Abstract
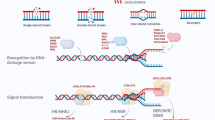
Eukaryotic cells repair thousands of lesions arising in the genome at each cell cycle. The most hazardous damage is likely DNA double-strand breaks (DSB) that cleave the double helix backbone. DSBs occur naturally during T cell receptor and immunoglobulin gene recombination in lymphocytes. DSBs can also arise as a consequence of exogenous stresses (e.g., ionizing irradiation, chemotherapeutic drugs, viruses) or oxidative processes. An increasing number of studies have reported that infection with pathogenic bacteria also alters the host genome, producing DSB and other modifications on DNA. This review focuses on recent data on bacteria-induced DNA damage and the known strategies used by these pathogens to maintain a physiological niche in the host. Even after DNA repair in infected cells, “scars” often remain on chromosomes and might generate genomic instability at the next cell division. Chronic inflammation in tissue, combined with infection and DNA damage, can give rise to genomic instability and eventually cancer. A functional link between the DNA damage response and the innate immune response has been recently established. Pathogenic bacteria also highjack the host cell cycle, often acting on the stability of the master regulator p53, or dampen the DNA damage response to support bacterial replication in an appropriate reservoir. Except in a few cases, the molecular mechanisms responsible for DNA lesions are poorly understood, although ROS release during infection is a serious candidate for generating DNA breaks. Thus, chronic or repetitive infections with genotoxic bacteria represent a common source of DNA lesions that compromise host genome integrity.



Similar content being viewed by others
Abbreviations
- ATM:
-
Ataxia telangiectasia mutated
- ATR:
-
Ataxia telangiectasia and Rad3-related
- 53BP1:
-
p53 binding protein 1
- Chk1/2:
-
Checkpoint kinase 1/2
- DNA-PK:
-
DNA protein kinase
- dsDNA:
-
Double-stranded DNA
- CDT:
-
Cytolethal distending toxin
- DDR:
-
DNA damage response
- DSB/SSB:
-
Double/Single strand breaks
- HR:
-
Homologous recombination
- IRIF:
-
Ionizing radiation induced foci
- LLO:
-
Lysteriolysin O
- MCD1:
-
Mediator of DNA-damage checkpoint 1
- M/HDM2:
-
Mouse/Human double minute 2 homolog
- MNR:
-
Mre11, Nbs1 and Rad50
- MOI:
-
Multiplicity of infection
- Mre11:
-
Mediator of DNA damage checkpoint
- NHEJ:
-
Non-Homologous End Joining
- OGG1:
-
8-oxoguanine DNA glycosylase
- PARP:
-
Poly(ADP-ribose) polymerase
- PRR:
-
Pattern recognition receptor
- PI3KK:
-
Phosphatidylinositol 3-kinase-like kinase
- ROS:
-
Reactive oxygen and nitrogen species
- STING:
-
Stimulator of interferon genes
- TTSS:
-
Type III secretion system
References
Ciccia A, Elledge SJ (2010) The DNA damage response: making it safe to play with knives. Mol Cell 40:179–204
Touati E (2010) When bacteria become mutagenic and carcinogenic: lessons from H. pylori. Mutat Res 703:66–70
Nougayrède JP, Homburg S, Taieb F, Boury M, Brzuszkiewicz E, Gottschalk G, Buchrieser C, Hacker J, Dobrindt U, Oswald E (2006) Escherichia coli induces DNA double-strand breaks in eukaryotic cells. Science 313:848–851
Price BD, D’Andrea AD (2013) Cell Chromatin remodeling at DNA double-strand breaks. Cell 14:1344–1354
Bernstein C, Prasad AR, Nfonsam V, Bernstein H (2013) DNA damage, DNA repair and cancer. In: Clark C (ed) New research directions in DNA repair, chap 16. InTech, ISBN: 978-953-51-1114-6, doi:10.5772/53919
Britton S, Coates J, Jackson SP (2013) A new method for high-resolution imaging of Ku foci to decipher mechanisms of DNA double-strand break repair. J Cell Biol 202:579–595
Bewersdorf J, Bennett BT, Knight KL (2006) H2AX chromatin structures and their response to DNA damage revealed by 4Pi microscopy. Proc Natl Acad Sci USA 103:18137–18142
Rogakou EP, Boon C, Redon C, Bonner WM (1999) Megabase chromatin domains involved in DNA double-strand breaks in vivo. J Cell Biol 146:905–916
Iacovoni JS, Caron P, Lassadi I, Nicolas E, Massip L, Trouche D, Legube G (2010) High-resolution profiling of gammaH2AX around DNA double strand breaks in the mammalian genome. EMBO 29:1446–1457
Cover TL, Blaser MJ (2009) Helicobacter pylori in health and disease. Gastroenterology 136:1863–1873
Kim JJ, Tao H, Carloni E, Leung WK, Graham DY, Sepulveda AR (2002) Helicobacter pylori impairs DNA mismatch repair in gastric epithelial cells. Gastroenterology 123:542–553
Machado AM, Figueiredo C, Touati E, Máximo V, Sousa S, Michel V, Carneiro F, Nielsen FC, Seruca R, Rasmussen LJ (2009) Helicobacter pylori infection induces genetic instability of nuclear and mitochondrial DNA in gastric cells. Clin Cancer Res 15:2995–3002
Machado AM, Figueiredo C, Seruca R, Rasmussen LJ (2010) Helicobacter pylori infection generates genetic instability in gastric cells. Biochem Biophys Acta 1806:58–65
Touati E, Michel V, Thiberge JM, Avé P, Huerre M, Bourgade F, Klungland A, Labigne A (2003) Chronic Helicobacter pylori infections induce gastric mutations in mice. Gastroenterology 124:1408–1419
Umeda M, Murata-Kamiya N, Saito Y, Ohba Y, Takahashi M, Hatakeyama M (2009) Helicobacter pylori CagA causes mitotic impairment and induces chromosomal instability. J Biol Chem 284:22166–22172
Wei J, Nagy TA, Vilgelm A, Ogden SR, Romero-Gallo J, Piazuelo MB, Correa P, Washington MK, El-Rifai W, Peek RM, Zaika A (2010) Regulation of p53 tumor suppressor by Helicobacter pylori in gastric epithelial cells. Gastroenterology 139:1333–1343
Buti L, Spooner E, Van der Veen AG, Rappuoli R, Covacci A, Ploegh HL (2011) Helicobacter pylori cytotoxin-associated gene A (CagA) subverts the apoptosis-stimulating protein of p53 (ASPP2) tumor suppressor pathway of the host. Proc Natl Acad Sci USA 108:9238–9243
Toller IM, Neelsen KJ, Steger M, Hartung ML, Hottiger MO, Stucki M, Kalali B, Gerhard M, Sartori AA, Lopes M, Müller A (2011) Carcinogenic bacterial pathogen Helicobacter pylori triggers DNA double-strand breaks and a DNA damage response in its host cells. Proc Natl Acad Sci USA 108:14944–14949
Tenaillon O, Skurnik D, Picard B, Denamur E (2010) The population genetics of commensal Escherichia coli. Nat Rev Microbiol 8:207–217
Cuevas-Ramos G, Petit CR, Marcq I, Marcq I, Boury M, Oswald E, Nougayrède JP (2010) Escherichia coli induces DNA damage in vivo and triggers genomic instability in mammalian cells. Proc Natl Acad Sci USA 107:11537–11542
Secher T, Samba-Louaka A, Oswald E, Nougayrède JP (2013) Escherichia coli producing colibactin triggers premature and transmissible senescence in mammalian cells. PLoS One 8:e77157
Burton DG, Krizhanovsky V (2014) Physiological and pathological consequences of cellular senescence. Cell Mol Life Sci 71:4373–4386
Cougnoux A, Dalmasso G, Martinez R, Buc E, Delmas J, Gibold L, Sauvanet P, Darcha C, Déchelotte P, Bonnet M, Pezet D, Wodrich H, Darfeuille-Michaud A, Bonnet R (2014) Bacterial genotoxin colibactin promotes colon tumour growth by inducing a senescence-associated secretory phenotype. Gut 63:1932–1942
Putze J, Hennequin C, Nougayrède J-P, Zhang W, Homburg S, Karch H, Bringer MA, Fayolle C, Carniel E, Rabsch W, Oelschlaeger TA, Oswald E, Forestier C, Hacker J, Dobrindt U (2009) Genetic structure of the colibactin genomic island among members of the family Enterobacteriaceae. Infect Immun 77:4696–4703
Lai Y-C, Lin A-C, Chiang MK, Dai YH, Hsu CC, Lu MC, Liau CY, Chen YT (2014) Genotoxic Klebsiella pneumoniae in Taiwan. PLoS One 9:e96292
Schwabe RF, Jobib C (2013) The microbiome and cancer. Nat Rev Cancer 13:800–812
Arthur JC, Perez-Chanona E, Mühlbauer M, Tomkovich S, Uronis JM, Fan TJ, Campbell BJ, Abujamel T, Dogan B, Rogers AB, Rhodes JM, Stintzi A, Simpson KW, Hansen JJ, Keku TO, Fodor AA, Jobin C (2012) Intestinal inflammation targets cancer-inducing activity of the microbiota. Science 338:120–123
Raisch J, Buc E, Bonnet M, Sauvanet P, Vazeille E, de Vallée A, Déchelotte P, Darcha C, Pezet D, Bonnet R, Bringer MA, Darfeuille-Michaud A (2014) Colon cancer-associated B2 Escherichia coli colonize gut mucosa and promote cell proliferation. World J Gastroenterol 20:6560–6572
Arthur JC, Gharaibeh RZ, Mühlbauer M, Perez-Chanona E, Uronis JM, McCafferty J, Fodor AA, Jobin C (2014) Microbial genomic analysis reveals the essential role of inflammation in bacteria-induced colorectal cancer. Nat Commun 5:4724. doi:10.1038/ncomms5724
Olier M, Marcq I, Salvador-Cartier C, Secher T, Dobrindt U, Boury M, Bacquié V, Pénary M, Gaultier E, Nougayrède JP, Fioramonti J, Oswald E (2013) Genotoxicity of Escherichia coli Nissle 1917 strain cannot be dissociated from its probiotic activity. Gut Microb 3:501–509
Bian X, Fu J, Plaza A, Herrmann J, Pistorius D, Stewart AF, Zhang Y, Müller R (2013) In vivo evidence for a prodrug activation mechanism during colibactin maturation. ChemBioChem 14:1194–1197
Brotherton CA, Wilson M, Balskus EP (2015) Isolation of a metabolite from pks island provides insights into colibactin biosynthesis and activity. Org Lett. doi:10.1021/acs.orglett.5b00432
Yan S, Sorrell M, Berman Z (2014) Functional interplay between ATM/ATR-mediated DNA damage response and DNA repair pathways in oxidative stress. Cell Mol Life Sci 71:3951–3967
Lara-Tejero M, Galán JE (2000) A bacterial toxin that controls cell cycle progression as a deoxyribonuclease I-like protein. Science 290:354–357
Comayras C, Tasca C, Pérès SY, Ducommun B, Oswald E, De Rycke J (1997) Escherichia coli cytolethal distending toxin blocks the HeLa cell cycle at the G2/M transition by preventing cdc2 protein kinase dephosphorylation and activation. Infect Immun 65:5088–5095
Guerra L, Cortes-Bratti X, Guidi R, Frisan T (2011) The biology of the cytolethal distending toxins. Toxins (Basel) 3:172–190
Li L, Sharipo A, Chaves-Olarte E, Masucci MG, Levitsky V, Thelestam M, Frisan T (2002) The Haemophilus ducreyi cytolethal distending toxin activates sensors of DNA damage and repair complexes in proliferating and non-proliferating cells. Cell Microbiol 4:87–99
Frisan T, Cortes-Bratti X, Chaves-Olarte E, Stenerlöw B, Thelestam M (2003) The Haemophilus ducreyi cytolethal distending toxin induces DNA double-strand breaks and promotes ATM-dependent activation of RhoA. Cell Microbiol 5:695–707
Blazkova H, Krejcikova K, Moudry P et al (2010) Bacterial intoxication evokes cellular senescence with persistent DNA damage and cytokine signalling. J Cell Mol Med 14:357–367
Fahrer J, Huelsenbeck J, Jaurich H, Dörsam B, Frisan T, Eich M, Roos WP, Kaina B, Fritz G (2014) Cytolethal distending toxin (CDT) is a radiomimetic agent and induces persistent lebels of DNA double-strand breaks in human fibroblasts. DNA Repair 18:31–43
Cortes-Bratti X, Karlsson C, Lagergård T, Thelestam M, Frisan T (2001) The Haemophilus ducreyi cytolethal distending toxin induces cell cycle arrest and apoptosis via the DNA damage checkpoint pathways. J Biol Chem 276:5296–5302
Kitagawa T, Hoshida H, Akada R (2007) Genome-wide analysis of cellular response to bacterial genotoxin CdtB in yeast. Infect Immun 75:1393–1402
Fedor Y, Vignard J, Nicolau-Travers ML, Boutet-Robinet E, Watrin C, Salles B, Mirey G (2013) From single-strand breaks to double-strand breaks during S-phase: a new mode of action of the Escherichia coli cytolethal distending toxin. Cell Microbiol 15:1–15
Bezine E, Vignard J, Mirey G (2014) The cytolethal distending toxin effects on mammalian cells: a DNA damage perspective. Cells 3:592–615
Guidi R, Guerra L, Levi L, Stenerlöw B, Fox JG, Josenhans C, Masucci M, Frisan T (2013) Chronic exposure to the cytolethal distending toxins of Gram-negative bacteria promotes genomic instability and altered DNA damage response. Cell Microbiol 15:98–113
Kerr KG, Snelling AM (2009) Pseudomonas aeruginosa: a formidable and ever-present adversary. J Hosp Infect 73:338–344
Kunz AN, Brook I (2010) Emerging resistant Gram-negative aerobic bacilli in hospital-acquired infections. Chemotherapy 56:492–500
Furukawa S, Kuchma SL, O’Toole GA (2014) Keeping their options open: acute versus persistent infections. J Bacteriol 188:1211–1217
Roy-Burman A, Savel RH, Racine S, Swanson BL, Revadigar NS, Fujimoto J, Sawa T, Frank DW, Wiener-Kronish JP (2001) Type III protein secretion is associated with death in lower respiratory and systemic Pseudomonas aeruginosa infections. J Infect Dis 183:1767–1774
Hauser AR, Cobb E, Bodi M, Mariscal D, Vallés J, Engel JN, Rello J (2002) Type III protein secretion is associated with poor clinical outcomes in patients with ventilator-associated pneumonia caused by Pseudomonas aeruginosa. Crit Care Med 30:521–528
Hauser AR (2009) The type III secretion system of Pseudomonas aeruginosa: infection by injection. Nat Rev Microbiol 7:654–665
Elsen S, Collin-Faure V, Gidrol X, Lemercier C (2013) The opportunistic pathogen Pseudomonas aeruginosa activates the DNA double-strand break signaling and repair pathway in infected cells. Cell Mol Life Sci 70:4385–4397
Wu M, Huang H, Zhang W, Kannan S, Weaver A, McKibben M, Herington D, Zeng H, Gao H (2011) Host DNA repair proteins in response to Pseudomonas aeruginosa in lung epithelial cells and in mice. Infect Immun 79:75–87
David SS, O’Shea VL, Kundu S (2007) Base-excision repar of oxidative DNA damage. Nature 447:941–950
Song J, Bent AF (2014) Microbial pathogens trigger host DNA double-strand breaks whose abundance is reduced by plant defense responses. PLoS Pathog 10:e1004030
Fu ZQ, Guo M, Jeong BR, Tian F, Elthon TE, Cerny RL, Staiger D, Alfano JR (2007) A type III effector ADP-ribosylates RNA-binding proteins and quells plant immunity. Nature 447:284–288
Markou P, Apidianakis Y (2014) Pathogenesis of intestinal Pseudomonas aeruginosa infection in patients with cancer. Front Cell Infect Microbiol 3:1–5
The Human Microbiome Projet Consortium (2012) Structure, function and diversity of the healthy human microbiome. Nature 486:207–214
Bertrand X, Thouverez M, Talon D, Boillot A, Capellier G, Floriot C, Hélias JP (2001) Endemicity, molecular diversity and colonisation routes of Pseudomonas aeruginosa in intensive care units. Intensive Care Med 27:1263–1268
Bergounioux J, Elisee R, Prunier AL, Donnadieu F, Sperandio B, Sansonetti P, Arbibe L (2012) Calpain activation by the Shigella flexneri effector VirA regulates key steps in the formation and life of the bacterium’s epithelial niche. Cell Host Microbe 11:240–252
Rudel T (2012) To die or not to die: Shigella has an answer. Cell Host Microbe 11:219–221
Word Health Organization, Department of Reproductive Health and Research (2012) Emergence of multi-drug resistant Neisseria gonorrhoeae. Threat of global rise in untreatable sexually transmitted infections. WHO reference number: WHO/RHR/11.14. Fact sheet11.14
Vielfort K, Söderholm N, Weyler L, Vare D, Löfmark S, Aro H (2013) Neisseria gonorrhoeae infection causes DNA damage and affects the expression of p21, p27 and p53 in non-tumor epithelial cells. J Cell Sci 126:339–347
Leitão E, Costa AC, Brito C, Costa L, Pombinho R, Cabanes D, Sousa S (2014) Listeria monocytogenes induces host DNA damage and delays the host cell cycle to promote infection. Cell Cycle 13:928–940
Samba-Louaka A, Pereira JM, Nahori MA, Villiers V, Deriano L, Hamon MA, Cossart P (2014) Listeria monocytogenes dampens the DNA damage response. PLoS Pathog 10(10):e1004470. doi:10.1371/journal.ppat.1004470
Lam GY, Fattouh R, Muise AM, Grinstein S, Higgins DE, Brumell JH (2011) Listeriolysin O suppresses phospholipase C-mediated activation of the microbicidal NADPH oxydase to promote Listeria monocytogenes infection. Cell Host Microbe 10:627–634
Eskandarian HA, Impens F, Nahori MA, Soubigou G, Coppée JY, Cossart P, Hamon MA (2013) A role for SIRT2-dependent histone H3K18 deacetylation in bacterial infection. Science 341:525. doi:10.1126/science.1238858
Bastidas RJ, Elwell CA, Engel JN, Valdivia RH (2013) Chlamydial intracellular survival strategies. Cold Spring Harb Perspect Med 3:a010256
Chumduri C, Gurumurthy RK, Zadora PK, Mi Y, Meyer TF (2013) Chlamydia infection promotes host DNA damage and proliferation but impairs the DNA damage response. Cell Host Microbe 13:746–758
Siegl C, Prusty BK, Karunakaran K, Wischhusen J, Rudel T (2014) Tumor suppressor p53 alters host cell metabolism to limit Chlamydia trachomatis infection. Cell Reports 9:918–929
González E, Rother M, Kerr MC, Al-Zeer MA, Abu-Lubad M, Kessler M, Brinkmann V, Loewer A, Meyer TF (2014) Chlamydia infection depends on a functional MDM2-p53 axis. Nat Commun. 5:5201. doi:10.1038/ncomms6201
Padberg I, Janßen S, Meyer TF (2013) Chlamydia trachomatis inhibits telomeric DNA damage signaling via transient hTERT upregulation. Int J Med Microbiol 303:463–474
Abdullah Z, Knolle PA (2014) Scaling of immune responses against intracellular bacterial infection. EMBO J 33:2283–2294
Paldudan SR, Bowie AG (2013) Immune sensing of DNA. Immunity 38:870–880
Chatzinikolaou G, Karakasilioti I, Garinis GA (2014) DNA damage and innate immunity: links and trade-offs. Trends Immunol 35:429–435
Jakobsen MR, Paludan SR (2015) IFI16: at the interphase between innate DNA sensing and genome regulation. Cytokine Growth Factor Rev 25:649–655
Kondo T, Kobayashi J, Saitoh T, Maruyama K, Ishii KJ, Barber GN, Komatsu K, Akira S, Kawai T (2013) DNA damage sensor MRE11 recognizes cytosolic double-stranded DNA and induces type I interferon by regulating STING trafficking. Proc Natl Acad Sci USA 110:2969–2974
Roth S, Rottach A, Lotz-Havla AS, Laux V, Muschaweckh A, Gersting SW, Muntau AC, Hopfner KP, Jin L, Vanness K, Petrini JH, Drexler I, Leonhardt H, Ruland J (2014) Rad50-CARD9 interactions link cytosolic DNA sensing to IL-1β production. Nat Immunol 15:538–545
Ferguson BJ, Mansur DS, Peters NE, Ren H, Smith GL (2012) DNA-PK is a DNA sensor for IRF-3-dependent innate immunity. Elife 1:e00047. doi:10.755/eLife.00047
Härtlova A, Erttmann SF, Raffi FA, Schmalz AM, Resch U, Anugula S, Lienenklaus S, Nilsson LM, Kröger A, Nilsson JA, Ek T, Weiss S, Gekara NO (2015) DNA damage primes the type I Interferon system via the cytosolic DNA sensor STING to promote anti-microbial innate immunity. Immunity 42:332–343
Acknowledgments
CL thanks Dr. Joël Gaffé (Adaptation and Pathogeny of Micro-organisms Laboratory, University Grenoble Alpes, CNRS) for critical reading and comments on the manuscript, Dr. Sandeep Nadendla for his careful reading and help with English language.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lemercier, C. When our genome is targeted by pathogenic bacteria. Cell. Mol. Life Sci. 72, 2665–2676 (2015). https://doi.org/10.1007/s00018-015-1900-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-015-1900-8