Abstract
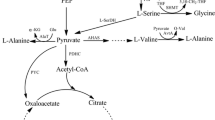
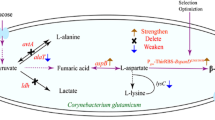
l-Serine plays a critical role as a building block for cell growth, and thus it is difficult to achieve the direct fermentation of l-serine from glucose. In this study, Corynebacterium glutamicum ATCC 13032 was engineered de novo by blocking and attenuating the conversion of l-serine to pyruvate and glycine, releasing the feedback inhibition by l-serine to 3-phosphoglycerate dehydrogenase (PGDH), in combination with the co-expression of 3-phosphoglycerate kinase (PGK) and feedback-resistant PGDH (PGDHr). The resulting strain, SER-8, exhibited a lower specific growth rate and significant differences in l-serine levels from Phase I to Phase V as determined for fed-batch fermentation. The intracellular l-serine pool reached (14.22±1.41) μmol gCDM−1, which was higher than glycine pool, contrary to fermentation with the wild-type strain. Furthermore, metabolic flux analysis demonstrated that the over-expression of PGK directed the flux of the pentose phosphate pathway (PPP) towards the glycolysis pathway (EMP), and the expression of PGDHr improved the l-serine biosynthesis pathway. In addition, the flux from l-serine to glycine dropped by 24%, indicating that the deletion of the activator GlyR resulted in down-regulation of serine hydroxymethyltransferase (SHMT) expression. Taken together, our findings imply that l-serine pool management is fundamental for sustaining the viability of C. glutamicum, and improvement of C1 units generation by introducing the glycine cleavage system (GCV) to degrade the excessive glycine is a promising target for l-serine production in C. glutamicum.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Furuya S. An essential role for de novo biosynthesis of l-serine in CNS development. Asia Pac Clin Nutr, 2008, 17: 312–315
Furuya S, Makino A, Hirabayashi Y. An improved method for culturing cerebellar Purkinje cells with differentiated dendrites under a mixed monolayer setting. Brain Res Protoc, 1998, 3: 192–198
Pizer L I, Potochny M L. Nutritional and regulatory aspects of serine metabolism in Escherichia coli. J Bacteriol, 1964, 88: 611–619
McNeil J B, Bognar A L, Pearlman R E. In vivo analysis of folate coenzymes and their compartmentation in Saccharomyces cerevisiae. Genetics, 1996, 142: 371–381
Gelling C L, Piper M D W, Hong S P, et al. Identification of a novel one-carbon metabolism regulon in Saccharomyces cerevisiae. J Biol Chem, 2004, 279: 7072–7081
Stauffer G V. Biosynthesis of serine, glycine, and one-carbon units. In: Neidhardt F C Curtiss III R, Ingraham J L, Lin E C C, et al., eds. Escherichia coli and Salmonella: Cellular and Molecular Biology. 2nd ed. Washington D.C.: ASM Press, 1996. 506–513
Marx A, deGraaf A A, Wiechert W, et al. Determination of the fluxes in the central metabolism of Corynebacterium glutamicum by nuclear magnetic resonance spectroscopy combined with metabolite balancing. Biotechnol Bioeng, 1996, 49: 111–129
Peters-Wendisch P, Netzer R, Eggeling L, et al. 3-Phosphoglycerate dehydrogenase from Corynebacterium glutamicum: the C-terminal domain is not essential for activity but is required for inhibition by l-serine. Appl Microbiol Biotechnol, 2002, 60: 437–441
Netzer R, Peters-Wendisch P, Eggeling L, et al. Cometabolism of a nongrowth substrate: l-serine utilization by Corynebacterium glutamicum. Appl Environ Microbiol, 2004, 70: 7148–7155
Haitani Y, Awano N, Yamazaki M, et al. Functional analysis of l-serine O-acetyltransferase from Corynebacterium glutamicum. FEMS Microbiol Lett, 2006, 255: 156–163
Ikeda M. Towards bacterial strains overproducing l-tryptophan and other aromatics by metabolic engineering. Appl Microbiol Biotechnol, 2006, 69: 615–626
Simic P, Willuhn J, Sahm H, et al. Identification of glyA (encoding serine hydroxymethyltransferase) and its use together with the exporter ThrE to increase l-threonine accumulation by Corynebacterium glutamicum. Appl Environ Microbiol, 2002, 68: 3321–3327
Schweitzer J E, Stolz M, Diesveld R, et al. The serine hydroxymethyltransferase gene glyA in Corynebacterium glutamicum is controlled by GlyR. J Biotechnol, 2009, 139: 214–221
Kubota K, Yokozeki K. Production of l-serine from glycine by Corynebacterium glycinophilum and properties of serine hydroxymethyltransferase, a key enzyme in l-serine production. J Ferment Bioeng, 1989, 67: 387–390
Izumi Y, Yoshida T, Miyazaki S S, et al.l-Serine production by a methylotroph and its related enzymes. Appl Microbiol Biotechnol, 1993, 39: 427–432
Stolz M, Peters-Wendisch P, Etterich H, et al. Reduced folate supply as a key to enhanced l-serine production by Corynebacterium glutamicum. Appl Environ Microbiol, 2007, 73: 750–755
Becker J, Zelder O, Häfner S, et al. From zero to hero—design-based systems metabolic engineering of Corynebacterium glutamicum for l-lysine production. Metab Eng, 2011, 13: 159–168
Holátko J, Elišáková V, Prouza M, et al. Metabolic engineering of the l-valine biosynthesis pathway in Corynebacterium glutamicum using promoter activity modulation. J Biotechnol, 2009, 139: 203–210
Park S D, Lee J Y, Sim S Y, et al. Characteristics of methionine production by an engineered Corynebacterium glutamicum strain. Metab Eng, 2007, 9: 327–336
Peters-Wendisch P, Stolz M, Etterich H, et al. Metabolic engineering of Corynebacterium glutamicum for l-serine production. Appl Environ Microbiol, 2005, 71: 7139–7144
Sambrook J, Fritsch E F, Maniatis T. Molecular Cloning: A Laboratory Manual. 2nd ed. New York: Cold Spring Harbor Laboratory Press, 1989
Schäfer A, Tauch A, Jäger W, et al. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Genetics, 1994, 145: 69–73
Jakoby M, Ngouoto-Nkili C E, Burkovski A. Construction and application of new Corynebacterium glutamicum vectors. Biotechnol Tech, 1999, 13: 437–441
Cremer J, Eggeling L, Sahm H. Control of the lysine biosynthesis sequence in Corynebacterium glutamicum as analyzed by overexpression of the individual corresponding genes. Appl Environ Microbiol, 1991, 57: 1746–1752
van der Rest M E, Lange C, Molenaar D. A heat shock following electroporation induces highly efficient transformation of Corynebacterium glutamicum with xenogeneic plasmid DNA. Appl Microbiol Biotechnol, 1999, 52: 541–545
Jäger W, Schäfer A, Pühler A, et al. Expression of the Bacillus subtilis sacB gene leads to sucrose sensitivity in the gram-positive bacterium Corynebacterium glutamicum but not in Streptomyces lividans. J Bacteriol, 1992, 174: 5462–5465
Zhang Y, Shang X, Deng A, et al. Genetic and biochemical characterization of Corynebacterium glutamicum ATP phosphoribosyltransferase and its three mutants resistant to feedback inhibition by histidine. Biochimie, 2012, 94: 829–838
Persicke M, Plassmeier J, Neuweger H, et al. Size exclusion chromatography—an improved method to harvest Corynebacterium glutamicum cells for the analysis of cytosolic metabolites. J Biotechnol, 2011, 155: 266–267
Taymaz-Nikerel H, de Mey M, Ras C, et al. Development and application of a differential method for reliable metabolome analysis in Escherichia coli. Anal Biochem, 2009, 386: 9–19
Klimacek M, Krahulec S, Sauer U, et al. Limitations in xylose-fermenting Saccharomyces cerevisiae, made evident through comprehensive metabolite profiling and thermodynamic analysis. Appl Environ Microbiol, 2010, 76: 7566–7574
Krömer J O, Wittmann C, Schröder H, et al. Metabolic pathway analysis for rational design of l-methionine production by Escherichia coli and Corynebacterium glutamicum. Metab Eng, 2006, 8: 353–369
Vallino J J, Stephanopoulos G. Metabolic flux distributions in Corynebacterium glutamicum during growth and lysine overproduction. Biotechnol Bioeng, 1993, 41: 633–646
Radmacher E, Vaitsikova A, Burger U, et al. Linking central metabolism with increased pathway flux: l-valine accumulation by Corynebacterium glutamicum. Appl Environ Microbiol, 2002, 68: 2246–2250
Stauffer L T, Stauffer G V. Role for the leucine-responsive regulatory protein (Lrp) as a structural protein in regulating the Escherichia coli gcvTHP operon. Microbiology, 1999, 145: 569–576
Kalinowski J, Bathe B, Bartels D, et al. The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of l-aspartate-derived amino acids and vitamins. J Biotechnol, 2003, 104: 5–25
Han M J, Lee S Y. The Escherichia coli proteome: past, present, and future prospects. Microbiol Mol Biol Rev, 2006, 70: 362–439
Kjeldsen K R, Nielsen J. In silico genome-scale reconstruction and validation of the Corynebacterium glutamicum metabolic network. Biotechnol Bioeng, 2009, 102: 583–597
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Lai, S., Zhang, Y., Liu, S. et al. Metabolic engineering and flux analysis of Corynebacterium glutamicum for L-serine production. Sci. China Life Sci. 55, 283–290 (2012). https://doi.org/10.1007/s11427-012-4304-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-012-4304-0