics 184:813–825
Coe E, Cone K, McMullen M, Chen S-S, Davis G, Gardiner J, Liscum E, Polacco M, Paterson A, Sanchez-Villeda H, Soderlund C, Wing R (2002) Access to the maize genome: an integrated physical and genetic map. Plant Physiol 128:9–12
Davis GL, McMullen MD, Baysdorfer C, Musket T, Grant D, Staebell M, Xu G, Polacco M, Koster M-HL, Houchins K, Chao S, Coe JEH (1999) A maize map standard with sequenced core markers, grass genome reference points and 932 expressed sequence tagged sites(ESTs) in a 1736-locus map. Genetics 152:1137–1172
Falque M (2005) IRILmap: linkage map distance correction for intermated recombinant inbred lines/advanced recombinant inbred strains. Bioinformatics 21:3441–3442
Flint-Garcia SA, Thuillet AC, Yu JM, Pressoir G, Romero SM, Mitchell SE, Doebley J, Kresovich S, Goodman MM, Buckler ES (2005) Maize association population: a high-resolution platform for quantitative trait locus dissection. Plant J 44:1054–1064
Frank T, Hallauer A (1999) Inter- and intrapopulation genetic variances after ten cycles of reciprocal full-sib recurrent selection in the BS10 and BS11 synthetic maize populations. Maydica 44:9–24
Gray J, Close PS, Briggs SP, Johal GS (1997) A novel suppressor of cell death in plants encoded by the Lls1 gene of maize. Cell 89:25–31
Hoisington DA, Neuffer MG, Walbot V (1982) Disease lesion mimics in maize: I. Effect of genetic background, temperature, developmental age, and wounding on necrotic spot formation with Les1. Dev Biol 93:381–388
Holland JB, Nyquist WE, Cervantes-Martinez CT (2003) Estimating and interpreting heritability for plant breeding: an update. Plant Breed Rev 22:9–112
Hu G, Richter TE, Hulbert SH, Pryor T (1996) Disease lesion mimicry caused by mutations in the rust resistance Gene rp1. Plant Cell 8:1367–1376
Jambunathan N, Siani JM, McNellis TW (2001) A humidity-sensitive Arabidopsis copine mutant exhibits precocious cell death and increased disease resistance. Plant Cell 13:2225–2240
Johal GS (2007) Disease lesion mimic mutants of maize. American Phytopahtological Society Feature Story. http://www.apsnet.org/online/feature/mimics/
Lander ES, Botstein D (1989) Mapping mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121:185–199
Lee M, Sharopova N, Beavis WD, Grant D, Katt M, Blair D, Hallauer A (2002) Expanding the genetic map of maize with the intermated B73 X Mo17 (IBM) population. Plant Mol Biol 48:453–461
Li H, Ribaut J-M, Li Z, Wang J (2008) Inclusive composite interval mapping (ICIM) for digenic epistasis of quantitative traits in biparental populations. Theor Appl Genet 116:243–260
Lorrain S, Vailleau F, Balagué C, Roby D (2003) Lesion mimic mutants: keys for deciphering cell death and defense pathways in plants? Trends Plant Sci 8:263–271
Makepeace JC, Oxley SJP, Havis ND, Hackett R, Burke JI, Brown JKM (2007) Associations between fungal and abiotic leaf spotting and the presence of mlo alleles in barley. Plant Pathol 56:934–942
McMullen MD, Kresovich S, Villeda HS, Bradbury P, Li HH, Sun Q, Flint-Garcia S, Thornsberry J, Acharya C, Bottoms C, Brown P, Browne C, Eller M, Guill K, Harjes C, Kroon D, Lepak N, Mitchell SE, Peterson B, Pressoir G, Romero S, Rosas MO, Salvo S, Yates H, Hanson M, Jones E, Smith S, Glaubitz JC, Goodman M, Ware D, Holland JB, Buckler ES (2009) Genetic properties of the maize nested association mapping population. Science 325:737–740
Moeder W, Yoshioka K (2008) Lesion mimic mutants. Plant Signal Behav 3:764–767
Mur LA, Kenton P, Lloyd AJ, Ougham H, Prats E (2007) The hypersensitive response; the centenary is upon us but how much do we know? J Exp Bot 59:501–520
Negeri A, Wang G-F, Benavente L, Kibiti C, Chaikam V, Johal G, Balint-Kurti P (2013) Characterization of temperature and light effects on the defense response phenotypes associated with the maize Rp1-D21 autoactive resistance gene. BMC Plant Biol 13:106
Neuffer MG, Calvert OH (1975) Dominant disease lesion mimics in maize. J Hered 66:265–270
Piffanelli P, Ramsay L, Waugh R, Benabdelmouna A, D’Hont A, Hollricher K, Jorgensen JH, Schulze-Lefert P, Panstruga R (2004) A barley cultivation-associated polymorphism conveys resistance to powdery mildew. Nature 430:887–891
Todesco M, Balasubramanian S, Hu TT, Traw MB, Horton M, Epple P, Kuhns C, Sureshkumar S, Schwartz C, Lanz C, Laitinen RAE, Huang Y, Chory J, Lipka V, Borevitz JO, Dangl JL, Bergelson J, Nordborg M, Weigel D (2010) Natural allelic variation underlying a major fitness trade-off in Arabidopsis thaliana. Nature 465:632–636
Winkler CR, Jensen NM, Cooper M, Podlich DW, Smith OS (2003) On the determination of recombination rates in intermated recombinant inbred populations. Genetics 164:741–745
Wright SAI, Azarang M, Falk AB (2013) Barley lesion mimics, supersusceptible or highly resistant to leaf rust and net blotch. Plant Pathol 62:982–992
Wu Y-X, von Tiedemann A (2004) Light-dependent oxidative stress determines physiological leaf spot formation in barley. Phytopathology 94:584–592
Yin Z, Chen J, Zeng L, Goh M, Leung H, Khush GS, Wang G-L (2000) Characterizing rice lesion mimic mutants and identifying a mutant with broad-spectrum resistance to rice blast and bacterial blight. Mol Plant-Microbe Interact 13:869–876
Zehr BE, Dudley JW, Rufener GK (1994) QTLs for degree of pollen-silk discordance, expression of disease lesion mimic, and leaf curl response to drought. Maize Genet Coop Newslett 68:109–200
Zhao Y, Lu X, Liu C, Guan H, Zhang M, Li Z, Cai H, Lai J (2012) Identification and fine mapping of rhm1 locus for resistance to southern corn leaf blight in maize. J Int Plant Biol 54:321–329
Acknowledgments
We thank Major Goodman and the Maize Genetics Cooperation Stock Center for provision of seed and David Rhyne, Ed Durren, Greg Marshall, Shannon Sermons, and Donna Stephens for their help with fieldwork. We thank Steve Moose for allowing us to observe and take lesion mimic data from his field trials and Steve Szalma for helpful discussions. We thank Cathy Herring and the staff of Central Crops in Clayton NC for maintaining the field trials in NC. This work was funded by the U.S. Department of Agriculture–Agricultural Research Service, NC State University, Purdue University, and by National Science Foundation Grant 0822495. This paper is dedicated to the memory of Steve Szalma.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by J. Yan.
Vijay Vontimitta and Bode A Olukolu contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Vontimitta, V., Olukolu, B.A., Penning, B.W. et al. The genetic basis of flecking and its relationship to disease resistance in the IBM maize mapping population. Theor Appl Genet 128, 2331–2339 (2015). https://doi.org/10.1007/s00122-015-2588-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-015-2588-8