Abstract
Purpose
The aims of the current study were to investigate the frequency and genetic diversity of Staphylococcus aureus from healthy horses, including both methicillin-resistant (MRSA) and -susceptible S. aureus (MSSA).
Methods
Three hundred-one nasal swabs were collected from healthy horses in three provinces, Iran. Sixty-one of the 301 tested samples contained S. aureus (20.3%), among which five were MRSA. Isolates were typed by spa PCR-RFLP and agr typing, followed by sequence-based spa typing and MLST on representative strains from each restriction pattern and SCCmec typing for MRSA strains. The presence of Panton-Valentine Leukocidin (PVL) encoding genes was also tested using PCR.
Results
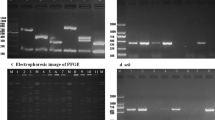
Eight distinct RFLP patterns (designated as N1-N8) were observed, with N2 (23/61; 37.7%) and N4 (18/61; 29.5%) the most common. On sequencing, N1-N8 patterns were found to be of clonal types ST15-t084, ST2151-t2484, ST291-t937, ST1-t127, and ST1-t1383, ST700-t11926, ST133-t1166, and ST1278-t12595, respectively. No PVL-positive S. aureus were detected. Five MRSA were identified as ST2151-t2484-SCCmecIVa (2 isolates), ST15-t084-SCCmecIVa, ST1-t1383-SCCmecIVa, and t12595-SCCmecIVa (one isolate each). Majority of S. aureus isolates were ascribed to agr types III (n = 30; 49.2%) and IV (n = 28; 45.9%), followed by types II (n = 2, 3.3%) and I (n = 1, 1.6%). The carriage of S. aureus was found to be associated with geographic locations.
Conclusions
This study for the first time describes the circulation of diverse clones of MSSA and MRSA among the Iranian horse population. This may pose a public health risk, which supports the need for their epidemiological monitoring.



Similar content being viewed by others
References
Agabou A, Ouchenane Z, Ngba Essebe C, Khemissi S, Chehboub MTE, Chehboub IB, Sotto A, Dunyach-Remy C, Lavigne JP (2017) Emergence of nasal carriage of ST80 and ST152 PVL+ Staphylococcus aureus isolates from livestock in Algeria. Toxins 9
AL-Tam F, Brunel AS, Bouzinbi N, Corne P, Banuls AL, Shahbazkia HR (2012) DNAGear-a free software for spa type identification in Staphylococcus aureus. BMC Res Notes 5:642
Asghar AH (2014) Molecular characterization of methicillin-resistant Staphylococcus aureus isolated from tertiary care hospitals. Pak J Med Sci 30:698–702
Axon J, Carrick J, Barton M, Collins N, Russell C, Kiehne J, Coombs G (2011) Methicillin-resistant Staphylococcus aureus in a population of horses in Australia. Aust Vet J 89:221–225
Barbagelata MS, Alvarez L, Gordiola M, Tuchscherr L, von Eiff C, Becker K, Sordelli D, Buzzola F (2011) Auxotrophic mutant of Staphylococcus aureus interferes with nasal colonization by the wild type. Microbes Infect 13:1081–1090
Basanisi MG, La Bella G, Nobili G, Franconieri I, La Salandra G (2017) Genotyping of methicillin-resistant Staphylococcus aureus (MRSA) isolated from milk and dairy products in South Italy. Food Microbiol 62:141–146
Boye K, Bartels MD, Andersen IS, Moller JA, Westh H (2007) A new multiplex PCR for easy screening of methicillin-resistant Staphylococcus aureus SCCmec types I-V. Clin Microbiol Infect 13:725–727
Brakstad OG, Aasbakk K, Maeland JA (1992) Detection of Staphylococcus aureus by polymerase chain reaction amplification of the nuc gene. J Clin Microbiol 30:1654–1660
Burton S, Reid-Smith R, McClure JT, Weese JS (2008) Staphylococcus aureus colonization in healthy horses in Atlantic Canada. Can Vet J 49:797–799
Carfora V, Caprioli A, Grossi I, Pepe M, Alba P, Lorenzetti S, Amoruso R, Sorbara L, Franco A, Battisti A (2016) A methicillin-resistant Staphylococcus aureus (MRSA) sequence type 8, spa type t11469 causing infection and colonizing horses in Italy. Pathog Dis 74:ftw025
CLSI (2009) Performance standards for antimicrobial Susceptability testing. Clinical and laboratory standards institute. Wayne, PA
Cohn LA, Middleton JR (2010) A veterinary perspective on methicillin-resistant staphylococci. J Vet Emerg Crit Care 20:31–45
Cuny C, Strommenger B, Witte W, Stanek C (2008) Clusters of infections in horses with MRSA ST1, ST254, and ST398 in a veterinary hospital. Microb Drug Resist 14:307–310
Cuny C, Friedrich A, Kozytska S, Layer F, Nubel U, Ohlsen K, Strommenger B, Walther B, Wieler L, Witte W (2010) Emergence of methicillin-resistant Staphylococcus aureus (MRSA) in different animal species. Int J Med Microbiol 300:109–117
Enright MC, Day NP, Davies CE, Peacock SJ, Spratt BG (2000) Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J Clin Microbiol 38:1008–1015
Fasihi Y, Kiaei S, Kalantar-Neyestanaki D (2017) Characterization of SCCmec and spa types of methicillin-resistant Staphylococcus aureus isolates from health-care and community-acquired infections in Kerman, Iran. J Epidemiol Glob Health 7:263–267
Gharsa H, Ben Sallem R, Ben Slama K, Gomez-Sanz E, Lozano C, Jouini A, Klibi N, Zarazaga M, Boudabous A, Torres C (2012) High diversity of genetic lineages and virulence genes in nasal Staphylococcus aureus isolates from donkeys destined to food consumption in Tunisia with predominance of the ruminant associated CC133 lineage. BMC Vet Res 8:203
Gilot P, Lina G, Cochard T, Poutrel B (2002) Analysis of the genetic variability of genes encoding the RNA III-activating components Agr and TRAP in a population of Staphylococcus aureus strains isolated from cows with mastitis. J Clin Microbiol 40:4060–4067
Gopal S, Divya KC (2017) Can methicillin-resistant Staphylococcus aureus prevalence from dairy cows in India act as potential risk for community-associated infections?: a review. Vet World 10:311–318
Harmsen D, Claus H, Witte W, Rothganger J, Claus H, Turnwald D, Vogel U (2003) Typing of methicillin-resistant Staphylococcus aureus in a university hospital setting by using novel software for spa repeat determination and database management. J Clin Microbiol 41:5442–5448
Havaei SA, Azimian A, Fazeli H, Naderi M, Ghazvini K, Samiee SM, Soleimani M (2013) Isolation of Asian endemic and livestock associated clones of methicillin resistant Staphylococcus aureus from ocular samples in northeastern Iran. Iran J Microbiol 5:227–232
Islam MZ, Espinosa-Gongora C, Damborg P, Sieber RN, Munk R, Husted L, Moodley A, Skov R, Larsen J, Guardabassi L (2017) Horses in Denmark are a reservoir of diverse clones of methicillin-resistant and -susceptible Staphylococcus aureus. Front Microbiol 8:543
Japoni-Nejad A, Rezazadeh M, Kazemian H, Fardmousavi N, van Belkum A, Ghaznavi-Rad E (2013) Molecular characterization of the first community-acquired methicillin-resistant Staphylococcus aureus strains from Central Iran. Int J Infect Dis 17:e949–e954
Johler S, Layer F, Stephan R (2011) Comparison of virulence and antibiotic resistance genes of food poisoning outbreak isolates of Staphylococcus aureus with isolates obtained from bovine mastitis milk and pig carcasses. J Food Prot 74:1852–1859
Karkaba A, Benschop J, Hill KE, Grinberg A (2016) Characterisation of methicillin-resistant Staphylococcus aureus clinical isolates from animals in New Zealand, 2012-2013, and subclinical colonisation in dogs and cats in Auckland. N Z Vet J:1–6
Katayama Y, Robinson DA, Enright MC, Chambers HF (2005) Genetic background affects stability of mecA in Staphylococcus aureus. J Clin Microbiol 43:2380–2383
Kiser KB, Cantey-Kiser JM, Lee JC (1999) Development and characterization of a Staphylococcus aureus nasal colonization model in mice. Infect Immun 67:5001–5006
Koreen L, Ramaswamy SV, Graviss EA, Naidich S, Musser JM, Kreiswirth BN (2004) spa typing method for discriminating among Staphylococcus aureus isolates: implications for use of a single marker to detect genetic micro- and macrovariation. J Clin Microbiol 42:792–799
Lakhundi S, Zhang K (2018) Methicillin-resistant Staphylococcus aureus: molecular characterization, evolution, and epidemiology. Clin Microbiol Rev 31:e00020–e00018
Lina G, Piemont Y, Godail-Gamot F, Bes M, Peter MO, Gauduchon V, Vandenesch F, Etienne J (1999) Involvement of Panton-valentine leukocidin-producing Staphylococcus aureus in primary skin infections and pneumonia. Clin Infect Dis 29:1128–1132
Lina G, Boutite F, Tristan A, Bes M, Etienne J, Vandenesch F (2003) Bacterial competition for human nasal cavity colonization: role of staphylococcal agr alleles. Appl Environ Microbiol 69:18–23
Liu J, Chen D, Peters BM, Li L, Li B, Xu Z, Shirliff ME (2016) Staphylococcal chromosomal cassettes mec (SCCmec): a mobile genetic element in methicillin-resistant Staphylococcus aureus. Microb Pathog 101:56–67
Loncaric I, Kunzel F, Licka T, Simhofer H, Spergser J, Rosengarten R (2014) Identification and characterization of methicillin-resistant Staphylococcus aureus (MRSA) from Austrian companion animals and horses. Vet Microbiol 168:381–387
Maddox TW, Clegg PD, Diggle PJ, Wedley AL, Dawson S, Pinchbeck GL, Williams NJ (2011) Cross-sectional study of antimicrobial-resistant bacteria in horses. Part 1: prevalence of antimicrobial-resistant Escherichia coli and methicillin-resistant Staphylococcus aureus. Equine Vet J 44:289–296
Manara S, Pasolli E, Dolce D, Ravenni N, Campana S, Armanini F, Asnicar F, Mengoni A, Galli L, Montagnani C, Venturini E, Rota-Stabelli O, Grandi G, Taccetti G, Segata N (2018) Whole-genome epidemiology, characterisation, and phylogenetic reconstruction of Staphylococcus aureus strains in a paediatric hospital. Genome Med 10:82
Mellmann A, Weniger T, Berssenbrugge C, Rothganger J, Sammeth M, Stoye J, Harmsen D (2007) Based upon repeat pattern (BURP): an algorithm to characterize the long-term evolution of Staphylococcus aureus populations based on spa polymorphisms. BMC Microbiol 7:98
Milheirico C, Oliveira DC, de Lencastre H (2007) Multiplex PCR strategy for subtyping the staphylococcal cassette chromosome mec type IV in methicillin-resistant Staphylococcus aureus: SCCmec IV multiplex. J Antimicrob Chemother 60 (1):42–48
Moridi M, Masoudi AA, Vaez Torshizi R, Hill EW (2013) Mitochondrial DNA D-loop sequence variation in maternal lineages of Iranian native horses. Anim Genet 44:209–213
Murakami K, Minamide W, Wada K, Nakamura E, Teraoka H, Watanabe S (1991) Identification of methicillin-resistant strains of staphylococci by polymerase chain reaction. J Clin Microbiol 29:2240–2244
Murugadas V, Toms CJ, Reethu SA, Lalitha KV (2017) Multilocus sequence typing and staphylococcal protein a typing revealed novel and diverse clones of methicillin-resistant Staphylococcus aureus in seafood and the aquatic environment. J Food Prot 80:476–481
Ohadian Moghadam S, Pourmand MR, Mahmoudi M, Sadighian H (2015) Molecular characterization of methicillin-resistant Staphylococcus aureus: characterization of major clones and emergence of epidemic clones of sequence type (ST) 36 and ST 121 in Tehran, Iran. FEMS Microbiol Lett 362:fnv043
O'Hara FP, Suaya JA, Ray GT, Baxter R, Brown ML, Mera RM, Close NM, Thomas E, Amrine-Madsen H (2016) Spa typing and multilocus sequence typing show comparable performance in a macroepidemiologic study of Staphylococcus aureus in the United States. Microb Drug Resist 22:88–96
Papadopoulos P, Papadopoulos T, Angelidis AS, Boukouvala E, Zdragas A, Papa A, Hadjichristodoulou C, Sergelidis D (2018) Prevalence of Staphylococcus aureus and of methicillin-resistant S. aureus (MRSA) along the production chain of dairy products in North-Western Greece. Food Microbiol 69:43–50
Peterson AE, Davis MF, Awantang G, Limbago B, Fosheim GE, Silbergeld EK (2012) Correlation between animal nasal carriage and environmental methicillin-resistant Staphylococcus aureus isolates at U.S. horse and cattle farms. Vet Microbiol 160:539–543
Sergelidis D, Papadopoulos T, Komodromos D, Sergelidou E, Lazou T, Papagianni M, Zdragas A, Papa A (2015) Isolation of methicillin-resistant Staphylococcus aureus from small ruminants and their meat at slaughter and retail level in Greece. Lett Appl Microbiol 61:498–503
Shambat S, Nadig S, Prabhakara S, Bes M, Etienne J, Arakere G (2012) Clonal complexes and virulence factors of Staphylococcus aureus from several cities in India. BMC Microbiol 12:64
Smith TC, Wardyn SE (2015) Human infections with Staphylococcus aureus CC398. Curr Environ Health Rep 2:41–51
Tokateloff N, Manning ST, Weese JS, Campbell J, Rothenburger J, Stephen C, Bastura V, Gow SP, Reid-Smith R (2009) Prevalence of methicillin-resistant Staphylococcus aureus colonization in horses in Saskatchewan, Alberta, and British Columbia. Can Vet J 50:1177–1180
Van den Eede A, Martens A, Feryn I, Vanderhaeghen W, Lipinska U, Gasthuys F, Butaye P, Haesebrouck F, Hermans K (2012) Low MRSA prevalence in horses at farm level. BMC Vet Res 8:213
Vandendriessche S, Vanderhaeghen W, Larsen J, de Mendonca R, Hallin M, Butaye P, Hermans K, Haesebrouck F, Denis O (2014) High genetic diversity of methicillin-susceptible Staphylococcus aureus (MSSA) from humans and animals on livestock farms and presence of SCCmec remnant DNA in MSSA CC398. J Antimicrob Chemother 69:355–362
Weese JS, van Duijkeren E (2010) Methicillin-resistant Staphylococcus aureus and Staphylococcus pseudintermedius in veterinary medicine. Vet Microbiol 140:418–429
Weese JS, Archambault M, Willey BM, Hearn P, Kreiswirth BN, Said-Salim B, McGeer A, Likhoshvay Y, Prescott JF, Low DE (2005) Methicillin-resistant Staphylococcus aureus in horses and horse personnel, 2000-2002. Emerg Infect Dis 11:430–435
Weese JS, Caldwell F, Willey BM, Kreiswirth BN, McGeer A, Rousseau J, Low DE (2006) An outbreak of methicillin-resistant Staphylococcus aureus skin infections resulting from horse to human transmission in a veterinary hospital. Vet Microbiol 114:160–164
Wichelhaus TA, Hunfeld KP, Boddinghaus B, Kraiczy P, Schafer V, Brade V (2001) Rapid molecular typing of methicillin-resistant Staphylococcus aureus by PCR-RFLP. Infect Control Hosp Epidemiol 22:294–298
Zunita Z, Bashir A, Hafizal A (2008) Occurrence of multidrug resistant Staphylococcus aureus in horses in Malaysia. Vet World 1:165–167
Acknowledgments
The authors are very grateful to Professor Alexander Mellmann from University of Münster, Institute of Hygiene, Germany, for conducting the BURP analysis. We are also thankful to Dr. M. Morovati, Dr. G. Jalilzadeh and Dr. S. Rostami for their help in sample collection, and Mrs. Mitra Panahi and Dr. S. Hosseinzadeh for technical assistance.
Funding
The Research Deputy of Urmia University financially supported the current investigation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The study protocol was approved by the Urmia University Animal Ethics Committee (ethical clearance number 1241).
Conflict of interest
The authors declare that they have no conflict of interest.
Research involving human participants and/or animals
Not applicable.
Informed consent
The authors confirm that this article’s content has no animal or human participants in research.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dastmalchi Saei, H., Safari, E. Methicillin resistance and clonal diversity of Staphylococcus aureus isolated from nasal samples of healthy horses in Iran. Ann Microbiol 69, 923–931 (2019). https://doi.org/10.1007/s13213-019-01484-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-019-01484-5