Abstract
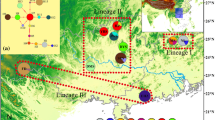
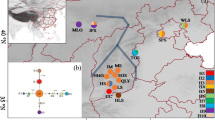
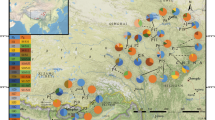
In order to investigate the genetic diversity and influence of climate oscillations on evolutionary processes of organisms in Northwest China, we selected Hexinia polydichotoma, a species endemic to China, and examined the phylogeographic structure and historical factors that influenced the evolutionary history of this species in its entire cover range, Tarim Basin and adjacent areas. In the study, 17 haplotypes were identified in H. polydichotoma on the basis of two chloroplast DNA sequences (trnH–psbA and ycf6–psbM). Shown in the network, the two common haplotypes, A and D, respectively, mainly distribute along the northern and southern rims of the basin. The analyses of molecular variance analysis suggest that genetic variation primarily occurs among populations, and all populations were subdivided into five groups by SAMOVA. Geographic range expansion along the southern and northern rims of the basin was supported by the significant value for Tajima’s D and by the unimodal mismatch distribution. It is possible that during the interglacial period of the middle Pleistocene, a large amount of snow and glacial ice melted from the mountains surrounding Tarim Basin. This increased water, the expanding desert, and the dispersal ability of H. polydichotoma were important factors driving not only geographic range expansion, but also the current phylogeographic structure of this species. It is possible that during the middle Pleistocene, the climatic fluctuations resulted in expansion and contraction cycles of river systems and oases, and may consequently have caused population fragmentation.




Similar content being viewed by others
References
Aizawa M, Yoshimaru H, Saito H, Katsuki T, Kawahara T, Kitamura K, Shi F, Sabirov R, Kaji M (2009) Range-wide genetic structure in a north-east Asian spruce (Picea jezoensis) determined using nuclear microsatellite markers. J Biogeogr 36:996–1007
Aris-Brosou S, Excoffier L (1996) The impact of population expansion and mutation rate heterogeneity on DNA sequence polymorphism. Mol Biol Evol 13:494–504
Avise JC (1987) Identification and interpretation of mitochondrial DNA stocks in marine species. In: Kumpf H, Nakamura EL (eds) Proceedings of the stock identification workshop. National Oceanographic and Atmospheric Administration, Panama City, FL, pp 105–136
Avise JC (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge, MA
Bandelt HJ, Forster P, Röhl A (1999) Median joining networks for inferring intraspecific phylogenies. Mol Biol Evol 14:37–48
Bittkau C, Comes HP (2005) Evolutionary processes in a continental island system: molecular phylogeography of the Aegean Nigella arvensis alliance (Ranunculaceae) inferred from chloroplast DNA. Mol Ecol 14:4065–4083
Broyles SB (1998) Postglacial migration and the loss of allozyme variation in Northern population of Asclepias exaltata (Asclepiadaceae). Am J Bot 85:1091–1097
Comes HP, Kadereit JW (1998) The effect of quaternary climatic changes on plant distribution and evolution. Trends Plant Sci 3:432–438
Cornman RS, Arnold ML (2007) Phylogeography of Iris missouriensis (Iridaceae) based on nuclear and chloroplast markers. Mol Ecol 16:4585–4598
Demesure B, Sodzi N, Petit RJ (1995) A set of universal primers for amplification of polymorphic non-coding regions of mitochondrial and chloroplast DNA in plants. Mol Ecol 4:129–131
Dong GR, Chen HZ, Wang GY, Li XZ, Shao YJ, Jin J (1997) The evolution of deserts with climatic changes in China since 150 ka BP. Sci China Ser D Earth Sci 40:370–382
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure from small quantities of fresh leaf tissues. Phytochem Bull 19:11–15
Dupanloup I, Schneider S, Excoffier L (2002) A simulated annealing approach to define the genetic structure of populations. Mol Ecol 11:2571–2581
Excoffier L, Smouse P, Quattro J (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: applications to human mitochondrial DNA restriction data. Genetics 131:479–491
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinforma 1:47–50
Feng Q, Su ZZ, Jin HJ (1999) Desert evolution and climatic changes in the Tarim River basin since 12 ka BP. Sci China Ser D Earth Sci 42:101–112
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking, and background selection. Genetics 147:915–925
Ge XJ, Hwang CC, Liu ZH, Huang CC, Huang WH, Hung KH, Wang WK, Chiang TY (2011) Conservation genetics and phylogeography of endangered and endemic shrub Tetraena mongolica (Zygophyllaceae) in Inner Mongolia, China. BMC Genet 12:1
Guo YP, Zhang R, Chen CY, Zhou DW, Liu JQ (2010) Allopatric divergence and regional range expansion of Juniperus sabina in China. J Syst Evol 48:153–160
Hewitt GM (2000) The genetic legacy of the Quaternary ice ages. Nature 405:907–913
Hewitt GM (2004) Genetic consequences of climatic oscillations in the Quaternary. Phil Trans R Soc Lond B Biol Sci 359:183–195
Huelsenbeck JP, Ronquist F (2001) MrBayes: a program for the Bayesian inference of phylogeny. Bioinformatics 17:754–755
Jaeger JR, Riddle BR, Bradford DF (2005) Cryptic neogene vicariance and Quaternary dispersal of the red-spotted toad (Bufo punctatus): insights on the evolution of North American warm desert biotas. Mol Ecol 14:3033–3048
Jin HL, Dong GR, Jin J, Li BS, Shao YJ (1994) Environmental and climatic changes in the interior of Taklimakan desert since Late Glacial Age. J Desert Res 14:31–37 (in Chinese with English abstract)
Jones ME, Shepherd M, Henry RJ, Delves A (2006) Chloroplast DNA variation and population structure in the widespread forest tree, Eucalyptus grandis. Conserv Genet 7:691–703
Li WC (1998) The Chinese Quaternary vegetation and environment. Science Press, Beijing
Li N, Fu L (1997) Notes on gymnosperms I. Taxonomic treatment of some Chinese conifers. Novon 7:261–264
Ling R, Shi Z (1997) Asteraceae. In: Wu ZY, Raven PH (eds) Flora of China, vol 80. Science Press, Beijing
Liu JQ, Qin XG (2005) Evolution of the environmental framework and oasis in the Tarim Basin. Quat Sci 25:533–539 (in Chinese with English abstract)
Luo C, Peng ZC, Yang D, Liu WG, Zhang ZF, He JF, Chou CL (2009) A lacustrine record from Lop Nur, Xinjiang, China: implications for paleoclimate change during Late Pleistocene. J Asian Earth Sci 34:38–45
Meng HH, Zhang ML (2011) Phylogeography of Lagochilus ilicifolius (Lamiaceae) in relation to Quaternary climatic oscillation and aridification in northern China. Biochem Syst Ecol 39:787–796
Mu GJ (1994) On the age and evolution of the Taklimakan desert. Arid Land Geogr 17:1–9 (in Chinese with English abstract)
Palmé AE, Semerikov V, Lascoux M (2003) Absence of geographical structure of chloroplast DNA variation in sallow, Salix caprea L. Heredity 91:465–474
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant-tissues. Plant Mol Biol 5:69–76
Rogers A, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol 9:552–569
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Sang T, Crawford DJ, Stuessy TF (1997) Chloroplast DNA phylogeny, reticulate evolution, and biogeography of Paeonia (Paeoniaceae). Am J Bot 84:1120–1136
Shi YF, Cui ZJ, Su Z (2005) The Quaternary glaciations and environmental variations in China. Hebei Science and Technology Publishing House, Hebei, pp 71–78
Simmons MP, Ochoterena H (2000) Gaps as characters in sequence-based phylogenetic analyses. Syst Biol 49:369–381
Slatkin M, Hudson RR (1991) Pairwise comparisons of mitochondrial DNA sequences in stable and exponentially growing populations. Genetics 129:555–562
Smith CI, Farrell BD (2005) Range expansions in the flightless longhorn cactus beetles, Moneilema gigas and Moneilema armatum, in response to Pleistocene climate changes. Mol Ecol 14:1025–1044
Soltis DE, Morris AB, McLachlan JS, Manos PS, Soltis PS (2006) Comparative phylogeography of unglaciated eastern North America. Mol Ecol 15:4261–4293
Sosa V, Ruiz-Sanchez E, Rodriguez-Gomez FC (2009) Hidden phylogeographic complexity in the Sierra Madre Oriental: the case of the Mexican tulip poppy Hunnemannia fumariifolia (Papaveraceae). J Biogeogr 36:18–27
Sun JM, Zhang LY, Deng CL, Zhu RX (2008) Evidence for enhanced aridity in the Tarim Basin of China since 5.3 Ma. Quat Sci Rev 27:1012–1023
Swofford DL (2002) PAUP*: phylogenetic analysis using parsimony (and other methods), version 4.0b10. Sinauer Associates, Sunderland
Taberlet P, Fumagalli L, Wust-Saucy AG, Cosson JF (1998) Comparative phylogeography and postglacial colonization routes in Europe. Mol Ecol 7:453–464
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tajima F (1996) The amount of DNA polymorphism maintained in a finite population when the neutral mutation rate varies among sites. Genetics 143:1457–1465
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal-W—improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Vidal-Russell R, Souto CP, Premoli AC (2011) Multiple Pleistocene refugia in the widespread Patagonian tree Embothrium coccineum (Proteaceae). Aust J Bot 59:299–314
Wang Y, Li S, Wang JH, Yan MC (1996) The uplift of the Qinghai-Xizang (Tibetan) Plateau and its effect on the formation and evolution of Chinese deserts. Arid Zone Res 13:20–24 (in Chinese with English abstract)
Wolfe KH, Li WH, Sharp PM (1987) Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAS. P Natl Acad Sci USA 84:9054–9058
Wu LL, Cui XK, Milne RI, Sun YS, Liu JQ (2010) Multiple autopolyploidizations and range expansion of Allium przewalskianum Regel. (Alliaceae) in the Qinghai-Tibetan Plateau. Mol Ecol 19:1691–1704
Wu YH, Xia L, Zhang Q, Yang QS, Meng XX (2011) Bidirectional introgressive hybridization between Lepus capensis and Lepus yarkandensis. Mol Phylogenet Evol 59:545–555
Yang XL (1992) Flora of desert from China. Science Press, Beijing
Yang XP, Zhu ZD, Jaekel D, Owen LA, Han JM (2002) Late Quaternary palaeoenvironment change and landscape evolution along the Keriya River, Xinjiang, China: the relationship between high mountain glaciation and landscape evolution in foreland desert regions. Quat Int 97–98:155–166
Zhang HY, Men GF (2002) Stratigraphic subdivision and climatic change of the Quaternary of the center Taklimakan Desert. Xinjiang Geol 20:256–261 (in Chinese with English abstract)
Zhang H, Wu JW, Zheng QH, Yu YJ (2003) A preliminary study of oasis evolution in the Tarim Basin, Xinjiang. China J Arid Environ 55:545–553
Zhang Q, Xia L, He JB, Fu JZ, Yang QS (2010) Comparison of phylogeographic structure and population history of two Phrynocephalus species in the Tarim Basin and adjacent areas. Mol Phylogenet Evol 57:1091–1104
Zhao YZ, Zhu ZY (2003) The endemic genera of desert region in the centre of Asia. Acta Botanica Yunnanica 25:113–121
Zheng HB, Powell CM, Butcher K, Cao JJ (2003) Late Neogene loess deposition in southern Tarim Basin: tectonic and palaeoenvironmental implications. Tectonophysics 375:49–59
Acknowledgments
We thank Honghu Meng and Jianfeng Huang at the Xinjiang Institute of Ecology and Geography, CAS, for their help with the experiment. Funding was provided by the CAS Important Direction for Knowledge Innovation Project (no. KZCX2-EW-305) and Xinjiang Institute of Ecology and Geography, CAS.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Su, Z., Zhang, M. & Cohen, J.I. Phylogeographic and demographic effects of Quaternary climate oscillations in Hexinia polydichotoma (Asteraceae) in Tarim Basin and adjacent areas. Plant Syst Evol 298, 1767–1776 (2012). https://doi.org/10.1007/s00606-012-0677-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-012-0677-6