Abstract
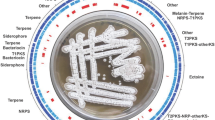
A draft genome sequence of Streptomyces ansochromogenes 7100 was generated using 454 sequencing technology. In combination with local BLAST searches and gap filling techniques, a comprehensive antiSMASH-based method was adopted to assemble the secondary metabolite biosynthetic gene clusters in the draft genome of S. ansochromogenes. A total of at least 35 putative gene clusters were identified and assembled. Transcriptional analysis showed that 20 of the 35 gene clusters were expressed in either or all of the three different media tested, whereas the other 15 gene clusters were silent in all three different media. This study provides a comprehensive method to identify and assemble secondary metabolite biosynthetic gene clusters in draft genomes of Streptomyces, and will significantly promote functional studies of these secondary metabolite biosynthetic gene clusters.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bentley S, Chater K, Cerdeno-Tarraga A M, et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3 (2). Nature, 2002, 417: 141–147
Ōmura S, Ikeda H, Ishikawa J, et al. Genome sequence of an industrial microorganism Streptomyces avermitilis: deducing the ability of producing secondary metabolites. Proc Natl Acad Sci USA, 2001, 98: 12215–12220
Nett M, Ikeda H, Moore B S. Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat Prod Rep, 2009, 26: 1362–1384
Song L, Barona-Gomez F, Corre C, et al. Type III polyketide synthase β-ketoacyl-ACP starter unit and ethylmalonyl-CoA extender unit selectivity discovered by Streptomyces coelicolor genome mining. J Am Chem Soc, 2006, 128: 14754–14755
Bok J W, Hoffmeister D, Maggio-Hall L A, et al. Genomic mining for Aspergillus natural products. Chem Bio, 2006, 13: 31–37
Gross H, Stockwell V O, Henkels M D, et al. The genomisotopic approach: a systematic method to isolate products of orphan biosynthetic gene clusters. Chem Bio, 2007, 14: 53–63
Starcevic A, Zucko J, Simunkovic J, et al. ClustScan: an integrated program package for the semi-automatic annotation of modular biosynthetic gene clusters and in silico prediction of novel chemical structures. Nucleic Acids Res, 2008, 36: 6882–6892
Weber T, Rausch C, Lopez P, et al. CLUSEAN: a computer-based frame-work for the automated analysis of bacterial secondary metabolite biosynthetic gene clusters. J Biotechnol, 2009, 140: 13–17
Li M, Ung P, Zajkowski J, et al. Automated genome mining for natural products. BMC Bioinformatics, 2009, 10: 185
Anand S, Prasad M, Yadav G, et al. SBSPKS: structure based sequence analysis of polyketide synthases. Nucleic Acids Res, 2010, 38: W487–W496
Khaldi N, Seifuddin F T, Turner G, et al. SMURF: genomic mapping of fungal secondary metabolite clusters. Fungal Genet Biol, 2010, 47: 736–741
Medema M H, Blin K, Cimermancic P, et al. AntiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res, 2011, 39: W339–W346
Pagani I, Liolios K, Jansson J, et al. The genomes online database (gold) v. 4: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Res, 2012, 40: D571–D579
Simpson J T, Wong K, Jackman S D, et al. ABySS: a parallel assembler for short read sequence data. Genome Res, 2009, 19: 1117–1123
Zerbino D R, Birney E. Velvet: algorithms for de novo short read assembly using de bruijn graphs. Genome Res, 2008, 18: 821–829
Chaisson M J, Pevzner P A. Short read fragment assembly of bacterial genomes. Genome Res, 2008, 18: 324–330
Li Y, Ling H, Li W, et al. Improvement of nikkomycin production by enhanced copy of sanU and sanV in Streptomyces ansochromogenes and characterization of a novel glutamate mutase encoded by sanU and sanV. Metab Eng, 2005, 7: 165–173
Kieser T, Bibb M J, Buttner M J, et al. Practical Streptomyces Genetics. Norwich: The John Innes Foundation, 2000
Aziz R, Bartels D, Best A, et al. The rast server: rapid annotations using subsystems technology. BMC Genomics, 2008, 9: 75
Delcher A L, Harmon D, Kasif S, et al. Improved microbial gene identification with glimmer. Nucleic Acids Res, 1999, 27: 4636–4641
Altschul S F, Gish W, Miller W, et al. Basic local alignment search tool. J Mol Biol, 1990, 215: 403–410
Ansari M Z, Yadav G, Gokhale R S, et al. NRPS-PKS: a knowledge-based resource for analysis of NRPS/PKS megasynthases. Nucleic Acids Res, 2004, 32: W405–W413
Eddy S R. Profile hidden markov models. Bioinformatics, 1998, 14: 755–763
Van Pée K H, Patallo E P. Flavin-dependent halogenases involved in secondary metabolism in bacteria. Appl Microbiol Biotechnol, 2006, 70: 631–641
Taylor B L, Zhulin I B. PAS domains: internal sensors of oxygen, redox potential, and light. Microbiol Mol Biol R, 1999, 63: 479–506
Walsh C T, Fischbach M A. Natural products version 2.0: connecting genes to molecules. J Am Chem Soc, 2010, 132: 2469–2493
Liu G, Chater K F, Chandra G, et al. Molecular regulation of antibiotic biosynthesis in Streptomyces. Microbiol Mol Biol R, 2013, 77: 112–143
Lai S, Zhang Y, Liu S, et al. Metabolic engineering and flux analysis of Corynebacterium glutamicum for L-serine production. Sci China Life Sci, 2012, 55: 283–290
Horinouchi S, Beppu T. A-factor as a microbial hormone that controls cellular differentiation and secondary metabolism in Streptomyces griseus. Mol Microbiol, 1994, 12: 859–864
Author information
Authors and Affiliations
Corresponding authors
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Zhong, X., Tian, Y., Niu, G. et al. Assembly and features of secondary metabolite biosynthetic gene clusters in Streptomyces ansochromogenes. Sci. China Life Sci. 56, 609–618 (2013). https://doi.org/10.1007/s11427-013-4506-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4506-0