Abstract
Objectives
Colorectal cancer (CRC) is one of the most common solid tumors worldwide, but its diagnosis and treatment are limited. The objectives of our study were to compare the metabolic differences between CRC patients and healthy controls (HC), and to identify potential biomarkers in the serum that can be used for early diagnosis and as effective therapeutic targets. The aim was to provide a new direction for CRC predictive, preventive, and personalized medicine (PPPM).
Methods
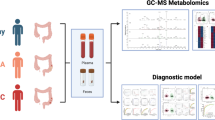
In this study, CRC patients (n = 30) and HC (n = 30) were recruited. Serum metabolites were assayed using an ultra-performance liquid chromatography coupled with quadrupole time-of-flight mass spectrometry (UPLC-Q-TOF/MS) technology. Subsequently, CRC cell lines (HCT116 and HCT8) were treated with metabolites to verify their function. Key targets were identified by molecular docking, thermal shift assay, and protein overexpression/inhibition experiments. The inhibitory effect of celastrol on tumor growth was also assessed, which included IC50 analysis, nude mice xenografting, molecular docking, protein overexpression/inhibition experiments, and network pharmacology technology.
Results
In the CRC group, 15 serum metabolites were significantly different in comparison with the HC group. The level of glycodeoxycholic acid (GDCA) was positively correlated with CRC and showed high sensitivity and specificity for the clinical diagnostic reference (AUC = 0.825). In vitro findings showed that GDCA promoted the proliferation and migration of CRC cell lines (HCT116 and HCT8), and Poly(ADP-ribose) polymerase-1 (PARP-1) was identified as one of the key targets of GDCA. The IC50 of celastrol in HCT116 cells was 121.1 nM, and the anticancer effect of celastrol was supported by in vivo experiments. Based on the potential of GDCA in PPPM, PARP-1 was found to be significantly correlated with the anticancer functions of celastrol.
Conclusion
These findings suggest that GDCA is an abnormally produced metabolite of CRC, which may provide an innovative molecular biomarker for the predictive identification and targeted prevention of CRC. In addition, PARP-1 was found to be an important target of GDCA that promotes CRC; therefore, celastrol may be a potential targeted therapy for CRC via its effects on PARP-1. Taken together, the pathophysiology and progress of tumor molecules mediated by changes in metabolite content provide a new perspective for predictive, preventive, and personalized medical of clinical cancer patients based on the target of metabolites in vivo.
Clinical trials registration number: ChiCTR2000039410.







Similar content being viewed by others
Data availability
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
All software applications used are included in this article.
Abbreviations
- PPPM:
-
Predictive, preventive, and personalized medicine
- CRC:
-
Colorectal cancer
- GDCA:
-
Glycodeoxycholic acid
- PARP-1:
-
Protein poly (ADP-ribose) polymerase-1
- PCA:
-
Principal component analysis
- OPLS-DA:
-
Orthogonal partial least squares discriminant analysis
- ROC:
-
Receiver operating characteristic
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- GO:
-
Gene Ontology
References
Edited by Wild CP, Weiderpass E, Stewart BW. World Cancer Report: Cancer Research for Cancer Prevention
Sharma R. An examination of colorectal cancer burden by socioeconomic status: evidence from GLOBOCAN 2018. EPMA J. 2019;11(1):95–117.
Akimoto N, Ugai T, Zhong R, Hamada T, Fujiyoshi K, Giannakis M, et al. Rising incidence of early-onset colorectal cancer-a call to action. Nat Rev Clin Oncol. 2021;18(4):230–43.
Reynolds A, Mann J, Cummings J, Winter N, Mete E, Te Morenga L. Carbohydrate quality and human health: a series of systematic reviews and meta-analyses. Lancet. 2019;393(10170):434–45.
Kerr J, Anderson C, Lippman SM. Physical activity, sedentary behaviour, diet, and cancer: an update and emerging new evidence. Lancet Oncol. 2017;18(8):e457–71.
Sveen A, Kopetz S, Lothe RA. Biomarker-guided therapy for colorectal cancer: strength in complexity. Nat Rev Clin Oncol. 2020;17(1):11–32.
Grech G, Zhan X, Yoo BC, Bubnov R, Hagan S, Danesi R, et al. EPMA position paper in cancer: current overview and future perspectives. EPMA J. 2015;6(1):9.
Wang J, Mouradov D, Wang X, Jorissen RN, Chambers MC, Zimmerman LJ, et al. Colorectal cancer cell line proteomes are representative of primary tumors and predict drug sensitivity. Gastroenterology. 2017;153(4):1082–95.
Aydin B, Caliskan A, Arga KY. Overview of omics biomarkers in pituitary neuroendocrine tumors to design future diagnosis and treatment strategies. EPMA J. 2021;12:383–401.
Lu M, Zhan X. The crucial role of multiomic approach in cancer research and clinically relevant outcomes. EPMA J. 2018;9(1):77–102.
Hull MA, Rees CJ, Sharp L, Koo S. A risk-stratified approach to colorectal cancer prevention and diagnosis. Nat Rev Gastroenterol Hepatol. 2020;17(12):773–80.
Simon K. Colorectal cancer development and advances in screening. Clin Interv Aging. 2016;11:967–76.
Levin B, Lieberman DA, McFarland B, Andrews KS, Brooks D, Bond J, et al. Screening and surveillance for the early detection of colorectal cancer and adenomatous polyps, 2008: a joint guideline from the American Cancer Society, the US Multi-Society Task Force on Colorectal Cancer, and the American College of Radiology. Gastroenterology. 2008;134(5):1570–95.
Needham BD, Adame MD, Serena G, Rose DR, Preston GM, Conrad MC, et al. Plasma and fecal metabolite profiles in autism spectrum disorder. Biol Psychiatry. 2021;89(5):451–62.
Danczak RE, Chu RK, Fansler SJ, Goldman AE, Graham EB, Tfaily MM, et al. Using metacommunity ecology to understand environmental metabolomes. Nat Commun. 2020;11(1):6369.
Ericksen RE, Lim SL, McDonnell E, Shuen WH, Vadiveloo M, White PJ, et al. Loss of BCAA catabolism during carcinogenesis enhances mTORC1 activity and promotes tumor development and progression. Cell Metab. 2019;29(5):1151-1165.e6.
Kaymak I, Williams KS, Cantor JR, Jones RG. Immunometabolic interplay in the tumor microenvironment. Cancer Cell. 2021;39(1):28–37.
Hamaidi I, Zhang L, Kim N, Wang MH, Iclozan C, Fang B, et al. Sirt2 inhibition enhances metabolic fitness and effector functions of tumor-reactive T cells. Cell Metab. 2020;32(3):420-436.e12.
Peitzsch C, Gorodetska I, Klusa D, Shi Q, Alves TC, Pantel K, et al. Metabolic regulation of prostate cancer heterogeneity and plasticity. Semin Cancer Biol. 2020;S1044–579X(20)30261–3.
Monedeiro F, Monedeiro-Milanowski M, Ligor T, Buszewski B. A review of GC-based analysis of non-invasive biomarkers of colorectal cancer and related pathways. J Clin Med. 2020;9(10):3191.
Yachida S, Mizutani S, Shiroma H, Shiba S, Nakajima T, Sakamoto T, et al. Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer. Nat Med. 2019;25(6):968–76.
Uchiyama K, Yagi N, Mizushima K, Higashimura Y, Hirai Y, Okayama T, et al. Serum metabolomics analysis for early detection of colorectal cancer. J Gastroenterol. 2017;52(6):677–94.
Wu JY, Huang TW, Hsieh YT, Wang YF, Yen CC, Lee GL, et al. Cancer-derived succinate promotes macrophage polarization and cancer metastasis via succinate receptor. Mol Cell. 2020;77(2):213-227.e5.
Koklesova L, Liskova A, Samec M, Qaradakhi T, Zulli A, Smejkal K, et al. Genoprotective activities of plant natural substances in cancer and chemopreventive strategies in the context of 3P medicine. EPMA J. 2020;11(2):261–87.
Dandawate PR, Subramaniam D, Jensen RA, Anant S. Targeting cancer stem cells and signaling pathways by phytochemicals: novel approach for breast cancer therapy. Semin Cancer Biol. 2016;40–41:192–208.
Samec M, Liskova A, Koklesova L, Samuel SM, Zhai K, Buhrmann C, et al. Flavonoids against the Warburg phenotype-concepts of predictive, preventive and personalised medicine to cut the Gordian knot of cancer cell metabolism. EPMA J. 2020;11(3):377–98.
Tong J, Shen Y, Zhang Z, Hu Y, Zhang X, Han L. Apigenin inhibits epithelial-mesenchymal transition of human colon cancer cells through NF-κB/Snail signaling pathway. Biosci Rep. 2019;39(5):BSR20190452.
Wang S, Ma K, Zhou C, Wang Y, Hu G, Chen L, et al. LKB1 and YAP phosphorylation play important roles in Celastrol-induced β-catenin degradation in colorectal cancer. Ther Adv Med Oncol. 2019;11:1758835919843736.
Zhu H, Liu XW, Ding WJ, Xu DQ, Zhao YC, Lu W, et al. Up-regulation of death receptor 4 and 5 by celastrol enhances the anti-cancer activity of TRAIL/Apo-2L. Cancer Lett. 2010;297(2):155–64.
Sun C, Li T, Song X, Huang L, Zang Q, Xu J, et al. Spatially resolved metabolomics to discover tumor-associated metabolic alterations. Proc Natl Acad Sci USA. 2019;116(1):52–7.
Chen PB, Black AS, Sobel AL, Zhao Y, Mukherjee P, Molparia B, et al. Directed remodeling of the mouse gut microbiome inhibits the development of atherosclerosis. Nat Biotechnol. 2020;38(11):1288–97.
Lavelle A, Sokol H. Gut microbiota-derived metabolites as key actors in inflammatory bowel disease. Nat Rev Gastroenterol Hepatol. 2020;17(4):223–37.
Funabashi M, Grove TL, Wang M, Varma Y, McFadden ME, Brown LC, et al. A metabolic pathway for bile acid dehydroxylation by the gut microbiome. Nature. 2020;582(7813):566–70.
Kühn T, Stepien M, López-Nogueroles M, Damms-Machado A, Sookthai D, Johnson T, et al. Prediagnostic plasma bile acid levels and colon cancer risk: a prospective study. J Natl Cancer Inst. 2020;112(5):516–24.
Nosho K, Yamamoto H, Mikami M, Taniguchi H, Takahashi T, Adachi Y, et al. Overexpression of poly(ADP-ribose) polymerase-1 (PARP-1) in the early stage of colorectal carcinogenesis. Eur J Cancer. 2006;42(14):2374–81.
Slyskova J, Korenkova V, Collins AR, Prochazka P, Vodickova L, Svec J, et al. Functional, genetic, and epigenetic aspects of base and nucleotide excision repair in colorectal carcinomas. Clin Cancer Res. 2012;18(21):5878–87.
Elmasry GF, Aly EE, Awadallah FM, El-Moghazy SM. Design and synthesis of novel PARP-1 inhibitors based on pyridopyridazinone scaffold. Bioorg Chem. 2019;87:655–66.
Kanth P, Inadomi JM. Screening and prevention of colorectal cancer. BMJ. 2021;374:n1855.
Keum N, Giovannucci E. Global burden of colorectal cancer: emerging trends, risk factors and prevention strategies. Nat Rev Gastroenterol Hepatol. 2019;16(12):713–32.
Menyhárt O, Győrffy B. Multi-omics approaches in cancer research with applications in tumor subtyping, prognosis, and diagnosis. Comput Struct Biotechnol J. 2021;19:949–60.
Liang L, Sun F, Wang H, Hu Z. Metabolomics, metabolic flux analysis and cancer pharmacology. Pharmacol Ther. 2021;224:107827.
Hao Y, Li D, Xu Y, Ouyang J, Wang Y, Zhang Y, et al. Investigation of lipid metabolism dysregulation and the effects on immune microenvironments in pan-cancer using multiple omics data. BMC Bioinformatics. 2019;20(Suppl 7):195.
Cheng C, Geng F, Cheng X, Guo D. Lipid metabolism reprogramming and its potential targets in cancer. Cancer Commun (Lond). 2018;38(1):27.
Snaebjornsson MT, Janaki-Raman S, Schulze A. Greasing the wheels of the cancer machine: the role of lipid metabolism in cancer. Cell Metab. 2020;31(1):62–76.
Costarelli V. Bile acids as possible human carcinogens: new tricks from an old dog. Int J Food Sci Nutr. 2009;60(Suppl 6):116–25.
Ocvirk S, O’Keefe SJ. Influence of bile acids on colorectal cancer risk: potential mechanisms mediated by diet - gut microbiota interactions. Curr Nutr Rep. 2017;6(4):315–22.
Schulthess J, Pandey S, Capitani M, Rue-Albrecht KC, Arnold I, Franchini F, et al. The short chain fatty acid butyrate imprints an antimicrobial program in macrophages. Immunity. 2019;50(2):432-445.e7.
Fan X, Jin Y, Chen G, Ma X, Zhang L. Gut microbiota dysbiosis drives the development of colorectal cancer. Digestion. 2021;102(4):508–15.
Yu J, Li S, Guo J, Xu Z, Zheng J, Sun X. Farnesoid X receptor antagonizes Wnt/β-catenin signaling in colorectal tumorigenesis. Cell Death Dis. 2020;11(8):640.
Gadaleta RM, Garcia-Irigoyen O, Moschetta A. Bile acids and colon cancer: is FXR the solution of the conundrum? Mol Aspects Med. 2017;56:66–74.
Mamianetti A, Garrido D, Carducci CN, Vescina MC. Fecal bile acid excretion profile in gallstone patients. Medicina (B Aires). 1999;59(3):269–73.
Cross AJ, Moore SC, Boca S, Huang WY, Xiong X, Stolzenberg-Solomon R, et al. A prospective study of serum metabolites and colorectal cancer risk. Cancer. 2014;120(19):3049–57.
Kam TI, Mao X, Park H, Chou SC, Karuppagounder SS, Umanah GE, et al. Poly(ADP-ribose) drives pathologic α-synuclein neurodegeneration in Parkinson’s disease. Science. 2018;362(6414):eaat8407.
Martín-Oliva D, O’Valle F, Muñoz-Gámez JA, Valenzuela MT, Nuñez MI, Aguilar M, et al. Crosstalk between PARP-1 and NF-kappaB modulates the promotion of skin neoplasia. Oncogene. 2004;23(31):5275–83.
Chan EM, Shibue T, McFarland JM, Gaeta B, Ghandi M, Dumont N, et al. WRN helicase is a synthetic lethal target in microsatellite unstable cancers. Nature. 2019;568(7753):551–6.
Ali T, Kim MO. Melatonin ameliorates amyloid beta-induced memory deficits, tau hyperphosphorylation and neurodegeneration via PI3/Akt/GSk3β pathway in the mouse hippocampus. J Pineal Res. 2015;59(1):47–59.
Pazzaglia S, Pioli C. Multifaceted role of PARP-1 in DNA repair and inflammation: pathological and therapeutic implications in cancer and non-cancer diseases. Cells. 2019;9(1):41.
Rojo F, García-Parra J, Zazo S, Tusquets I, Ferrer-Lozano J, Menendez S, et al. Nuclear PARP-1 protein overexpression is associated with poor overall survival in early breast cancer. Ann Oncol. 2012;23(5):1156–64.
Schiewer MJ, Goodwin JF, Han S, Brenner JC, Augello MA, Dean JL, et al. Dual roles of PARP-1 promote cancer growth and progression. Cancer Discov. 2012;2(12):1134–49.
Dörsam B, Seiwert N, Foersch S, Stroh S, Nagel G, Begaliew D, et al. PARP-1 protects against colorectal tumor induction, but promotes inflammation-driven colorectal tumor progression. Proc Natl Acad Sci U S A. 2018;115(17):E4061–70.
Pan SR, Chen ZY, Zhao K, Liu YC, Wang PY. Clinical research progress on disappearing colorectal liver metastases. Zhonghua Wei Chang Wai Ke Za Zhi. 2021;24(11):1028–34.
Ohmachi T, Inoue H, Mimori K, Tanaka F, Sasaki A, Kanda T, et al. Fatty acid binding protein 6 is overexpressed in colorectal cancer. Clin Cancer Res. 2006;12(17):5090–5.
Li H, Dai W, Xia X, Wang R, Zhao J, Han L, et al. Modeling tumor development and metastasis using paired organoids derived from patients with colorectal cancer liver metastases. J Hematol Oncol. 2020;13(1):119.
Rodríguez MI, Majuelos-Melguizo J, Martí Martín-Consuegra JM, Ruiz de Almodóvar M, López-Rivas A, Javier Oliver F. Deciphering the insights of poly(ADP-ribosylation) in tumor progression. Med Res Rev. 2015;35(4):678–697.
Rodríguez MI, González-Flores A, Dantzer F, Collard J, de Herreros AG, Oliver FJ. Poly(ADP-ribose)-dependent regulation of Snail1 protein stability. Oncogene. 2011;30(42):4365–72.
Rodríguez MI, Peralta-Leal A, O'Valle F, Rodriguez-Vargas JM, Gonzalez-Flores A, Majuelos-Melguizo J, et al. 2013 PARP-1 regulates metastatic melanoma through modulation of vimentin-induced malignant transformation PLoS Genet 9 (6) e1003531
Abdelrahman AE, Ibrahim DA, El-Azony A, Alnagar AA, Ibrahim A. ERCC1, PARP-1, and AQP1 as predictive biomarkers in colon cancer patients receiving adjuvant chemotherapy. Cancer Biomark. 2020;27(2):251–64.
Umar A, Dunn BK, Greenwald P. Future directions in cancer prevention. Nat Rev Cancer. 2012;12(12):835–48.
Amin AR, Kucuk O, Khuri FR, Shin DM. Perspectives for cancer prevention with natural compounds. J Clin Oncol. 2009;27(16):2712–25.
Huang XM, Yang ZJ, Xie Q, Zhang ZK, Zhang H, Ma JY 2019 Natural products for treating colorectal cancer: a mechanistic review Biomed Pharmacother 117 109142
Pang G, Wang F, Zhang LW. Dose matters: direct killing or immunoregulatory effects of natural polysaccharides in cancer treatment. Carbohydr Polym. 2018;195:243–56.
Huang M, Lu JJ, Huang MQ, Bao JL, Chen XP, Wang YT. Terpenoids: natural products for cancer therapy. Expert Opin Investig Drugs. 2012;21(12):1801–18.
Khan H, Ullah H, Castilho PCMF, Gomila AS, D’Onofrio G, Filosa R, et al. Targeting NF-κB signaling pathway in cancer by dietary polyphenols. Crit Rev Food Sci Nutr. 2020;60(16):2790–800.
Efferth T, Oesch F. Repurposing of plant alkaloids for cancer therapy: pharmacology and toxicology. Semin Cancer Biol. 2021;68:143–63.
Liu X, Zhao P, Wang X, Wang L, Zhu Y, Song Y, et al. Celastrol mediates autophagy and apoptosis via the ROS/JNK and Akt/mTOR signaling pathways in glioma cells. J Exp Clin Cancer Res. 2019;38(1):184.
Huang YL, Zhou YX, Zhou D, Zuo YX, Jia YP. Celastrol in the inhibition of neovascularization. Chinese Journal of Oncology. 2003;25(5):429–32.
Xu ZY, Shi JF, Xian J, Zhang C, Zhang JM. Research progress on anti-tumor mechanism of celastrol alone and in combination. Chinese Traditional and Herbal Drugs. 2021;52(14):4372–5.
Zhang H, Li J, Li G, Wang S. Effects of celastrol on enhancing apoptosis of ovarian cancer cells via the downregulation of microRNA-21 and the suppression of the PI3K/Akt-NF-κB signaling pathway in an in vitro model of ovarian carcinoma. Mol Med Rep. 2016;14(6):5363–8.
Pang X, Yi Z, Zhang J, Lu B, Sung B, Qu W, et al. Celastrol suppresses angiogenesis-mediated tumor growth through inhibition of AKT/mammalian target of rapamycin pathway. Cancer Res. 2010;70(5):1951–9.
Lin L, Sun Y, Wang D, Zheng S, Zhang J, Zheng C. Celastrol ameliorates ulcerative colitis-related colorectal cancer in mice via suppressing inflammatory responses and epithelial-mesenchymal transition. Front Pharmacol. 2016;6:320.
Johnson CH, Dejea CM, Edler D, Hoang LT, Santidrian AF, Felding BH, et al. Metabolism links bacterial biofilms and colon carcinogenesis. Cell Metab. 2015;21(6):891–7.
Chen F, Dai X, Zhou CC, Li KX, Zhang YJ, Lou XY, et al. Integrated analysis of the faecal metagenome and serum metabolome reveals the role of gut microbiome-associated metabolites in the detection of colorectal cancer and adenoma. Gut. 2021; gutjnl-2020–323476.
Zhang L, Wei TT, Li Y, Li J, Fan Y, Huang FQ, et al. Functional metabolomics characterizes a key role for n-acetylneuraminic acid in coronary artery diseases. Circulation. 2018;137(13):1374–90.
Liu CX. Scientific understanding of toxicity and safety of Chinese medicines. Chinese Herbal Medicines. 2018;2(10):107.
Funding
This work was supported by the National Natural Science Foundation of China (81573825, 81902826, 81672781 and 81973356) and Tianjin Development Program for Innovation and Entrepreneurship (20190617), the Fundamental Research Funds for the Central Universities, Nankai University (3206054, 91923101, 63213082 and 92122017), and the National Key R&D Program of China (No. 2018YFC2002000).
National Natural Science Foundation of China,81573825,Yubo Li,81902826,Shuai Zhang,81672781,Shuai Zhang,81973356,Shuai Zhang,Tianjin Development Program for Innovation and Entrepreneurship,20190617,Yubo Li,Fundamental Research Funds for the Central Universities,3206054,Changliang Shan,91923101,Changliang Shan,63213082,Changliang Shan,92122017,Changliang Shan,National Key R&D Program of China,2018YFC2002000,Changliang Shan
Author information
Authors and Affiliations
Contributions
SZ and YL conceived the study and designed the experiments. YY and CY performed most experiments and analyzed the data. YY, CY, YW, SS and LL assisted with cells culture. GS and JH collected clinical samples. YY, SS, and CB assisted with UPLC-Q-TOF/MS analysis. YM and MS performed animal experiments. CS, SZ, and YL performed writing. The author(s) read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the Institutional Ethics Committee of the Second Hospital of Tianjin Medical University (Ethical approval number: KY2019K144) and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Consent to participate
Written informed consent was obtained from individual or guardian participants.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file 1 Figure 1:
The base peak intensity (BPI) chromatogram of serum in the QC sample. (ai 481 KB)
Supplementary file 2 Figure 2:
Effect of GDCA on the activity of FHC cell line. Cell viability was measured with MTT assay in FHC cells. (ai 175 KB)
Supplementary file 3 Figure 3:
Anticancer effect of celastrol on CRC. A: The docking results of PARP-1 protein (PDB:5DS3) with celastro (2D). B: Weight changes of xenograft mice. (ai 200 KB)
Supplementary file 4
(XLSX 18.8 KB)
Supplementary file 5
(XLSX 34.4 KB)
Supplementary file 6
(XLSX 11.7 KB)
Supplementary file 7
(XLSX 12.8 KB)
Rights and permissions
About this article
Cite this article
Yuan, Y., Yang, C., Wang, Y. et al. Functional metabolome profiling may improve individual outcomes in colorectal cancer management implementing concepts of predictive, preventive, and personalized medical approach. EPMA Journal 13, 39–55 (2022). https://doi.org/10.1007/s13167-021-00269-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13167-021-00269-8