Summary
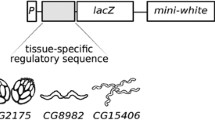
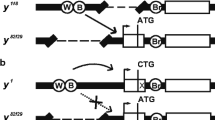
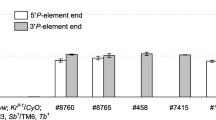
We have assessed the DNA sequence requirements for the correct spatial pattern and phenotypic expression of y in the late embryo/larvae. The wild-type larval phenotype requires both the regions between-294 bp and-92 bp and a portion of the intron; the sequence element(s) located within the intron can act in a position independent manner to effect the wild-type larval phenotype. The larval expression pattern was examined by tissue experiments in situ and by staining germline transformants derived from various y/lacZ fusion constructs. The larval expression of y is restricted to the mouthparts, microsetae and anal plates. While the-495 bp to+194 bp region alone cannot effect a wild-type larval expression pattern, this region in conjunction with the intron appears to be sufficient to drive β-gal expression in an essentially wild-type pattern. Our data further suggest that the-294 bp to-92 bp region contains elements which specify the larval pattern and that the element(s) in the intron normally act to enhance the level of expression necessary for the wild-type larval phenotype. We also present a phenotypic analysis of the adult cuticle structures of germline transformants derived from a variety of deletion and rearrangement constructs of the y gene. This analysis has revealed several new features associated with the regulation of y expression.
Similar content being viewed by others
References
Akam M (1983) The location of Ultrabithorax transcripts in Drosophila tissue sections. EMBO J 2:2075–2084
Biessmann H (1985) Molecular analysis of the yellow gene (y) region of Drosophila melanogaster. Proc Natl Acad Sci USA 82:7369–7373
Biessmann H, Mason JM (1988) Progressive loss of DNA sequences from terminal chromosome deficiencies in Drosophila melanogaster. EMBO J 7:1081–1086
Burnet B, Wilson R (1980) Pattern mosaicism for behaviour controlled by the yellow locus in Drosophila melanogaster. Genet Res 36:235–247
Campuzano S, Carramolino L, Cabrera CV, Ruiz-Gomez M, Villares R, Boronat A, Modelell J (1985) Molecular genetics of the achaete-scute gene complex of D. melanogaster. Cell 40:327–338
Chia W, Karp R, McGill S, Ashburner M (1985) Molecular analysis of the Adh region of the genome of Drosophila melanogaster. J Mol Biol 186:689–706
Chia W, Howes G, Martin M, Meng YB, Moses K, Tsubota S (1986) Molecular analysis of the yellow locus of Drosophila. EMBO J 5:3597–3605
Geyer PK, Corces VG (1987) Separate regulatory elements are responsible for the complex pattern of tissue-specific and developmental transcription of the yellow locus in Drosophila melanogaster. Genes Dev 1:996–1004
Geyer P, Spana C, Corces VG (1986) On the molecular mechanism of gypsy-induced mutations at the yellow locus of Drosophila melanogaster. EMBO J 5:2657–2662
Giangrande A, Mettling C, Richards G (1987) Sgs-3 transcript levels are determined by multiple remote sequence elements. EMBO J 6:3079–3084
Green MM (1961) Complementation at the Yellow locus in Drosophila melanogaster. Genetics 46:1385–1388
Hafen E, Levine M (1986) The localization of RNAs in Drosophila tissue sections by in situ hybridization. In: Roberts DB (ed) Drosophila — a practical approach. IRL Press, Oxford, pp 139–157
Hiromi Y, Kuroiwa A, Gehring WJ (1985) Control elements of the Drosophila segmentation gene fushi tarazu. Cell 43:603–613
Howes G, O'Connor M, Chia W (1988) On the specificity and effects on transcription of P-element insertions at the yellow locus of Drosophila melanogaster. Nucleic Acid Res 16:3039–3052
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Meyerowitz EM, Raghavan KV, Mathers PH, Roark M (1987) How Drosophila larvae make glue: control of Sgs-3 gene expression. TIG 3:288–293
Nash WG (1976) Patterns of pigmentation color states regulated by the y locus in Drosophila melanogaster. Dev Biol 48:336–343
Nash WG, Yarkin RJ (1974) Genetic regulation and pattern formation: A study of the yellow locus in Drosophila melanogaster. Genet Res 24:19–26
Parkhurst SM, Corces VG (1985) Forked, Gypsys, and suppressors in Drosophila. Cell 41:429–437
Parkhurst SM, Corces VG (1986) Interactions among the gypsy transposable element and the yellow and the suppressor of hairy-wing loci in Drosophila melanogaster. Mol Cell Biol 6:47–53
Robertson HM, Preston CR, Phillis RW, Johnson-Schlitz DM, Benz WK, Engels WR (1988) A stable genomic source of P element transposase in Drosophila melanogaster. Genetics 118:461–470
Rubin GM, Spradling AC (1982) Genetic transformation of Drosophila with transposable element vector. Science 218:348–353
Spradling AC, Rubin GM (1982) Transposition of cloned P elements into Drosophila germ line chromosomes. Science 218:341–348
Steller H, Pirrotta V (1985) A transposable P vector that confers selectable G418 resistance to Drosophila larvae. EMBO J 4:167–171
Steller H, Pirrotta V (1986) P transposons controlled by the heat shock promoter. Mol Cell Biol 6:1640–1649
Vijay Raghavan K, Crosby MA, Mathers PH, Meyerowitz EM (1986) Sequences sufficient for correct regulation of Sgs-3 lie close to or within the gene. EMBO J 5:3321–3326
Wilson R, Burnet B, Eastwood L, Connolly K (1976) Behavioural pleiotropy of the yellow gene in Drosophila melanogaster. Genet Res 28:75–88
Author information
Authors and Affiliations
Additional information
Communicated by d.J. Finnegan
Rights and permissions
About this article
Cite this article
Martin, M., Meng, Y.B. & Chia, W. Regulatory elements involved in the tissue-specific expression of the yellow gene of Drosophila . Mol Gen Genet 218, 118–126 (1989). https://doi.org/10.1007/BF00330574
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00330574