Abstract
Introduction
Autism spectrum disorder (ASD) is a group of neurodevelopmental disorders characterized by deficiencies in social interactions and communication, combined with restricted and repetitive behavioral issues.
Objectives
As little is known about the etiopathophysiology of ASD and early diagnosis is relatively subjective, we aim to employ a targeted, fully quantitative metabolomics approach to biochemically profile post-mortem human brain with the overall goal of identifying metabolic pathways that may have been perturbed as a result of the disease while uncovering potential central diagnostic biomarkers.
Methods
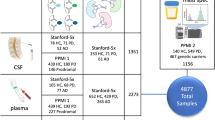
Using a combination of 1H NMR and DI/LC–MS/MS we quantitatively profiled the metabolome of the posterolateral cerebellum from post-mortem human brain harvested from people who suffered with ASD (n = 11) and compared them with age-matched controls (n = 10).
Results
We accurately identified and quantified 203 metabolites in post-mortem brain extracts and performed a metabolite set enrichment analyses identifying 3 metabolic pathways as significantly perturbed (p < 0.05). These include Pyrimidine, Ubiquinone and Vitamin K metabolism. Further, using a variety of machine-based learning algorithms, we identified a panel of central biomarkers (9-hexadecenoylcarnitine (C16:1) and the phosphatidylcholine PC ae C36:1) capable of discriminating between ASD and controls with an AUC = 0.855 with a sensitivity and specificity equal to 0.80 and 0.818, respectively.
Conclusion
For the first time, we report the use of a multi-platform metabolomics approach to biochemically profile brain from people with ASD and report several metabolic pathways which are perturbed in the diseased brain of ASD sufferers. Further, we identified a panel of biomarkers capable of distinguishing ASD from control brains. We believe that these central biomarkers may be useful for diagnosing ASD in more accessible biomatrices.




Similar content being viewed by others
Data availability
The 1H NMR spectroscopy and mass spectrometry metabolomics data have been deposited to the MetaboLights. Archive (https://www.ebi.ac.uk/metabolights/mysubmissions?status=PRIVATE) via the MetaboLights partner repository with the data set MTBL960.
References
Altman, D. G., & Bland, J. M. (1995). Absence of evidence is not evidence of absence. BMJ,311, 485.
Bahado-Singh, R., Poon, L. C., Yilmaz, A., Syngelaki, A., Turkoglu, O., Kumar, P., et al. (2017a). Integrated proteomic and metabolomic prediction of term preeclampsia. Scientific Reports,7, 16189.
Bahado-Singh, R. O., Graham, S. F., Han, B., Turkoglu, O., Ziadeh, J., Mandal, R., et al. (2016). Serum metabolomic markers for traumatic brain injury: A mouse model. Metabolomics,12, 100.
Bahado-Singh, R. O., Syngelaki, A., Mandal, R., Graham, S. F., Akolekar, R., Han, B., et al. (2017b). Metabolomic determination of pathogenesis of late-onset preeclampsia. The Journal of Maternal-Fetal & Neonatal Medicine,30, 658–664.
Baio, J., Wiggins, L., Christensen, D. L., Maenner, M. J., Daniels, J., Warren, Z., et al. (2014). Prevalence of autism spectrum disorder among children aged 8 years—autism and developmental disabilities monitoring network, 11 sites, United States. MMWR Surveillance Summaries, 67(6), 1–23. https://doi.org/10.15585/mmwr.ss6706a1.
Blatt, G. J. (2012). The neuropathology of autism. Scientifica (Cairo),2012, 703675.
Brown, C. E. (1981). Interactions among carnosine, anserine, ophidine and copper in biochemical adaptation. Journal of Theoretical Biology,88, 245–256.
Chauhan, A., & Chauhan, V. (2006). Oxidative stress in autism. Pathophysiology,13, 171–181.
Chong, J., Soufan, O., Li, C., Caraus, I., Li, S., Bourque, G., et al. (2018). MetaboAnalyst 4.0: Towards more transparent and integrative metabolomics analysis. Nucleic Acids Research,46, W486–W494.
Coleman, M., & Blass, J. P. (1985). Autism and lactic acidosis. Journal of Autism and Developmental Disorders,15, 1–8.
Deth, R., Muratore, C., Benzecry, J., Power-Charnitsky, V. A., & Waly, M. (2008). How environmental and genetic factors combine to cause autism: A redox/methylation hypothesis. Neurotoxicology,29, 190–201.
Diémé, B., Mavel, S., Blasco, H., Tripi, G., Bonnet-Brilhault, F., Malvy, J., et al. (2015). Metabolomics study of urine in autism spectrum disorders using a multiplatform analytical methodology. Journal of Proteome Research,14, 5273–5282.
Dolske, M. C., Spollen, J., Mckay, S., Lancashire, E., & Tolbert, L. (1993). A preliminary trial of ascorbic acid as supplemental therapy for autism. Progress in Neuro-Psychopharmacology and Biological Psychiatry,17, 765–774.
Fatemi, S. H., Halt, A. R., Realmuto, G., Earle, J., Kist, D. A., Thuras, P., et al. (2002). Purkinje cell size is reduced in cerebellum of patients with autism. Cellular and Molecular Neurobiology,22, 171–175.
Fiandaca, M. S., Zhong, X., Cheema, A. K., Orquiza, M. H., Chidambaram, S., Tan, M. T., et al. (2015). Plasma 24-metabolite panel predicts preclinical transition to clinical stages of alzheimer's disease. Frontiers in Neurology,6, 237.
Frye, R. E. (2012). Biomarker of abnormal energy metabolism in children with autism spectrum disorder. North American Journal of Medicine & Science,5, 141–147.
Frye, R. E., Melnyk, S., & Macfabe, D. F. (2013). Unique acyl-carnitine profiles are potential biomarkers for acquired mitochondrial disease in autism spectrum disorder. Translational Psychiatry,3, e220.
Fumagalli, M., Lecca, D., Abbracchio, M. P., & Ceruti, S. (2017). Pathophysiological role of purines and pyrimidines in neurodevelopment: Unveiling new pharmacological approaches to congenital brain diseases. Frontiers in Pharmacology,8, 941–941.
Gevi, F., Zolla, L., Gabriele, S., & Persico, A. M. (2016). Urinary metabolomics of young Italian autistic children supports abnormal tryptophan and purine metabolism. Molecular Autism,7, 47–47.
Goldani, A. A. S., Downs, S. R., Widjaja, F., Lawton, B., & Hendren, R. L. (2014). Biomarkers in autism. Frontiers in Psychiatry,5, 100.
Golnik, A. E., & Ireland, M. (2009). Complementary alternative medicine for children with autism: A physician survey. Journal of Autism and Developmental Disorders,39, 996–1005.
Graham, S. F., Chevallier, O. P., Kumar, P., Turkoglu, O., & Bahado-Singh, R. O. (2017a). Metabolomic profiling of brain from infants who died from sudden infant death syndrome reveals novel predictive biomarkers. Journal of Perinatology,37, 91–97.
Graham, S. F., Chevallier, O. P., Kumar, P., Türkoğlu, O., & Bahado-Singh, R. O. (2016a). High resolution metabolomic analysis of ASD human brain uncovers novel biomarkers of disease. Metabolomics,12, 62.
Graham, S. F., Chevallier, O. P., Roberts, D., Holscher, C., Elliott, C. T., & Green, B. D. (2013). Investigation of the human brain metabolome to identify potential markers for early diagnosis and therapeutic targets of Alzheimer's disease. Analytical Chemistry,85, 1803–1811.
Graham, S. F., Kumar, P. K., Bjorndahl, T., Han, B., Yilmaz, A., Sherman, E., et al. (2016b). Metabolic signatures of Huntington's disease (HD): (1)H NMR analysis of the polar metabolome in post-mortem human brain. Biochimica et Biophysica Acta,1862, 1675–1684.
Graham, S. F., Pan, X., Yilmaz, A., Macias, S., Robinson, A., Mann, D., et al. (2018a). Targeted biochemical profiling of brain from Huntington's disease patients reveals novel metabolic pathways of interest. Biochimica et Biophysica Acta,1864, 2430–2437.
Graham, S. F., Rey, N. L., Yilmaz, A., Kumar, P., Madaj, Z., Maddens, M., et al. (2018b). Biochemical profiling of the brain and blood metabolome in a mouse model of prodromal Parkinson's disease reveal distinct metabolic profiles. Journal of Proteome Research,17, 2460.
Graham, S. F., Turkoglu, O., Kumar, P., Yilmaz, A., Bjorndahl, T. C., Han, B., et al. (2017b). Targeted metabolic profiling of post-mortem brain from infants who died from sudden infant death syndrome. Journal of Proteome Research,16, 2587–2596.
Grunewald, R. A. (1993). Ascorbic acid in the brain. Brain Research, Brain Research Reviews,18, 123–133.
Harrison, F. E., & May, J. M. (2009). Vitamin C function in the brain: Vital role of the ascorbate transporter (SVCT2). Free Radical Biology & Medicine,46, 719–730.
Kemper, T. L., & Bauman, M. L. (1993). The contribution of neuropathologic studies to the understanding of autism. Neurologic Clinics,11, 175–187.
Kern, J. K., & Jones, A. M. (2006). Evidence of toxicity, oxidative stress, and neuronal insult in autism. Journal of Toxicology and Environmental Health, Part B: Critical Reviews,9, 485–499.
Klein, J. A., & Ackerman, S. L. (2003). Oxidative stress, cell cycle, and neurodegeneration. Journal of Clinical Investigation,111, 785–793.
Kohen, R., Yamamoto, Y., Cundy, K. C., & Ames, B. N. (1988). Antioxidant activity of carnosine, homocarnosine, and anserine present in muscle and brain. Proceedings of the National Academy of Sciences of the United States of America,85, 3175–3179.
Kurochkin, I., Khrameeva, E., Tkachev, A., Stepanova, V., Vanyushkina, A., Stekolshchikova, E., et al. (2019). Metabolome signature of autism in the human prefrontal cortex. Communications Biology,2, 234.
Lecca, D., & Ceruti, S. (2008). Uracil nucleotides: From metabolic intermediates to neuroprotection and neuroinflammation. Biochemical Pharmacology,75, 1869–1881.
Liu, A., Zhou, W., Qu, L., He, F., Wang, H., Wang, Y., et al. (2019). Altered urinary amino acids in children with autism spectrum disorders. Frontiers in Cellular Neuroscience,13, 7–7.
Loffler, M., Carrey, E. A., & Zameitat, E. (2015). Orotic acid, more than just an intermediate of pyrimidine de novo synthesis. Journal of Genetics and Genomics,42, 207–219.
Lombard, J. (1998). Autism: A mitochondrial disorder? Medical Hypotheses,50, 497–500.
Macfabe, D. F. (2015). Enteric short-chain fatty acids: Microbial messengers of metabolism, mitochondria, and mind: Implications in autism spectrum disorders. Microbial Ecology in Health and Disease,26, 28177.
Mapstone, M., Cheema, A. K., Fiandaca, M. S., Zhong, X., Mhyre, T. R., Macarthur, L. H., et al. (2014). Plasma phospholipids identify antecedent memory impairment in older adults. Nature Medicine,20, 415–418.
Ming, X., Stein, T. P., Barnes, V., Rhodes, N., & Guo, L. (2012). Metabolic perturbance in autism spectrum disorders: A metabolomics study. Journal of Proteome Research,11, 5856–5862.
Pan, X., Nasaruddin, M. B., Elliott, C. T., Mcguinness, B., Passmore, A. P., Kehoe, P. G., et al. (2016a). Alzheimer's disease-like pathology has transient effects on the brain and blood metabolome. Neurobiology of Aging,38, 151–163.
Ratajczak, H. V. (2011). Theoretical aspects of autism: Biomarkers—A review. J Immunotoxicol,8, 80–94.
Ravanbakhsh, S., Liu, P., Bjordahl, T. C., Mandal, R., Grant, J. R., Wilson, M., et al. (2015). Accurate, fully-automated NMR spectral profiling for metabolomics. PLoS ONE,10, e0124219.
Rebec, G. V., & Pierce, R. C. (1994). A vitamin as neuromodulator: Ascorbate release into the extracellular fluid of the brain regulates dopaminergic and glutamatergic transmission. Progress in Neurobiology,43, 537–565.
Ritvo, E. R., Freeman, B. J., Scheibel, A. B., Duong, T., Robinson, H., Guthrie, D., et al. (1986). Lower Purkinje cell counts in the cerebella of four autistic subjects: Initial findings of the UCLA-NSAC Autopsy Research Report. American Journal of Psychiatry,143, 862–866.
Rose, S., Niyazov, D. M., Rossignol, D. A., Goldenthal, M., Kahler, S. G., & Frye, R. E. (2018). Clinical and molecular characteristics of mitochondrial dysfunction in autism spectrum disorder. Molecular Diagnosis & Therapy,22, 571–593.
Rossignol, D. A., & Frye, R. E. (2012a). Mitochondrial dysfunction in autism spectrum disorders: A systematic review and meta-analysis. Molecular Psychiatry,17, 290–314.
Rotholz, D. A., Kinsman, A. M., Lacy, K. K., & Charles, J. (2017). Improving early identification and intervention for children at risk for autism spectrum disorder. Pediatrics,139(2), e20161061. https://doi.org/10.1542/peds.2016-1061.
Tibshirani, R. (1996). Regression shrinkage and selection via the Lasso. Journal of the Royal Statistical Society Series B (Methodological),58, 267–288.
Urban, M., Enot, D. P., Dallmann, G., Körner, L., Forcher, V., Enoh, P., et al. (2010). Complexity and pitfalls of mass spectrometry-based targeted metabolomics in brain research. Analytical Biochemistry,406, 124–131.
Vabalas, A., Gowen, E., Poliakoff, E., & Casson, A. J. (2019). Machine learning algorithm validation with a limited sample size. PLoS ONE,14, e0224365.
Varghese, M., Keshav, N., Jacot-Descombes, S., Warda, T., Wicinski, B., Dickstein, D. L., et al. (2017). Autism spectrum disorder: Neuropathology and animal models. Acta Neuropathologica,134, 537–566.
Varma, V. R., Oommen, A. M., Varma, S., Casanova, R., An, Y., Andrews, R. M., et al. (2018). Brain and blood metabolite signatures of pathology and progression in Alzheimer disease: A targeted metabolomics study. PLoS Medicine, 15(1), e1002482. https://doi.org/10.1371/journal.pmed.1002482.
Wang, H., Liang, S., Wang, M., Gao, J., Sun, C., Wang, J., et al. (2016). Potential serum biomarkers from a metabolomics study of autism. Journal of Psychiatry & Neuroscience:JPN,41, 27–37.
Wang, L., Angley, M. T., Gerber, J. P., & Sorich, M. J. (2011). A review of candidate urinary biomarkers for autism spectrum disorder. Biomarkers,16, 537–552.
West, P. R., Amaral, D. G., Bais, P., Smith, A. M., Egnash, L. A., Ross, M. E., et al. (2014). Metabolomics as a tool for discovery of biomarkers of autism spectrum disorder in the blood plasma of children. PLoS ONE,9, e112445–e112445.
Wishart, D. S. (2010). Computational approaches to metabolomics. Methods in Molecular Biology,593, 283–313.
Xia, J., Broadhurst, D. I., Wilson, M., & Wishart, D. S. (2013). Translational biomarker discovery in clinical metabolomics: An introductory tutorial. Metabolomics,9, 280–299.
Zeviani, M., Bertagnolio, B., & Uziel, G. (1996). Neurological presentations of mitochondrial diseases. Journal of Inherited Metabolic Disease,19, 504–520.
Acknowledgements
We would like to thank the NICHD Brain and Tissue Bank for Developmental Disorder ad NICH Contract #HHSN275200900011C, Ref. No. N01-HD-9-0011 for supplying the tissue used herein. In addition, this work was partly funded by the generous contribution made by the Fred A. & Barbara M. Erb Foundation.
Author information
Authors and Affiliations
Contributions
Designing research studies (SFG, ROB-S), conducting experiments (AY, TB, RM, ZU), statistical analysis (BH, IU), analyzing data (SFG, BH, IU, AY), and writing the manuscript (All authors contributed to the writing of the manuscript).
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Ethical approval
This study was performed in accordance with the 1964 Helsinki declaration and its later amendments, and ethical approval was obtained from the Beaumont Institutional Review Board (IRB# 2014-142).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11306_2020_1685_MOESM3_ESM.docx
Supplementary file3 (DOCX 15 kb) Table S1. Clinical and demographic information of patient and control groups. The donor highlighted in Bold was excluded from the study.
Rights and permissions
About this article
Cite this article
Graham, S.F., Turkoglu, O., Yilmaz, A. et al. Targeted metabolomics highlights perturbed metabolism in the brain of autism spectrum disorder sufferers. Metabolomics 16, 59 (2020). https://doi.org/10.1007/s11306-020-01685-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-020-01685-z