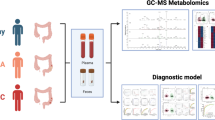
Abstract
Introduction
Epithelial ovarian cancer (EOC) remains the leading cause of death from gynecologic malignancies and has an alarming global fatality rate. Besides the differences in underlying pathogenesis, distinguishing between high grade (HG) and low grade (LG) EOC is imperative for the prediction of disease progression and responsiveness to chemotherapy.
Objectives
The aim of this study was to investigate, the tissue metabolome associated with HG and LG serous epithelial ovarian cancer.
Methods
A combination of one dimensional proton nuclear magnetic resonance (1D H NMR) spectroscopy and targeted mass spectrometry (MS) was employed to profile the tissue metabolome of HG, LG serous EOCs, and controls.
Results
Using partial least squares-discriminant analysis, we observed significant separation between all groups (p < 0.05) following cross validation. We identified which metabolites were significantly perturbed in each EOC grade as compared with controls and report the biochemical pathways which were perturbed due to the disease. Among these metabolic pathways, ascorbate and aldarate metabolism was identified, for the first time, as being significantly altered in both LG and HG serous cancers. Further, we have identified potential biomarkers of EOC and generated predictive algorithms with AUC (CI) = 0.940 and 0.929 for HG and LG, respectively.
Conclusion
These previously unreported biochemical changes provide a framework for future metabolomic studies for the development of EOC biomarkers. Finally, pharmacologic targeting of the key metabolic pathways identified herein could lead to novel and effective treatments of EOC.



Similar content being viewed by others
Data availability
The1H NMR spectroscopy and mass spectrometry metabolomics data have been deposited to the MetaboLights. Archive (https://www.ebi.ac.uk/metabolights/mysubmissions?status=PRIVATE) via the MetaboLights partner repository with the data set MTBLS640. Username: ali.yilmaz@beaumont.org and study ID is MTBLS640.
References
Amelio, I., Cutruzzolá, F., Antonov, A., Agostini, M., & Melino, G. (2014). Serine and glycine metabolism in cancer. Trends in Biochemical Sciences, 39(4), 191–198. https://doi.org/10.1016/j.tibs.2014.02.004.
Anastasi, E., Granato, T., Falzarano, R., Storelli, P., Ticino, A., Frati, L., et al. (2013). The use of HE4, CA125 and CA72-4 biomarkers for differential diagnosis between ovarian endometrioma and epithelial ovarian cancer. Journal of Ovarian Research, 6(1), 44–44. https://doi.org/10.1186/1757-2215-6-44.
Andrisic, L., Dudzik, D., Barbas, C., Milkovic, L., Grune, T., & Zarkovic, N. (2018). Short overview on metabolomics approach to study pathophysiology of oxidative stress in cancer. Redox Biology, 14, 47–58. https://doi.org/10.1016/j.redox.2017.08.009.
Atrih, A., Mudaliar, M. A. V., Zakikhani, P., Lamont, D. J., Huang, J. T. J., Bray, S. E., et al. (2014). Quantitative proteomics in resected renal cancer tissue for biomarker discovery and profiling. British Journal of Cancer, 110(6), 1622–1633. https://doi.org/10.1038/bjc.2014.24.
Bast, R. C., Hennessy, B., & Mills, G. B. (2009). The biology of ovarian cancer: New opportunities for translation. Nature Reviews Cancer, 9(6), 415–428.
Benjamin, D. I., Cravatt, B. F., & Nomura, D. K. (2012). Global profiling strategies for mapping dysregulated metabolic pathways in cancer. Cell Metabolism, 16(5), 565–577. https://doi.org/10.1016/j.cmet.2012.09.013.
Bosse, K., Rhiem, K., Wappenschmidt, B., Hellmich, M., Madeja, M., Ortmann, M., et al. (2006). Screening for ovarian cancer by transvaginal ultrasound and serum CA125 measurement in women with a familial predisposition: A prospective cohort study. Gynecologic Oncology, 103(3), 1077–1082. https://doi.org/10.1016/j.ygyno.2006.06.032.
Cho, K. R., & Shih, I.-M. (2009). Ovarian cancer. Annual Review of Pathology, 4, 287–313. https://doi.org/10.1146/annurev.pathol.4.110807.092246.
Das, P. M., & Bast, R. C. Jr. (2008). Early detection of ovarian cancer. Biomarkers in Medicine, 2(3), 291–303. https://doi.org/10.2217/17520363.2.3.291.
Dolce, V., Cappello, A. R., Lappano, R., & Maggiolini, M. (2011). Glycerophospholipid synthesis as a novel drug target against cancer. Current Molecular Pharmacology, 4(3), 167–175.
Elzek, M. A., & Rodland, K. D. (2015). Proteomics of ovarian cancer: Functional insights and clinical applications. Cancer and Metastasis Reviews, 34(1), 83–96. https://doi.org/10.1007/s10555-014-9547-8.
Fan, L., Zhang, W., Yin, M., Zhang, T., Wu, X., Zhang, H., et al. (2012). Identification of metabolic biomarkers to diagnose epithelial ovarian cancer using a UPLC/QTOF/MS platform. Acta Oncologica, 51(4), 473–479. https://doi.org/10.3109/0284186X.2011.648338.
Faul, F., Erdfelder, E., Buchner, A., & Lang, A.-G. (2009). Statistical power analyses using G*Power 3.1: Tests for correlation and regression analyses. Behavior Research Methods, 41(4), 1149–1160. https://doi.org/10.3758/brm.41.4.1149.
Flavin, R., Peluso, S., Nguyen, P. L., & Loda, M. (2010). Fatty acid synthase as a potential therapeutic target in cancer. Future Oncology (London, England), 6(4), 551–562. https://doi.org/10.2217/fon.10.11.
Gaul, D. A., Mezencev, R., Long, T. Q., Jones, C. M., Benigno, B. B., Gray, A., et al. (2015). Highly-accurate metabolomic detection of early-stage ovarian cancer. Scientific Reports, 5, 16351. https://doi.org/10.1038/srep16351.
Glunde, K., Ackerstaff, E., Mori, N., Jacobs, M. A., & Bhujwalla, Z. M. (2006). Choline phospholipid metabolism in cancer: Consequences for molecular pharmaceutical interventions. Molecular Pharmaceutics, 3(5), 496–506. https://doi.org/10.1021/mp060067e.
Gowda, G. A. N., Zhang, S., Gu, H., Asiago, V., Shanaiah, N., & Raftery, D. (2008). Metabolomics-based methods for early disease diagnostics. Expert Review of Molecular Diagnostics, 8(5), 617–633. https://doi.org/10.1586/14737159.8.5.617.
Graham, S. F., Kumar, P., Bahado-Singh, R. O., Robinson, A., Mann, D., & Green, B. D. (2016). Novel metabolite biomarkers of Huntington’s disease as detected by high-resolution mass spectrometry. Journal of Proteome Research, 15(5), 1592–1601. https://doi.org/10.1021/acs.jproteome.6b00049.
Hensley, K., & Denton, T. T. (2015). Alternative functions of the brain transsulfuration pathway represent an underappreciated aspect of brain redox biochemistry with significant potential for therapeutic engagement. Free Radical Biology & Medicine, 0, 123–134. https://doi.org/10.1016/j.freeradbiomed.2014.10.581.
Hishikawa, D., Hashidate, T., Shimizu, T., & Shindou, H. (2014). Diversity and function of membrane glycerophospholipids generated by the remodeling pathway in mammalian cells. Journal of Lipid Research, 55(5), 799–807. https://doi.org/10.1194/jlr.R046094.
Hoffmann, E. K., Lambert, I. H., & Pedersen, S. F. (2009). Physiology of cell volume regulation in vertebrates. Physiological Reviews, 89(1), 193–277. https://doi.org/10.1152/physrev.00037.2007.
Jacobs, I. J., & Menon, U. (2004). Progress and challenges in screening for early detection of ovarian cancer. Molecular & Cellular Proteomics, 3(4), 355–366. https://doi.org/10.1074/mcp.R400006-MCP200.
Jarboe, E., Folkins, A., Nucci, M. R., Kindelberger, D., Drapkin, R., Miron, A., et al. (2008). Serous carcinogenesis in the fallopian tube: A descriptive classification. International Journal of Gynecological Pathology, 27(1), 1–9. https://doi.org/10.1097/pgp.0b013e31814b191f.
Ke, C., Hou, Y., Zhang, H., Fan, L., Ge, T., Guo, B., et al. (2015). Large-scale profiling of metabolic dysregulation in ovarian cancer. International Journal of Cancer, 136(3), 516–526. https://doi.org/10.1002/ijc.29010.
Ke, C., Li, A., Hou, Y., Sun, M., Yang, K., Cheng, J., et al. (2016). Metabolic phenotyping for monitoring ovarian cancer patients. Scientific Reports, 6, 23334. https://doi.org/10.1038/srep23334.
Kim, A., Ueda, Y., Naka, T., & Enomoto, T. (2012). Therapeutic strategies in epithelial ovarian cancer. Journal of Experimental & Clinical Cancer Research, 31(1), 14. https://doi.org/10.1186/1756-9966-31-14.
Köbel, M., Kalloger, S. E., Boyd, N., McKinney, S., Mehl, E., Palmer, C., et al. (2008). Ovarian carcinoma subtypes are different diseases: Implications for biomarker studies. PLoS Medicine, 5(12), e232. https://doi.org/10.1371/journal.pmed.0050232.
Kozar, N., Kruusmaa, K., Bitenc, M., Argamasilla, R., Adsuar, A., Goswami, N., et al. (2018). Metabolomic profiling suggests long chain ceramides and sphingomyelins as a possible diagnostic biomarker of epithelial ovarian cancer. Clinica Chimica Acta, 481, 108–114. https://doi.org/10.1016/j.cca.2018.02.029.
Kurman, R. J., Vang, R., Junge, J., Hannibal, C. G., Kjaer, S. K., & Ie, S., M (2011). Papillary tubal hyperplasia: The putative precursor of ovarian atypical proliferative (borderline) serous tumors, noninvasive implants, and endosalpingiosis. The American Journal of Surgical Pathology, 35(11), 1605–1614. https://doi.org/10.1097/PAS.0b013e318229449f.
Liu, C.-C., Chen, W.-S. E., Lin, C.-C., Liu, H.-C., Chen, H.-Y., Yang, P.-C., et al. (2006). Topology-based cancer classification and related pathway mining using microarray data. Nucleic Acids Research, 34(14), 4069–4080. https://doi.org/10.1093/nar/gkl583.
Lushchak, V. I. (2012). Glutathione homeostasis and functions: Potential targets for medical interventions. Journal of Amino Acids. https://doi.org/10.1155/2012/736837.
Maddocks, O. D. K., Berkers, C. R., Mason, S. M., Zheng, L., Blyth, K., Gottlieb, E., et al. (2013). Serine starvation induces stress and p53-dependent metabolic remodelling in cancer cells. Nature, 493(7433), 542–546.
Malpica, A., Deavers, M. T., Lu, K., Bodurka, D. C., Atkinson, E. N., Gershenson, D. M., et al. (2004). Grading ovarian serous carcinoma using a two-tier system. The American Journal of Surgical Pathology, 28(4), 496–504.
Moore, R. G., McMeekin, D. S., Brown, A. K., DiSilvestro, P., Miller, M. C., Allard, W. J., et al. (2009). A novel multiple marker bioassay utilizing HE4 and CA125 for the prediction of ovarian cancer in patients with a pelvic mass. Gynecologic Oncology, 112(1), 40–46. https://doi.org/10.1016/j.ygyno.2008.08.031.
Moreno, A., & Arus, C. (1996). Quantitative and qualitative characterization of 1H NMR spectra of colon tumors, normal mucosa and their perchloric acid extracts: Decreased levels of myo-inositol in tumours can be detected in intact biopsies. NMR in Biomedicine, 9(1), 33–45, https://doi.org/10.1002/(sici)1099-1492(199602)9:1%3C33::aid-nbm391%3E3.0.co;2-g.
Muñoz, K. A., Harlan, L. C., & Trimble, E. L. (1997). Patterns of care for women with ovarian cancer in the United States. Journal of Clinical Oncology, 15(11), 3408–3415. https://doi.org/10.1200/JCO.1997.15.11.3408.
Nieman, K. M., Kenny, H. A., Penicka, C. V., Ladanyi, A., Buell-Gutbrod, R., Zillhardt, M. R., et al. (2011). Adipocytes promote ovarian cancer metastasis and provide energy for rapid tumor growth. Nature Medicine, 17(11), 1498–1503.
Odunsi, K., Wollman, R. M., Ambrosone, C. B., Hutson, A., McCann, S. E., Tammela, J., et al. (2005). Detection of epithelial ovarian cancer using 1H-NMR-based metabonomics. International Journal of Cancer, 113(5), 782–788. https://doi.org/10.1002/ijc.20651.
Patti, G. J., Yanes, O., & Siuzdak, G. (2012). Innovation: Metabolomics: The apogee of the omics trilogy. Nature Reviews Molecular Cell Biology, 13(4), 263–269. https://doi.org/10.1038/nrm3314.
Pencina, M. J., D’Agostino, R. B. Sr., D’Agostino, R. B. Jr., & Vasan, R. S. (2008). Evaluating the added predictive ability of a new marker: From area under the ROC curve to reclassification and beyond. Statistics in Medicine, 27(2), 157–172. https://doi.org/10.1002/sim.2929 (discussion 207–112).
Perets, R., Wyant, G. A., Muto, K. W., Bijron, Jonathan, G., Poole, B. B., Chin, K. T., et al. (2013). Transformation of the fallopian tube secretory epithelium leads to high-grade serous ovarian cancer in Brca; Tp53; Pten models. Cancer Cell, 24(6), 751–765. https://doi.org/10.1016/j.ccr.2013.10.013.
Petrillo, M., Anchora, L. P., Scambia, G., & Fagotti, A. (2016). Cytoreductive surgery plus platinum-based hyperthermic intraperitoneal chemotherapy in epithelial ovarian cancer: A promising integrated approach to improve locoregional control. The Oncologist, 21(5), 532–534. https://doi.org/10.1634/theoncologist.2015-0500.
Piotto, M., Moussallieh, F.-M., Dillmann, B., Imperiale, A., Neuville, A., Brigand, C., et al. (2008). Metabolic characterization of primary human colorectal cancers using high resolution magic angle spinning 1H magnetic resonance spectroscopy. Metabolomics, 5(3), 292. https://doi.org/10.1007/s11306-008-0151-1.
Platten, M., Wick, W., & Van den Eynde, B. J. (2012). Tryptophan catabolism in cancer: Beyond IDO and tryptophan depletion. Cancer Research, 72(21), 5435–5440. https://doi.org/10.1158/0008-5472.can-12-0569.
Prabhu, A., Sarcar, B., Kahali, S., Yuan, Z., Johnson, J. J., Adam, K.-P., et al. (2014). Cysteine catabolism: A novel metabolic pathway contributing to glioblastoma growth. Cancer Research, 74(3), 787–796. https://doi.org/10.1158/0008-5472.can-13-1423.
Rauh-Hain, J. A., Krivak, T. C., del Carmen, M. G., & Olawaiye, A. B. (2011). Ovarian cancer screening and early detection in the general population. Reviews in Obstetrics & Gynecology, 4(1), 15–21.
Ravanbakhsh, S., Liu, P., Bjordahl, T. C., Mandal, R., Grant, J. R., Wilson, M., et al. (2015). Accurate, fully-automated NMR spectral profiling for metabolomics. PLoS ONE, 10(5), e0124219. https://doi.org/10.1371/journal.pone.0124219.
Rivenbark, A. G., O’Connor, S. M., & Coleman, W. B. (2013). Molecular and cellular heterogeneity in breast cancer: Challenges for personalized medicine. The American Journal of Pathology, 183(4), 1113–1124. https://doi.org/10.1016/j.ajpath.2013.08.002.
Roy, D., Mondal, S., Wang, C., He, X., Khurana, A., Giri, S., et al. (2014). Loss of HSulf-1 promotes altered lipid metabolism in ovarian cancer. Cancer & Metabolism, 2(1), 13. https://doi.org/10.1186/2049-3002-2-13.
Rubin, S. C., Randall, T. C., Armstrong, K. A., Chi, D. S., & Hoskins, W. J. (1999). Ten-year follow-up of ovarian cancer patients after second-look laparotomy with negative findings. Obstetrics & Gynecology, 93(1), 21–24. https://doi.org/10.1016/S0029-7844(98)00334-2.
Salani, R., Kurman, R. J., Giuntoli, R. 2nd, Gardner, G., Bristow, R., Wang, T. L., et al. (2008). Assessment of TP53 mutation using purified tissue samples of ovarian serous carcinomas reveals a higher mutation rate than previously reported and does not correlate with drug resistance. International Journal of Gynecological Cancer, 18(3), 487–491. https://doi.org/10.1111/j.1525-1438.2007.01039.x.
Schmeler, K. M., Sun, C. C., Bodurka, D. C., Deavers, M. T., Malpica, A., Coleman, R. L., et al. (2008). Neoadjuvant chemotherapy for low-grade serous carcinoma of the ovary or peritoneum. Gynecologic Oncology, 108(3), 510–514. https://doi.org/10.1016/j.ygyno.2007.11.013.
Seidman, J. D. (2015). Serous tubal intraepithelial carcinoma localizes to the tubal-peritoneal junction: A pivotal clue to the site of origin of extrauterine high-grade serous carcinoma (ovarian cancer). International Journal of Gynecological Pathology, 34(2), 112–120. https://doi.org/10.1097/pgp.0000000000000123.
Singer, G., Oldt, R. 3rd, Cohen, Y., Wang, B. G., Sidransky, D., Kurman, R. J., et al. (2003). Mutations in BRAF and KRAS characterize the development of low-grade ovarian serous carcinoma. Journal of the National Cancer Institute, 95(6), 484–486.
Souba, W. W. (1993). Glutamine and cancer. Annals of Surgery, 218(6), 715–728.
Stipanuk, M. H., & Ueki, I. (2011). Dealing with methionine/homocysteine sulfur: Cysteine metabolism to taurine and inorganic sulfur. Journal of Inherited Metabolic Disease, 34(1), 17–32. https://doi.org/10.1007/s10545-009-9006-9.
Timmermans, M., Sonke, G. S., Van de Vijver, K. K., van der Aa, M. A., & Kruitwagen, R. (2018). No improvement in long-term survival for epithelial ovarian cancer patients: A population-based study between 1989 and 2014 in the Netherlands. European Journal of Cancer, 88, 31–37. https://doi.org/10.1016/j.ejca.2017.10.030.
Tu, S., Zhang, X.-L., Wan, H.-F., Xia, Y.-Q., Liu, Z.-Q., Yang, X.-H., et al. (2018). Effect of taurine on cell proliferation and apoptosis human lung cancer A549 cells. Oncology Letters, 15(4), 5473–5480. https://doi.org/10.3892/ol.2018.8036.
Turkoglu, O., Zeb, A., Graham, S., Szyperski, T., Szender, J. B., Odunsi, K., et al. (2016). Metabolomics of biomarker discovery in ovarian cancer: A systematic review of the current literature. Metabolomics, 12(4), 60. https://doi.org/10.1007/s11306-016-0990-0.
Uetaki, M., Tabata, S., Nakasuka, F., Soga, T., & Tomita, M. (2015). Metabolomic alterations in human cancer cells by vitamin C-induced oxidative stress. Scientific Reports, 5, 13896. https://doi.org/10.1038/srep13896.
Uyttenhove, C., Pilotte, L., Theate, I., Stroobant, V., Colau, D., Parmentier, N., et al. (2003). Evidence for a tumoral immune resistance mechanism based on tryptophan degradation by indoleamine 2,3-dioxygenase. Nature Medicine, 9(10), 1269–1274.
Vang, R., Shih, I.-M., & Kurman, R. J. (2009). Ovarian low-grade and high-grade serous carcinoma: Pathogenesis, clinicopathologic and molecular biologic features, and diagnostic problems. Advances in Anatomic Pathology, 16(5), 267–282. https://doi.org/10.1097/PAP.0b013e3181b4fffa.
Williams, T. I., Toups, K. L., Saggese, D. A., Kalli, K. R., Cliby, W. A., & Muddiman, D. C. (2007). Epithelial ovarian cancer: Disease etiology, treatment, detection, and investigational gene, metabolite, and protein biomarkers. Journal of Proteome Research, 6(8), 2936–2962. https://doi.org/10.1021/pr070041v.
Woo, H. M., Kim, K. M., Choi, M. H., Jung, B. H., Lee, J., Kong, G., et al. (2009). Mass spectrometry based metabolomic approaches in urinary biomarker study of women’s cancers. Clinica Chimica Acta, 400(1–2), 63–69. https://doi.org/10.1016/j.cca.2008.10.014.
Xia, J., Mandal, R., Sinelnikov, I. V., Broadhurst, D., & Wishart, D. S. (2012). MetaboAnalyst 2.0—A comprehensive server for metabolomic data analysis. Nucleic Acids Research, 40(W1), W127–W133. https://doi.org/10.1093/nar/gks374.
Xia, J., Psychogios, N., Young, N., & Wishart, D. S. (2009). MetaboAnalyst: A web server for metabolomic data analysis and interpretation. Nucleic Acids Research, 37(suppl 2), W652–W660. https://doi.org/10.1093/nar/gkp356.
Xia, J., Sinelnikov, I. V., Han, B., & Wishart, D. S. (2015). MetaboAnalyst 3.0—Making metabolomics more meaningful. Nucleic Acids Research, 43(W1), W251–W257. https://doi.org/10.1093/nar/gkv380.
Yang, M., & Vousden, K. H. (2016). Serine and one-carbon metabolism in cancer. Nature Reviews Cancer, 16(10), 650–662. https://doi.org/10.1038/nrc.2016.81.
Yarchoan, M., Johnson B. A. III, Lutz, E. R., Laheru, D. A., & Jaffee, E. M. (2017). Targeting neoantigens to augment antitumour immunity. Nature Reviews Cancer. https://doi.org/10.1038/nrc.2016.154.
Zhang, T., Wu, X., Ke, C., Yin, M., Li, Z., Fan, L., et al. (2013). Identification of potential biomarkers for ovarian cancer by urinary metabolomic profiling. Journal of Proteome Research, 12(1), 505–512. https://doi.org/10.1021/pr3009572.
Acknowledgements
This research was supported by seed grant funding from the Cancer Research Seed Grant Award at Beaumont Health.
Author information
Authors and Affiliations
Contributions
RBS supervised and designed the experiment, SFG and GG supervised all experimental procedures, AY and PK collected the metabolomics data, AY wrote the manuscript, AY performed statistical data analysis and bioinformatics, all authors read and reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Garg, G., Yilmaz, A., Kumar, P. et al. Targeted metabolomic profiling of low and high grade serous epithelial ovarian cancer tissues: a pilot study. Metabolomics 14, 154 (2018). https://doi.org/10.1007/s11306-018-1448-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-018-1448-3