Abstract
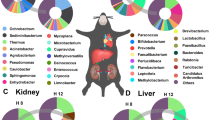
Death does not occur instantaneously and organs do not decompose at the same rate or in the same way. Nulligravid human uteri and prostate glands are the last internal organs to deteriorate during decomposition; however, the reason for this very important observation is still enigmatic. Recent studies have elucidated that the composition and abundance of microbes in the human thanatomicrobiome (microbiome of death) varies by organ and changes as a function of time and temperature. The ileocecal area has the largest absolute postmortem burden that spreads to the liver and spleen and continues to the heart and brain depending on the cause of death. To truly understand the mechanisms of microbial assembly during decomposition, a thorough examination of different strategies utilized by the trillions of microbes that colonize decaying tissues is needed from a multi-organ and multidisciplinary approach. In this review, we highlight interdisciplinary research and provide an overview of human decomposition investigations of thanatomicrobiomic changes in internal organs.


Similar content being viewed by others
References
Ford WW. On the bacteriology of normal organs. Epidemiol Infect. 1901;1:277–84.
Stewart EJ. Growing unculturable bacteria. J Bacteriol. 2012;194:4151–60.
Gevers W. Biochemical aspects of cell death. Forensic Sci. 1975;6:25–9.
Can I, Javan GT, Pozhitkov AE, Noble PA. Distinctive thanatomicrobiome signatures found in the blood and internal organs of humans. J Microbiol Methods. 2014;106:1–7.
Javan GT, Finley SJ, Can I, Wilkinson JE, Hanson JD, Tarone AM. Human thanatomicrobiome succession and time since death. Sci Rep. 2016;6:29598.
Javan GT, Finley SJ, Abidin Z, Mulle JG. The thanatomicrobiome: a missing piece of the microbial puzzle of death. Front Microbiol. 2016;7:225.
Adserias Garriga J, Quijada NM, Hernandez M, Lázaro DR, Steadman D, Garcia-Gil J. Dynamics of the oral microbiota as a tool to estimate time since death. Mol Oral Microbiol. 2017;32:511–6.
DeBruyn JM, Hauther KA. Postmortem succession of gut microbial communities in deceased human subjects. PeerJ. 2017;5:e3437.
Vass AA. Beyond the grave-understanding human decomposition. Microbiol Today. 2001;28:190–3.
Stefanuto PH, Perrault KA, Grabherr S, Varlet V, Focant JF. Postmortem internal gas reservoir monitoring using GC× GC-HRTOF-MS. Separations. 2016;3:24.
Carter DO, Tomberlin JK, Benbow ME, Metcalf JL. Perspectives on the future of forensic microbiology. In: Carter DO, Tomberlin JK, Benbow ME, Metcalf JL, editors. Forensic microbiology. New York City: Wiley; 2017. p. 376–8.
Pascual J, von Hoermann C, Rottler-Hoermann AM, et al. Function of bacterial community dynamics in the formation of cadaveric semiochemicals during in situ carcass decomposition. Environ Microbiol. 2017;9:3310–22.
Javan GT, Finley SJ, Smith T, Miller J, Wilkinson JE. Cadaver thanatomicrobiome signatures: the ubiquitous nature of Clostridium species in human decomposition. Front Microbiol. 2017;8:2096.
Bell CR, Wilkinson JE, Robertson BK, Javan GT. Sex-related differences in the thanatomicrobiome in postmortem heart samples using bacterial gene regions V1-2 and V4. J Appl Microbiol. 2018;67:144–53.
Pechal JL, Schmidt CJ, Jordan HR, Benbow ME. A large-scale survey of the postmortem human microbiome, and its potential to provide insight into the living health condition. Sci Rep. 2018;8:5724.
Tuomisto S, Pessi T, Collin P, Vuento R, Aittoniemi J, Karhunen PJ. Changes in gut bacterial populations and their translocation into liver and ascites in alcoholic liver cirrhotics. BMC Gastroenterol. 2014;14:40.
Belk A, Xu ZZ, Carter DO, Lynne A, Bucheli S, Knight R, et al. Microbiome data accurately predicts the postmortem interval using random forest regression models. Genes. 2018;9:104.
Zhou C, Byard RW. Factors and processes causing accelerated decomposition in human cadavers–an overview. J Forensic Legal Med. 2011;18:6–9.
Ellis KJ. Human body composition: in vivo methods. Physiol Rev. 2000;80:649–80.
Matuszewski S, Konwerski S, Frątczak K, Szafałowicz M. Effect of body mass and clothing on decomposition of pig carcasses. Int J Legal Med. 2014;128:1039–48.
Vass AA, Barshick SA, Sega G, et al. Decomposition chemistry of human remains: a new methodology for determining the postmortem interval. J For Sci. 2002;47:542–53.
Olakanye AO, Nelson A, Ralebitso-Senior TK. A comparative in situ decomposition study using still born piglets and leaf litter from a deciduous forest. Forensic Sci Int. 2017;276:85–92.
Dent BB, Forbes SL, Stuart BH. Review of human decomposition processes in soil. Environ Geol. 2004;45:576–85.
Goff ML. Early post-mortem changes and stages of decomposition in exposed cadavers. Exp Appl Acarol. 2009;49:21–36.
Clemente JC, Ursell LK, Parfrey LW, Knight R. The impact of the gut microbiota on human health: an integrative view. Cell. 2012;148:1258–70.
DeVault TL, Rhodes OE Jr, Shivik JA. Scavenging by vertebrates: behavioral, ecological, and evolutionary perspectives on an important energy transfer pathway in terrestrial ecosystems. Oikos. 2003;102:225–34.
Wilson EE, Wolkovich EM. Scavenging: how carnivores and carrion structure communities. Trends Ecol Evol. 2011;26:129–35.
Benbow ME, Lewis AJ, Tomberlin JK, Pechal JL. Seasonal necrophagous insect community assembly during vertebrate carrion decomposition. J Med Entomol. 2013;50:440–50.
Damann FE, Williams DE, Layton AC. Potential use of bacterial community succession in decaying human bone for estimating postmortem interval. J Forensic Sci. 2015;60:844–50.
Molina DK, DiMaio VJ. Normal organ weights in men: part II—the brain, lungs, liver, spleen, and kidneys. Am J Forensic Med Pathol. 2012;33:368–72.
Gunn A, Pitt SJ. Microbes as forensic indicators. Trop Biomed. 2012;29:311–30.
Huang C, Sloan EA, Boerkoel CF. Chromatin remodeling and human disease. Curr Opin Genet Dev. 2003;13:246–52.
Bär W, Kratzer A, Mächler M, Schmid W. Postmortem stability of DNA. Forensic Sci Int. 1988;39:59–70.
Hynd MR, Lewohl JM, Scott HL, Dodd PR. Biochemical and molecular studies using human autopsy brain tissue. J Neurochem. 2003;85:543–62.
Williams T, Soni S, White J, Can G, Javan GT. Evaluation of DNA degradation using flow cytometry: promising tool for postmortem interval determination. Am J Forensic Med Pathol. 2015;36:104–10.
Di Nunno NR, Costantinides F, Bernasconi P, Bottin C, Melato M. Is flow cytometric evaluation of DNA degradation a reliable method to investigate the early postmortem period? Am J Forensic Med Pathol. 1998;19:50–3.
Molina DK, DiMaio VJ. Normal organ weights in women: part I-the heart. Am J Forensic Med Pathol. 2015;36:176–81.
Payne-James J, Simpson K. Simpson’s forensic medicine. Boca Raton: CRC Press; 2011.
Levy AD, Harcke HT Jr. Essentials of forensic imaging: a text-atlas. Boca Raton: CRC Press; 2010.
Trotter SA, Brill Ii LB, Bennett JP. Stability of gene expression in postmortem brain revealed by cDNA gene array analysis. Brain Res. 2002;942:120–3.
Yasojima K, McGeer EG, McGeer PL. High stability of mRNAs postmortem and protocols for their assessment by RT-PCR. Brain Res Protocol. 2001;8:212–8.
Donaldson AE, Lamont IL. Biochemistry changes that occur after death: potential markers for determining post-mortem interval. PLoS One. 2013;8:e82011.
Cocariu EA, Mageriu V, Stăniceanu F, Bastian A, Socoliuc C, Zurac S. Correlations between the autolytic changes and postmortem interval in refrigerated cadavers. Rom J Intern Med. 2016;54:105–12.
Sibulesky L. Normal liver anatomy. Clin Liver Dis. 2013;2(S1).
Tuomisto S, Karhunen PJ, Vuento R, Aittoniemi J, Pessi T. Evaluation of postmortem bacterial migration using culturing and real-time quantitative PCR. J Forensic Sci. 2013;58:910–6.
Carpenter HM, Wilkins RM. Autopsy bacteriology: review of 2,033 cases. Arch Pathol. 1964;77:73–81.
Wilson SJ, Wilson ML, Reller LB. Diagnostic utility of postmortem blood cultures. Arch Pathol Lab Med. 1993;117:986–8.
Boutin EL, Battle E, Cunha GR. The response of female urogenital tract epithelia to mesenchymal inductors is restricted by the germ layer origin of the epithelium: prostatic inductions. Differentiation. 1991;48:99–105.
Chang K, Zhang L. Steroid hormones and uterine vascular adaptation to pregnancy. Reprod Sci. 2008;15:336–48.
Casper JL. A handbook of the practice of forensic medicine: based upon personal experience. London: New Sydenham Society; 1861.
Tolbert M, Finley SJ, Visonà SD, Soni S, Osculati A, Javan GT. The thanatotranscriptome: gene expression of male reproductive organs after death. Gene. 2018;675:191–6.
Lee CH, Akin-Olugbade O, Kirschenbaum A. Overview of prostate anatomy, histology, and pathology. Endocrinol Metab Clin N Am. 2011;40:565–75.
Suzuki Y, Tsujimoto Y, Matsui H, Watanabe K. Decomposition of extremely hard-to-degrade animal proteins by thermophilic bacteria. J Biosci Bioeng. 2006;102:73–81.
Hyde ER, Haarmann DP, Lynne AM, Bucheli SR, Petrosino JF. The living dead: bacterial community structure of a cadaver at the onset and end of the bloat stage of decomposition. PLoS One. 2013;8:e77733.
Hyde E, Haarmann DP, Petrosino JF, Lynne AM, Bucheli SR. Initial insights into bacterial succession during human decomposition. Int J Legal Med. 2015;129:661–71.
Metcalf JL, Parfrey LW, Gonzalez A, et al. A microbial clock provides an accurate estimate of the postmortem interval in a mouse model system. elife. 2013;2:e01104.
Thomas TB, Finley SJ, Wilkinson JE, Wescott DJ, Gorski A, Javan GT. Postmortem microbial communities in burial soil layers of skeletonized humans. J Forensic Legal Med. 2017;49:43–9.
Pechal JL, Crippen TL, Tarone AM. Microbial community functional change during vertebrate carrion decomposition. PLoS One. 2013;8:e79035.
Pechal JL, Crippen TL, Benbow ME, Tarone AM, Dowd S, Tomberlin JK. The potential use of bacterial community succession in forensics as described by high throughput metagenomic sequencing. Int J Legal Med. 2014;128:193–205.
Benbow ME, Pechal JL, Lang JM, Erb R, Wallace JR. The potential of high-throughput metagenomic sequencing of aquatic bacterial communities to estimate the postmortem submersion interval. J Forensic Sci. 2015;60:1500–10.
Fredette JW. Bacteremias in the agonal period. Transl Res. 1916;2:180–8.
Fouts DE, Torralba M, Nelson KE, Brenner DA, Schnabl B. Bacterial translocation and changes in the intestinal microbiome in mouse models of liver disease. J Hepatol. 2012;56:1283–92.
Heimesaat MM, Boelke S, Fischer A, et al. Comprehensive postmortem analyses of intestinal microbiota changes and bacterial translocation in human flora associated mice. PLoS One. 2012;7:e40758.
Burcham ZM, Hood JA, Pechal JL, et al. Fluorescently labeled bacteria provide insight on post-mortem microbial transmigration. Forensic Sci Int. 2016;264:63–9.
Dix J, Graham M. Time of death, decomposition and identification: an atlas. Boca Raton: CRC press; 1999.
Latil M, Rocheteau P, Châtre L, et al. Skeletal muscle stem cells adopt a dormant cell state post mortem and retain regenerative capacity. Nat Commun. 2012;3:903.
Mostafa EM, El-Elemi AH, El-Beblawy MA, Dawood AE. Adult sex identification using digital radiographs of the proximal epiphysis of the femur at Suez Canal University Hospital in Ismailia, Egypt. Egypt J Forensic Sci. 2012;2:81–8.
Đurić M, Rakočević Z, Đonić D. The reliability of sex determination of skeletons from forensic context in the Balkans. Forensic Sci Int. 2005;147:159–64.
Campobasso CP, Introna F. The forensic entomologist in the context of the forensic pathologist’s role. Forensic Sci Int. 2001;120:132–9.
Acknowledgements
This review paper was supported by National Science Foundation HRD 1401075 and National Institute of Justice 2017-MU-MU-0042 grants.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Human and animal studies
The authors declare that no research involving human participants and/or animals was conducted in this review.
No identifying information about participants was used in this review.
Rights and permissions
About this article
Cite this article
Javan, G.T., Finley, S.J., Tuomisto, S. et al. An interdisciplinary review of the thanatomicrobiome in human decomposition. Forensic Sci Med Pathol 15, 75–83 (2019). https://doi.org/10.1007/s12024-018-0061-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12024-018-0061-0