Abstract
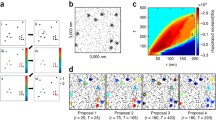
We demonstrate a combined univariate and bivariate Getis and Franklin’s local point pattern analysis method to investigate the co-clustering of membrane proteins in two-dimensional single-molecule localisation data. This method assesses the degree of clustering of each molecule relative to its own species and relative to a second species. Using simulated data, we show that this approach can quantify the degree of cluster overlap in multichannel point patterns. The method is validated using photo-activated localisation microscopy and direct stochastic optical reconstruction microscopy data of the proteins Lck and CD45 at the T cell immunological synapse. Analysing co-clustering in this manner is generalizable to higher numbers of fluorescent species and to three-dimensional or live cell data sets.





Similar content being viewed by others
References
Annibale P et al (2010) Photoactivatable fluorescent protein mEos2 displays repeated photoactivation after a long-lived dark state in the red photoconverted form. J Phys Chem Lett 1:1506–1510
Annibale P et al (2011a) Identification of clustering artifacts in photoactivated localization microscopy. Nat Methods 8(7):527–528
Annibale P et al (2011b) Quantitative photo activated localization microscopy: unraveling the effects of photoblinking. PLoS ONE 6(7):e22678
Annibale P et al (2012) Identification of the factors affecting co-localization precision for quantitative multicolor localization microscopy. Opt Nanoscopy 1(9):1–13
Betzig E et al (2006) Imaging intracellular fluorescent proteins at nanometer resolution. Science 313:1642–1645
Bolte S, Cordeliere FP (2006) A guided tour into subcellular colocalization analysis in light microscopy. J Microsc 224:213–232
Getis A, Franklin J (1987) Second-order neighbourhood analysis of mapped point patterns. Ecology 68(3):473–477
Heilemann M et al (2008) Subdiffraction-resolution fluorescence imaging with conventional fluorescent probes. Angew Chem Int Ed 47(33):6172–6176
Hell SW, Wichmann J (1994) Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt Lett 19(11):780–782
Huang B et al (2008) Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319(5864):810–813
Malkusch S et al (2012) Coordinate-based colocalization analysis of single-molecule localization microscopy data. Histochem Cell Biol 137(1):1–10
Owen DM et al (2010) PALM imaging and cluster analysis of protein heterogeneity at the cell surface. J Biophotonics 3(7):446–454
Owen MD et al (2013) Quantitative analysis of three-dimensional fluorescence localization microscopy data. Biophys J 105(2):L05–L07
Pavani SRP et al (2009) Three-dimensional, single-molecule fluorescence imaging beyond the diffraction limit by using a double-helix point spread function. Proc Natl Acad Sci 106(9):2995–2999
Ripley BD (1977) Modelling spatial patterns. J R Stat Soc B 39:172–192
Rossy J et al (2013) Conformational states of the kinase Lck regulate clustering in early T cell signalling. Nat Immunol 14(1):82–89
Rust MJ, Bates M, Zhuang X (2006) Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat Methods 3(10):793–796
Sengupta P, Lippincott-Schwartz J (2012) Quantitative analysis of photoactivated localization microscopy (PALM) datasets using pair-correlation analysis. BioEssays 34(5):396–405
Sengupta P et al (2011) Probing protein heterogeneity in the plasma membrane using PALM and pair correlation analysis. Nat Methods 8(11):969–975
Thompson RE, Larson DR, Webb WW (2002) Precise nanometer localization analysis for individual fluorescent probes. Biophys J 82(5):2775–2783
Veatch SL et al (2012) Correlation functions quantify super-resolution images and estimate apparent clustering due to over-counting. PLoS ONE 7(2):e31457
Vicidomini G et al (2011) Sharper low-power STED nanoscopy by time gating. Nat Methods 8(7):571–573
Williamson DJ et al (2011) Pre-existing clusters of the adaptor Lat do not participate in early T cell signalling events. Nat Immunol 12(7):655–662
Xu K, Babcock HP, Zhuang X (2012) Dual-objective STORM reveals three-dimensional filament organization in the actin cytoskeleton. Nat Methods 9(2):185–188
Acknowledgments
D.M.O. is supported by a Marie Curie Career Integration Grant (CIG) Ref 334303. KG is supported by the National Health and Medical Research Council of Australia and the Australian Research Council.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Rossy, J., Cohen, E., Gaus, K. et al. Method for co-cluster analysis in multichannel single-molecule localisation data. Histochem Cell Biol 141, 605–612 (2014). https://doi.org/10.1007/s00418-014-1208-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-014-1208-z