Abstract
Background
Hippocalcin-like 1 (HPCAL1), a neuronal calcium sensor protein family member, has been reported to regulate cancer growth. As yet, however, the biological functions of HPCAL1 and its molecular mechanisms have not been investigated in non-small cell lung carcinoma (NSCLC).
Methods
HPCAL1 expression in NSCLC samples was detected using immunohistochemistry, Western blotting and RT-PCR. The anticancer effects of HPCAL1 knockdown were determined by MTT, soft agar, cell cycle, oxygen consumption and reactive oxygen species assays. The effect of HPCAL1 knockdown on in vivo tumor growth was assessed using NSCLC cancer patient-derived xenograft models. Potentially interacting protein partners of HPCAL1 were identified using IP-MS/MS, immunoprecipitation and Western blotting assays. Metabolic alterations resulting from HPCAL1 knockdown were investigated using non-targeted metabolomics and RNA sequencing analyses.
Results
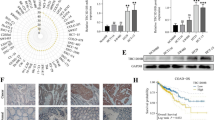
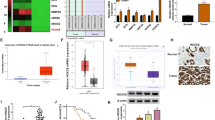
We found that HPCAL1 is highly expressed in NSCLC tissues and is positively correlated with low survival rates and AJCC clinical staging in lung cancer patients. Knockdown of HPCAL1 strongly increased oxygen consumption rates and the production of reactive oxygen species. HPCAL1 knockdown also inhibited NSCLC cell growth and patient-derived NSCLC tumor growth in vivo. Mechanistically, we found that HPCAL1 can directly bind to LDHA and enhance SRC-mediated phosphorylation of LDHA at tyrosine 10. The metabolomics and RNA sequencing analyses indicated that HPCAL1 knockdown reduces amino acid levels and induces fatty acid synthesis through regulating the expression of metabolism-related genes. Additionally, rescued cells expressing wild-type or mutant LDHA in HPCAL1 knockdown cells suggest that LDHA may serve as the main substrate of HPCAL1.
Conclusions
Our data indicate that the effect of HPCAL1 knockdown on reducing SRC-mediated LDHA activity attenuates NSCLC growth. Our findings reveal novel biological functions and a mechanism underlying the role of HPCAL1 in NSCLC growth in vitro and in vivo.






Similar content being viewed by others
Abbreviations
- HPCAL1:
-
hippocalcin-like 1
- LDHA:
-
lactate dehydrogenase A
- SRC:
-
proto-oncogene tyrosine-protein kinase Src
- NSCLC:
-
non-small cell lung carcinoma
References
F. Bray, J. Ferlay, I. Soerjomataram, R.L. Siegel, L.A. Torre, A. Jemal, CA Cancer J Clin 68, 394–424 (2018). https://doi.org/10.3322/caac.21492
X. Gu, G. Wang, H. Shen, X. Fei, Exp Ther Med 16, 3220–3226 (2018). https://doi.org/10.3892/etm.2018.6565
A. Roointan, T. Ahmad Mir, S. Ibrahim Wani, R. Mati Ur, K.K. Hussain, B. Ahmed, S. Abrahim, A. Savardashtaki, G. Gandomani, M. Gandomani, R. Chinnappan, M.H. Akhtar, J Pharm Biomed Anal 164, 93–103 (2018). https://doi.org/10.1016/j.jpba.2018.10.017
J.Z. Huang, M. Chen, X.C. Chen, S. Gao, H. Zhu, M. Huang, H.Z. Hu, G.R. Yan, Mol Cell 68, 171–184 e176 (2017). https://doi.org/10.1016/j.molcel.2017.09.015
J. Liu, G. Chen, Z. Liu, S. Liu, Z. Cai, P. You, Y. Ke, L. Lai, Y. Huang, H. Gao, L. Zhao, H. Pelicano, P. Huang, W.L. McKeehan, C.L. Wu, C. Wang, W. Zhong, F. Wang, Cancer Res 78, 4459–4470 (2018). https://doi.org/10.1158/0008-5472.CAN-17-3226
Y. Ji, C. Yang, Z. Tang, Y. Yang, Y. Tian, H. Yao, X. Zhu, Z. Zhang, J. Ji, X. Zheng, Nat Commun 8, 15308 (2017). https://doi.org/10.1038/ncomms15308
Y. Feng, Y. Xiong, T. Qiao, X. Li, L. Jia, Y. Han, Cancer Med 7, 6124–6136 (2018). https://doi.org/10.1002/cam4.1820
R. Arseneault, A. Chien, J.T. Newington, T. Rappon, R. Harris, R.C. Cumming, Cancer Lett 338, 255–266 (2013). https://doi.org/10.1016/j.canlet.2013.03.034
F. Jafary, M.R. Ganjalikhany, A. Moradi, M. Hemati, S. Jafari, Sci Rep 9, 4686 (2019). https://doi.org/10.1038/s41598-019-38854-7
G. Pathria, D.A. Scott, Y. Feng, J. Sang Lee, Y. Fujita, G. Zhang, A.D. Sahu, E. Ruppin, M. Herlyn, A.L. Osterman, Z.A. Ronai, EMBO J 37, e99735 (2018). https://doi.org/10.15252/embj.201899735
Z. Huang, N. Ma, Y.L. Xiong, L. Wang, W.M. Li, Y.Y. Lai, C.X. Zhang, Z.P. Zhang, X.F. Li, J.B. Zhao, Onco Targets Ther 12, 10299–10309 (2019). https://doi.org/10.2147/OTT.S210014
L. Jin, J. Chun, C. Pan, G.N. Alesi, D. Li, K.R. Magliocca, Y. Kang, Z.G. Chen, D.M. Shin, F.R. Khuri, J. Fan, S. Kang, Oncogene 36, 3797–3806 (2017). https://doi.org/10.1038/onc.2017.6
D. Zhao, S.W. Zou, Y. Liu, X. Zhou, Y. Mo, P. Wang, Y.H. Xu, B. Dong, Y. Xiong, Q.Y. Lei, K.L. Guan, Cancer Cell 23, 464–476 (2013). https://doi.org/10.1016/j.ccr.2013.02.005
J. Fan, T. Hitosugi, T.W. Chung, J. Xie, Q. Ge, T.L. Gu, R.D. Polakiewicz, G.Z. Chen, T.J. Boggon, S. Lonial, F.R. Khuri, S. Kang, J. Chen, Mol Cell Biol 31, 4938–4950 (2011). https://doi.org/10.1128/mcb.06120-11
X.M. Li, W.H. Xiao, H.X. Zhao, Med Chem Comm 8, 599–605 (2017). https://doi.org/10.1039/c6md00670a
K.H. Braunewell, A.J. Klein-Szanto, Cell Tissue Res 335, 301–316 (2009). https://doi.org/10.1007/s00441-008-0716-3
R.D. Burgoyne, Nat Rev Neurosci 8, 182–193 (2007). https://doi.org/10.1038/nrn2093
W. Wang, Q. Zhong, L. Teng, N. Bhatnagar, B. Sharma, X. Zhang, W. Luther 2nd, L.P. Haynes, R.D. Burgoyne, M. Vidal, S. Volchenboum, D.E. Hill, R.E. George, Oncogene 33, 3316–3324 (2014). https://doi.org/10.1038/onc.2013.290
D. Zhang, X. Liu, X. Xu, J. Xu, Z. Yi, B. Shan, B. Liu, J Cell Mol Med 23, 3108–3117 (2019). https://doi.org/10.1111/jcmm.14083
Y. Zhang, Y. Liu, J. Duan, H. Yan, J. Zhang, H. Zhang, Q. Fan, F. Luo, G. Yan, K. Qiao, J. Liu, Hepatology 63, 880–897 (2016). https://doi.org/10.1002/hep.28395
A.F. Gazdar, J.D. Minna, J Natl Cancer Inst 91, 299–301 (1999). https://doi.org/10.1093/jnci/91.4.299
C.V. Dang, Genes Development 26, 877–890 (2012). https://doi.org/10.1101/gad.189365.112
H. Makinoshima, M. Takita, S. Matsumoto, A. Yagishita, S. Owada, H. Esumi and K. Tsuchihara, J Biol Chem 289, 20813-20823 (2014). https://doi.org/10.1074/jbc.M114.575464
H.Y. Min, H.Y. Lee, Biomol Therap 26, 45–56 (2018). https://doi.org/10.4062/biomolther.2017.211
A. Martin-Bernabe, R. Cortes, S.G. Lehmann, M. Seve, M. Cascante, S. Bourgoin-Voillard, J Prot Res 13, 4695–4704 (2014). https://doi.org/10.1021/pr500327v
J.R. Doherty, J.L. Cleveland, J Clin Invest 123, 3685–3692 (2013). https://doi.org/10.1172/JCI69741
G. Kayser, A. Kassem, W. Sienel, L. Schulte-Uentrop, D. Mattern, K. Aumann, E. Stickeler, M. Werner, B. Passlick, A.Z. Hausen, Diagn Pathol 5, 22 (2010). https://doi.org/10.1186/1746-1596-5-22
M.I. Koukourakis, A. Giatromanolaki, C. Simopoulos, A. Polychronidis, E. Sivridis, Clin Exp Metast 22, 25–30 (2005). https://doi.org/10.1007/s10585-005-2343-7
H. Xie, J. Hanai, J.G. Ren, L. Kats, K. Burgess, P. Bhargava, S. Signoretti, J. Billiard, K.J. Duffy, A. Grant, X. Wang, P.K. Lorkiewicz, S. Schatzman, M. Bousamra 2nd, A.N. Lane, R.M. Higashi, T.W. Fan, P.P. Pandolfi, V.P. Sukhatme, P. Seth, Cell Metab 19, 795–809 (2014). https://doi.org/10.1016/j.cmet.2014.03.003
A. Le, C.R. Cooper, A.M. Gouw, R. Dinavahi, A. Maitra, L.M. Deck, R.E. Royer, D.L. Vander Jagt, G.L. Semenza, C.V. Dang, Proc Natl Acad Sci USA 107, 2037–2042 (2010). https://doi.org/10.1073/pnas.0914433107
M.G. Vander Heiden, L.C. Cantley, C.B. Thompson, Science 324, 1029–1033 (2009). https://doi.org/10.1126/science.1160809
Z. Li, H. Zhang, Cell Mol Life Sci 73, 377–392 (2016). https://doi.org/10.1007/s00018-015-2070-4
N.N. Pavlova, C.B. Thompson, Cell Metab 23, 27–47 (2016). https://doi.org/10.1016/j.cmet.2015.12.006
A. Fang, Q. Zhang, H. Fan, Y. Zhou, Y. Yao, Y. Zhang, X. Huang, Med Chem Comm 8, 1720–1726 (2017). https://doi.org/10.1039/c7md00222j
K. Vanhove, E. Derveaux, G.J. Graulus, L. Mesotten, M. Thomeer, J.P. Noben, W. Guedens, P. Adriaensens, Int J Mol Sci 20, 252 (2019). https://doi.org/10.3390/ijms20020252
Acknowledgements
We greatly appreciate the help of Mrs. Ran Yang for preparing immunohistochemistry slides.
Availability of data and materials
Supplementary figures (1-9), supplementary table (1-5), supplementary methods and associated figure legends are provided as supplementary material and are available online with the paper.
Funding
This work was supported by the Henan Joint Fund, China [grant number U1804196], the General project, National Natural Science Foundation China (NSFC) [grant number 82073075, 81872335, and 82103193] and the Youth Science Foundation of Natural Science Foundation of Henan Province, China [grant number 212300410315].
Author information
Authors and Affiliations
Contributions
X.W. performed the in vitro experiments and assisted in the cell-based and in vivo experiments, and prepared the manuscript; X.X., Y.Z. and M.P. assisted in the cell-based assays and in vivo studies; F.M. assisted in the in vivo studies; K.V.L., X.L. and K.L. assisted in data analysis and editing the manuscript; Z.D. supervised the overall experimental design; D.J.K. developed the idea, supervised designed experiments.
Corresponding authors
Ethics declarations
Ethics approval
All experiments involving animals were performed with permission and under strict guidance of the Zhengzhou University Institutional Animal Care and Use Committee (Zhengzhou, Henan, China). A written consent to participate for these studies was provided by all participants.
Consent for publication
All authors have given their consent for the publication of this article
Competing financial interest statement
None of the authors have any competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 20 kb)
ESM 2
Supplementary Figure 1. The expression of HPCAL1 in stable HPCAL1 knockdown cell lines. Cells were infected with shControl or shHPCAL1 and selected with puromycin for 2 days. The expression level of HPCAL1 (shControl, shHPCAL1 #2 and shHPCAL1 #4) in A549 and H460 cell lines was detected by Western blotting. (JPG 433 kb)
ESM 3
Supplementary Figure 2. The effect of HPCAL1 knockdown on colony formation ability and cell cycle distribution. (A and B) Representative images of (A) colony formation and (B) cell cycle distribution in stable HPCAL1 knockdown and shControl cell lines. (JPG 2284 kb)
ESM 4
Supplementary Figure 3. Effect of HPCAL1 overexpression on A549 NSCLC cell growth. (A) The expression of HPCAL1 in cells stably expressing the HPCAL1-flag vector. The expression of HPCAL1 and flag was detected by Western blotting. (B) Effect of HPCAL1 over-expression on cell growth. Cells were seeded and incubated for 24, 48, or 72 h. Cell growth was measured at an absorbance of 570 nm. (C). Effect of HPCAL1 over-expression on anchorage-independent cell growth. Cells were seeded and incubated for 2 weeks. Colonies were photographed using an inverted microscope and the number of colonies was quantified with Image-Pro PLUS (v.6) computer software program; representative images are shown. For B and C, data are shown as means ± S.D. of triplicate values from 3 independent experiments and the asterisks indicate a significant difference (*, p < 0.05). (JPG 1972 kb)
ESM 5
Supplementary Figure 4. The effect of HPCAL1 knockdown on NSCLC PDX tumor growth and mice body weight. (A) HPCAL1 protein levels in various NSCLC PDX tissues were assessed by Western blotting. (B) The effect of HPCAL1 knockdown on LG14, LG52, or LG55 NSCLC PDX tissues. (C) The effect of HPCAL1 knockdown on mice body weight. Data are shown as means ± S.E. of values obtained from experiments. (JPG 3090 kb)
ESM 6
Supplementary Figure 5. Immunoprecipitation of HPCAL1 in H1299 NSCLC cells. (A and B) Cells were transfected and incubated for 48 h. The HPCAL1-flag protein (A, 1st IP experiment; B, 2nd IP experiment) was immunoprecipitated and the gels were stained with Coomassie blue. (JPG 1143 kb)
ESM 7
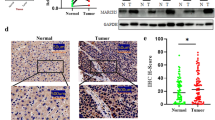
Supplementary Figure 6. Expression of LDHA and phosphorylated LDHA in normal, adjacent, and NSCLC tissues. (A and B) The expression of (A) LDHA and (B) phosphorylated LDHA protein in normal, adjacent, and NSCLC tissues was analyzed by immunohistochemistry (N, normal; AT, adjacent tissue; T, cancer tissue). Tissues stained with antibodies specific for LDHA or phosphorylated LDHA were photographed using an inverted microscope; staining intensity was quantified using the Image-Pro PLUS (v.6) computer software program. Quantification of LDHA and phosphorylated LDHA protein levels based on staining intensity is shown as a dot graph. For A and B, the asterisks (*, **) indicate a significant difference (p < 0.05, 0.01) respectively. (JPG 2401 kb)
ESM 8
Supplementary Figure 7. Effect of HPCAL1 knockdown on the expression of LDHA upstream kinases and its interaction with LDHA. (A) The protein expression levels of phosphorylated and total-SRC, -FGFR1, and -ERBB2 were measured in HPCAL1 knockdown and shControl cell lines by Western blotting. (B, C) Effect of HPCAL1 knockdown on the interaction between LDHA upstream kinases and LDHA. Cells were transfected with LDHA and ERBB2-flag (B) or FGFR1-flag (C) for 48 h. LDHA protein was immunoprecipitated in shControl and shHPCAL1 H1299 NSCLC cells. Expression of flag and myc was detected by Western blotting. Similar results were obtained from 3 independent experiments. (JPG 2560 kb)
ESM 9
Supplementary Figure 8. HPCAL1-mediated gene profiling. (A) Heatmap visualizing the HPCAL1 knockdown-mediated gene profile in H1299 NSCLC cells. Stable HPCAL1 knockdown or shControl cell lines were seeded for 48 h and then harvested. Differentially expressed genes identified by RNA sequencing were clustered (n = 3). (B) KEGG pathway classification of HPCAL1 knockdown-mediated genes. Differentially expressed genes were annotated. (JPG 1542 kb)
ESM 10
Supplementary Figure 9. Effect of glutamine deprivation on cell growth in HCPAL1 knockdown or shControl cells. Stable HPCAL1 knockdown cells were seeded and incubated for 24 h. Cell culture medium was replaced with glutamine-free medium supplemented with 10% FBS. Cell growth was analyzed by MTT assay. (JPG 795 kb)
ESM 11
Supplementary Table 1. The list of primers used for RT-PCR and gene cloning. (XLSX 11 kb)
ESM 12
Supplementary Table 2. The clinical parameters associated with HPCAL1 expression. (DOCX 15 kb)
ESM 13
Supplementary Table 3. The IP-MS/MS analysis in A549 NSCLC cells stably expressing HPCAL1-flag or Control-flag. (XLSX 26 kb)
ESM 14
Supplementary Table 4. The list of metabolic changes observed by HPCAL1 knockdown in H1299 NSCLC cells. (XLSX 51 kb)
ESM 15
Supplementary Table 5. The list of genes affected by HPCAL1 knockdown in H1299 NSCLC cells. (XLSX 947 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Xie, X., Zhang, Y. et al. Hippocalcin-like 1 is a key regulator of LDHA activation that promotes the growth of non-small cell lung carcinoma. Cell Oncol. 45, 179–191 (2022). https://doi.org/10.1007/s13402-022-00661-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-022-00661-0