Abstract
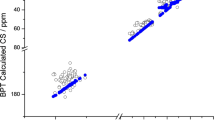
We performed density functional calculations of backbone 15N shielding tensors in the regions of beta-sheet and turns of protein G. The calculations were carried out for all twenty-four beta-sheet residues and eight beta-turn residues in the protein GB3 and the results were compared with the available experimental data from solid-state and solution NMR measurements. Together with the alpha-helix data, our calculations cover 39 out of the 55 residues (or 71%) in GB3. The applicability of several computational models developed previously (Cai et al. in J Biomol NMR 45:245–253, 2009) to compute 15N shielding tensors of alpha-helical residues is assessed. We show that the proposed quantum chemical computational model is capable of predicting isotropic 15N chemical shifts for an entire protein that are in good correlation with experimental data. However, the individual components of the predicted 15N shielding tensor agree with experiment less well: the computed values show much larger spread than the experimental data, and there is a profound difference in the behavior of the tensor components for alpha-helix/turns and beta-sheet residues. We discuss possible reasons for this.






Similar content being viewed by others
References
Becke AD (1988) Density-functional exchange-energy approximation with correct asymptotic behavior. Phys Rev A 38(6):3098–3100
Becke AD (1993) Density-functional thermochemistry. III. The role of exact exchange. J Chem Phys 98:5648–5652
Bernado P, Blackledge M (2004) Local dynamic amplitudes on the protein backbone from dipolar couplings: toward the elucidation of slower motions in biomolecules. J Am Chem Soc 126(25):7760–7761
Brender JR, Taylor DM, Ramamoorthy A (2001) Orientation of amide-nitrogen-15 chemical shift tensors in peptides: a quantum chemical study. J Am Chem Soc 123(5):914–922
Cai L, Fushman D, Kosov DS (2008) Density functional calculations of 15 N chemical shifts in solvated dipeptides. J Biomol NMR 41(2):77–88
Cai L, Fushman D, Kosov DS (2009) Density functional calculations of chemical shielding of backbone 15 N in helical residues of protein G. J Biomol NMR 45(3):245–253
Clore GM, Schwieters CD (2004) Amplitudes of protein backbone dynamics and correlated motions in a small alpha/beta protein: correspondence of dipolar coupling and heteronuclear relaxation measurements. Biochemistry 43(33):10678–10691
Cornell WD, Cieplak P, Bayly CI, Gould IR, Merz KM, Ferguson DM, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) A second generation force-field for the simulation of proteins, nucleic acids, and organic molecules. J Am Chem Soc 117:5179–5197
Cornilescu G, Delaglio F, Bax A (1999) Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J Biomol NMR 13(3):289–302
de Dios AC, Pearson JG, Oldfield E (1993) Secondary and tertiary structural effects on protein NMR chemical shifts: an ab initio approach. Science 260(5113):1491–1496
Derrick JP, Wigley DB (1994) The third IgG-binding domain from streptococcal protein G. An analysis by X-ray crystallography of the structure alone and in a complex with Fab. J Mol Biol 243(5):906–918
Franks WT, Zhou DH, Wylie BJ, Money BG, Graesser DT, Frericks HL, Sahota G, Rienstra CM (2005) Magic-angle spinning solid-state NMR spectroscopy of the beta1 immunoglobulin binding domain of protein G (GB1): 15 N and 13C chemical shift assignments and conformational analysis. J Am Chem Soc 127(35):12291–12305
Frisch M, Trucks G, Schlegel H, Scuseria G, Robb M, Cheeseman J, Montgomery J, Vreven T, Kudin K, Burant J, Millam J, Iyengar S, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson G, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox J, Hratchian H, Cross J, Adamo C, Jaramillo J, Gomperts R, Stratmann R, Yazyev O, Austin A, Cammi R, Pomelli C, Ochterski J, Ayala P, Morokuma K, Voth G, Salvador P, Dannenberg J, Zakrzewski V, Dapprich S, Daniels A, Strain M, Farkas O, Malick D, Rabuck A, Raghavachari K, Foresman J, Ortiz JQ, Baboul A, Clifford S, Cioslowski J, Stefanov B, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin R, Fox D, Keith T, Al-Laham M, Peng C, Nanayakkara A, Challacombe M, Gill P, Johnson B, Chen W, Wong M, Gonzalez C, Pople J (2004) Gaussian 03, Revision C.02. Gaussian, Inc, Wallingford, CT
Fushman D, Cowburn D (2001) Nuclear magnetic resonance relaxation in determination of residue-specific 15 N chemical shift tensors in proteins in solution: protein dynamics, structure, and applications of transverse relaxation optimized spectroscopy. Methods Enzymol 339:109–126
Fushman D, Tjandra N, Cowburn D (1998) Direct measurement of 15N chemical shift anisotropy in solution. J Am Chem Soc 120(42):10947–10952
Fushman D, Tjandra N, Cowburn D (1999) An approach to direct determination of protein dynamics from 15 N NMR relaxation at multiple fields, independent of variable 15 N chemical shift anisotropy and chemical exchange contributions. J Am Chem Soc 121:8577–8582
Gallagher T, Alexander P, Bryan P, Gilliland GL (1994) Two crystal structures of the B1 immunoglobulin-binding domain of streptococcal protein G and comparison with NMR. Biochemistry 33(15):4721–4729
Hall JB, Fushman D (2003) Characterization of the overall and local dynamics of a protein with intermediate rotational anisotropy: Differentiating between conformational exchange and anisotropic diffusion in the B3 domain of protein G. J Biomol NMR 27(3):261–275
Hall JB, Fushman D (2006) Variability of the 15 N chemical shielding tensors in the B3 domain of protein G from 15 N relaxation measurements at several fields. Implications for backbone order parameters. J Am Chem Soc 128(24):7855–7870
Harris RK, Becker ED, De Menezes SM, Granger P, Hoffman RE, Zilm KW (2008) Further conventions for NMR shielding and chemical shifts (IUPAC Recommendations 2008). Magn Reson Chem 46(6):582–598
Hiyama Y, Niu C, Silverton JV, Bavoso A, Torchia DA (1988) Determination of 15 N chemical shift tensor via 15 N–2H dipolar coupling in Boc-glycylglycyl[15 N glycine]benzyl ester. J Am Chem Soc 110:2378–2383
Jameson CJ, Jameson AK, Oppusunggu D, Wille S, Burrell PM, Mason J (1981) 15 N nuclear magnetic shielding scale from gas-phase studies. J Chem Phys 74:81–88
Le HB, Oldfield E (1996) Ab initio studies of amide-N-15 chemical shifts in dipeptides: applications to protein NMR spectroscopy. J Phys Chem 100:16423–16428
Lee DK, Ramamoorthy A (1998) A simple one-dimensional solid-state NMR method to characterize the nuclear spin interaction tensors associated with the peptide bond. J Magn Reson 133(1):204–206
Lee C, Yang W, Parr RG (1988) Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B Condens Matter 37(2):785–789
Lee D-K, Wei Y, Ramamoorthy A (2001) A Two-dimensional magic-angle decoupling and magic-angle turning solid-state NMR method: an application to study chemical shift tensors from peptides that are nonselectively labeled with 15 N isotope. J Phys Chem B 105(20):4752–4762
Lipsitz RS, Tjandra N (2003) 15 N chemical shift anisotropy in protein structure refinement and comparison with NH residual dipolar couplings. J Magn Reson 164(1):171–176
Markley JL, Bax A, Arata Y, Hilbers CW, Kaptein R, Sykes BD, Wright PE, Wuthrich K (1998) Recommendations for the presentation of NMR structures of proteins and nucleic acids. IUPAC-IUBMB-IUPAB Inter-union task group on the standardization of data bases of protein and nucleic acid structures determined by NMR spectroscopy. J Biomol NMR 12(1):1–23
Nadaud PS, Helmus JJ, Jaroniec CP (2007) C-13 and N-15 chemical shift assignments and secondary structure of the B3 immunoglobulin-binding domain of streptococcal protein G by magic-angle spinning solid-state NMR spectroscopy. Biomol NMR Assign 1:117–120
Neal S, Nip AM, Zhang H, Wishart DS (2003) Rapid and accurate calculation of protein 1H, 13C and 15 N chemical shifts. J Biomol NMR 26(3):215–240
Oas TG, Hartzell CJ, Dahlquist FW, Drobny GP (1987) The amide nitrogen-15 chemical shift tensors of four peptides determined from carbon-13 dipole-coupled chemical shift powder patterns. J Am Chem Soc 109:5962–5966
Oldfield E (1995) Chemical shifts and three-dimensional protein structures. J Biomol NMR 5(3):217–225
Poon A, Birn J, Ramamoorthy A (2004) How does an amide-N chemical shift tensor vary in peptides? J Phys Chem B 108(42):16577–16585
Ramamoorthy A, Wu CH, Opella SJ (1995) Three-dimensional solid-state NMR experiment that correlates the chemical shift and dipolar coupling frequencies of two heteronuclei. J Magn Reson B 107(1):88–90
Sadqi M, Fushman D, Munoz V (2006) Atom-by-atom analysis of global downhill protein folding. Nature 442(7100):317–321
Saito H, Ando I, Ramamoorthy A (2010) Chemical shift tensor—the heart of NMR: Insights into biological aspects of proteins. Prog Nucl Magn Reson Spectrosc 57(2):181–228
Shen Y, Bax A (2007) Protein backbone chemical shifts predicted from searching a database for torsion angle and sequence homology. J Biomol NMR 38(4):289–302
Shen Y, Lange O, Delaglio F, Rossi P, Aramini JM, Liu G, Eletsky A, Wu Y, Singarapu KK, Lemak A, Ignatchenko A, Arrowsmith CH, Szyperski T, Montelione GT, Baker D, Bax A (2008) Consistent blind protein structure generation from NMR chemical shift data. Proc Natl Acad Sci USA 105(12):4685–4690
Shen Y, Bryan PN, He Y, Orban J, Baker D, Bax A (2009a) De novo structure generation using chemical shifts for proteins with high sequence identity but different folds. Protein Sci 19(2): 349–356
Shen Y, Delaglio F, Cornilescu G, Bax A (2009b) TALOS + : a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44(4):213–223
Shen Y, Vernon R, Baker D, Bax A (2009c) De novo protein structure generation from incomplete chemical shift assignments. J Biomol NMR 43(2):63–78
Spronk CAEM, Nabuurs SB, Krieger E, Vriend G, Vuister GW (2004) Validation of protein structures derived by NMR spectroscopy. Progr NMR Spect 45:315–337
Ulmer TS, Ramirez BE, Delaglio F, Bax A (2003) Evaluation of backbone proton positions and dynamics in a small protein by liquid crystal NMR spectroscopy. J Am Chem Soc 125(30):9179–9191
Varadan R, Walker O, Pickart C, Fushman D (2002) Structural properties of polyubiquitin chains in solution. J Mol Biol 324(4):637–647
Varadan R, Assfalg M, Haririnia A, Raasi S, Pickart C, Fushman D (2004) Solution conformation of Lys63-linked di-ubiquitin chain provides clues to functional diversity of polyubiquitin signaling. J Biol Chem 279(8):7055–7063
Varadan R, Assfalg M, Raasi S, Pickart C, Fushman D (2005) Structural determinants for selective recognition of a Lys48-linked polyubiquitin chain by a UBA domain. Mol Cell 18(6):687–698
Vasos PR, Hall JB, Kümmerle R, Fushman D (2006) Measurement of 15 N relaxation in deuterated amide groups in proteins using direct nitrogen detection. J Biomol NMR 36(1):27–36
Vila JA, Scheraga HA (2008) Factors affecting the use of 13C(alpha) chemical shifts to determine, refine, and validate protein structures. Proteins 71(2):641–654
Villegas ME, Vila JA, Scheraga HA (2007) Effects of side-chain orientation on the 13C chemical shifts of antiparallel beta-sheet model peptides. J Biomol NMR 37(2):137–146
Wishart DS, Sykes BD, Richards FM (1992) The chemical shift index: a fast and simple method for the assignment of protein secondary structure through NMR spectroscopy. Biochemistry 31(6):1647–1651
Wishart DS, Bigam CG, Yao J, Abildgaard F, Dyson HJ, Oldfield E, Markley JL, Sykes BD (1995) 1H, 13C and 15 N chemical shift referencing in biomolecular NMR. J Biomol NMR 6(2):135–140
Wishart DS, Watson MS, Boyko RF, Sykes BD (1997) Automated 1H and 13C chemical shift prediction using the BioMagResBank. J Biomol NMR 10(4):329–336
Wylie BJ, Sperling LJ, Frericks HL, Shah GJ, Franks WT, Rienstra CM (2007) Chemical-shift anisotropy measurements of amide and carbonyl resonances in a microcrystalline protein with slow magic-angle spinning NMR spectroscopy. J Am Chem Soc 129(17):5318–5319
Wylie BJ, Schwieters CD, Oldfield E, Rienstra CM (2009) Protein structure refinement using 13C alpha chemical shift tensors. J Am Chem Soc 131(3):985–992
Xu XP, Case DA (2001) Automated prediction of 15 N, 13 Calpha, 13 Cbeta and 13 C’ chemical shifts in proteins using a density functional database. J Biomol NMR 21(4):321–333
Xu XP, Case DA (2002) Probing multiple effects on 15 N, 13C alpha, 13C beta, and 13C’ chemical shifts in peptides using density functional theory. Biopolymers 65(6):408–423
Yao L, Grishaev A, Cornilescu G, Bax A (2010) Site-specific backbone amide (15)N chemical shift anisotropy tensors in a small protein from liquid crystal and cross-correlated relaxation measurements. J Am Chem Soc 132(12):4295–4309
Zhang D, Raasi S, Fushman D (2008) Affinity makes the difference: nonselective interaction of the UBA domain of Ubiquilin-1 with monomeric ubiquitin and polyubiquitin chains. J Mol Biol 377(1):162–180
Zhang D, Chen T, Ziv I, Rosenzweig R, Matiuhin Y, Bronner V, Glickman MH, Fushman D (2009) Together, Rpn10 and Dsk2 Can Serve as a Polyubiquitin Chain-Length Sensor. Mol Cell 36:1018–1033
Zuiderweg ER (2002) Mapping protein-protein interactions in solution by NMR spectroscopy. Biochemistry 41(1):1–7
Acknowledgments
Supported by NIH grant GM065334 to DF and Francqui Foundation to DSK. We thank Gabriel Cornilescu, Chad Rienstra, and Ben Wylie for making available detailed experimental chemical shift data for comparison and Christopher Jaroniec for insightful discussions.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cai, L., Kosov, D.S. & Fushman, D. Density functional calculations of backbone 15N shielding tensors in beta-sheet and turn residues of protein G. J Biomol NMR 50, 19–33 (2011). https://doi.org/10.1007/s10858-011-9474-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-011-9474-8