Abstract
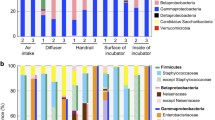
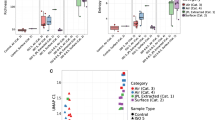
Potentially pathogenic microbes and so-called technophiles may form a serious threat in advanced life support systems, such as the International Space Station (ISS). They not only pose a threat to the health of the crew, but also to the technical equipment and materials of the space station. The development of fast and easy to use molecular detection and quantification methods for application in manned spacecraft is therefore desirable and may also be valuable for applications on Earth. In this paper we present the preliminary results of the SAMPLE experiment in which we performed molecular microbial analysis on environmental samples of the ISS as part of an ESA-MAP project.
Similar content being viewed by others
References
Pierson D. L.: Microbial contamination of spacecraft. Gravit. Space Biol. Bull. vol. 14, pp. 1–6 (2001).
Castro V. A., Thrasher A. N., Healy M., Ott C. M., Pierson D. L.: Microbial characterization during the early habitation of the International Space Station. Microb. Ecol. vol. 47, pp. 119–126 (2004).
Novikova N. D.: Review of the knowledge of microbial contamination of the Russian manned spacecraft. Microb. Ecol. vol. 47, pp. 127–132 (2004).
Ciferri O., Tiboni O., Di Pasquale G., Orlandoni A. M., Marchesi M. L.: Effects of microgravity on genetic recombination in Escherichia coli. Naturwissenschaften vol. 73, pp. 418–421 (1986).
Klaus D., Simske S., Todd P., Stodieck L.: Investigation of space flight effects onEscherichia coli and a proposed model of underlying physical mechanisms. Microbiology vol. 143, pp. 449–455 (1997).
Mennigmann H. D., Lange M.: Growth and differentiation of Bacillus subtilis under microgravity. Naturwissenschaften vol. 73, pp. 415–417 (1986).
Nickerson C. A., Ott C. M., Wilson J. W., Rantamurthy R., Pierson D. L.: Microbial responses to microgravity and other low-shear environments. Microbiol. Mol. Biol. Rev. vol. 68, pp. 345–361 (2004).
Tixador R., Gasset G., Eche B. et al.: Behavior of bacteria and antibiotics under space conditions. Aviat. Space Environ. Med. vol. 65, pp. 551–556 (1994).
Holdeman L. V., Good I. J., Moore W. E.: Human fecal flora: variation in bacterial composition within individuals and a possible effect of emotional stress. Appl. Environ. Microbiol. vol. 31, pp. 359–375 (1976).
Kaur I., Simons E. R., Castro V. A., Mark Ott C., Pierson D. L.: Changes in neutrophil functions in astronauts. Brain Behav. Immun. vol. 18, pp. 443–450 (2004).
Kaur I., Simons E. R., Castro V. A., Ott C. M., Pierson D. L.: Changes in monocyte functions of astronauts. Brain Behav. Immun. vol. 19, pp. 547–554 (2005).
Giulietti A., Overbergh L., Valckx D., Decallonne B., Bouillon R., Mathieu C.: An overview of real-time quantitative PCR: applications to quantify cytokine gene expression. Methods vol. 25, pp. 386–401 (2001).
Amann R. L., Ludwig W., Schleifer K. H.: Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. vol. 59, pp. 143–169 (1995).
Harmsen H. J. M., Raangs G. C., He T., Degener J. E., Welling G. W.: Extensive set of 16S rRNA-based probes for detection of bacteria in human feces. Appl. Environ. Microbiol. vol. 68, pp. 2982–2990 (2002).
Ludwig, W., Schleifer, K. H.: How quantitative is quantitative PCR with respect to cell counts? Syst. Appl. Microbiol. vol. 23, pp. 556–562 (2000).
Reischl, U., Linde, H. J., Metz, M., Leppmeier, B., Lehn, N.: Rapid identification of methicillin-resistantStaphylococcus aureus and simultaneous species confirmation using real-time fluorescence PCR. J. Clin. Microbiol. vol. 38, pp. 2429–2433 (2000).
Brakstad, O. G., Aasbakk, K., Maeland, J. A.: Detection ofStaphylococcus aureus by polymerase chain reaction amplification of thenuc gene. J. Clin. Microbiol. vol. 30, pp. 1654–1660 (1992).
Brasher C. W., De Paola, A., Jones, D. D., Bej, A. K.: Detection of microbial pathogens in shellfish with multiplex PCR. Curr. Microbiol. vol. 37, pp. 101–107 (1998).
Ballard, A. L., Fry, N. K., Chan, L., Surman, S. B., Lee, J. V., Harrison T. G., Towner, K. J.: Detection ofLegionella pneumophila using a Real-Time PCR hybridization assay. J. Clin. Microbiol. vol. 38, pp. 4215–4218 (2000).
Eishi, Y., Suga, M., Ishige, I., Kobayashi, D., Yamada, T., Takemura, T., Takizawa, T., Koike, M., Kudoh, S., Costabel, U., Guzman, J., Rizzato, G., Gambacorta, M., du Bois, R., Nicholson, A. G., Sharma, O. P., Ando, M.: Quantitative analysis of mycobacterial and propionibacterial DNA in lymph nodes of Japanese and European patients with sarcoidosis. J. Clin. Microb. vol. 40, pp. 198–204 (2002).
Costa, C., Vidaud, D., Olivi, M., Bart-Delabesse, E., Vidaud, M., Bretagne, S.: Development of two real-time quantitative Taqman PCR assays to detect circulatingAspergillus fumigatus DNA in serum. J. Microbiol. Methods, vol. 44, pp. 263–269 (2001).
De Vos, D., Lim, A. Jr.,Pirnay, J.-P., Struelens, M., Vandenvelde, C., Duinslaeger, L., Vanderkelen, A., Cornelis, P.: Direct detection and identification ofPseudomonas aeruginosa in clinical samples such as skin biopsy specimens and expectorations by multiplex PCR based on two outer membrane lipoprotein genes, oprl and oprl. J. Clin. Microbiol. vol. 35, pp. 1295–1299 (1997).
Nadkarni, M. A., Martin, F. E., Jacques, N. A., Hunter, N.: Determination of bacterial load by real-time PCR using a broad-range (universal) probe and primers set. Microbiology, vol. 148, pp. 257–266 (2002).
Lindsley, M. D., Hurst S. F., Iqbal, N. J., Morrison, C. J.: Rapid identification of dimorphic and yeast-like fungal pathogens using specific DNA probes. J. Clin. Microbiol. vol. 39, pp. 3505–3511 (2001).
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is a reprint of Microgravity sci. technol. XVIII-3/4 (2006). Tab. 1 has been corrected.
Rights and permissions
About this article
Cite this article
van Tongeren, S.P., Krooneman, J., Raangs, G.C. et al. Microbial detection and monitoring in advanced life support systems like the International Space Station. Microgravity sci. Technol. 19, 45–48 (2007). https://doi.org/10.1007/BF02911866
Issue Date:
DOI: https://doi.org/10.1007/BF02911866