Abstract
Key message
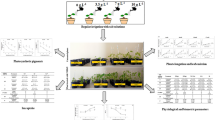
Herbaspirillum rubrisubalbicans decreases growth of rice. Inoculation of rice with H. rubrisubalbicans increased the ACCO mRNA levels and ethylene production. The H. rubrisubalbicans rice interactions were further characterized by proteomic approach.
Abstract
Herbaspirillum rubrisubalbicans is a well-known growth-promoting rhizobacteria that can also act as a mild phyto-pathogen. During colonisation of rice, RT-qPCR analyses showed that H. rubrisubalbicans up-regulates the methionine recycling pathway as well as phyto-siderophore synthesis genes. mRNA levels of ACC oxidase and ethylene levels also increased in rice roots but inoculation with H. rubrisubalbicans impaired growth of the rice plant. A proteomic approach was used to identify proteins specifically modulated by H. rubrisubalbicans in rice and amongst the differentially expressed proteins a V-ATPase and a 14-3-3 protein were down-regulated. Several proteins of H. rubrisubalbicans were identified, including the type VI secretion system effector Hcp1, suggesting that protein secretion play a role colonisation in rice. Finally, the alkyl hydroperoxide reductase, a primary scavenger of endogenous hydrogen peroxide was also identified. Monitoring the levels of reactive oxygen species in the epiphytic bacteria by flow cytometry revealed that H. rubrisubalbicans is subjected to oxidative stress, suggesting that the alkyl hydroperoxide reductase is an important regulator of redox homeostasis in plant-bacteria interactions.








Similar content being viewed by others
References
Alberton D, Müller-Santos M, Brusamarello-Santos LC, Valdameri G, Cordeiro FA, Yates MG, Pedrosa FO, Souza EM (2013) Comparative proteomics analysis of the rice roots colonized by Herbaspirillum seropedicae strain SmR1 reveals induction of the methionine recycling in the plant host. J Proteome Res 12:4757–4768. doi:10.1021/pr400425f
Almagro L, Ros LVG, Belchi-Navarro S, Bru R, Barcelo AR, Pedreno MA (2009) Class III peroxidases in plant defence reactions. J Exp Bot 60:377–390. doi:10.1093/jxb/ern277
Alsterfjord M, Sehnke PC, Arkell A, Larsson H, Svennelid F, Rosenquist M, Ferl RJ, Sommarin M, Larsson C (2004) Plasma membrane H+-ATPase and 14-3-3 Isoforms of Arabidopsis leaves: evidence for isoform specificity in the 14-3-3/H+-ATPase interaction. Plant Cell Physiol 45:1202–1210. doi:10.1093/pcp/pch136
Baldani JI, Baldani VLD, Seldin L, Dobereiner J (1986) Characterization of Herbaspirillum-seropedicae gen. nov., sp. nov., a root-associated nitrogen-fixing bacterium. Int J Syst Bacteriol 36:86–93
Baldani VLD, Baldani JI, Olivares F, Dobereiner J (1992) Identification and ecology of Herbaspirillum-seropedicae and the closely related Pseudomonas rubrisubalbicans. Symbiosis 13:65–73
Berezin I, Mizrachy-Dagry T, Brook E, Mizrahi K, Elazar M, Zhuo S, Saul-Tcherkas V, Shaul O (2008) Overexpression of AtMHX in tobacco causes increased sensitivity to Mg2+, Zn2+, and Cd2 + ions, induction of V-ATPase expression, and a reduction in plant size. Plant Cell Rep 27:939–949. doi:10.1007/s00299-007-0502-9
Book AJ, Gladman NP, Lee SS, Scalf M, Smith LM, Vierstra RD (2010) Affinity purification of the Arabidopsis 26S proteasome reveals a diverse array of plant proteolytic complexes. J Biol Chem 285:25554–25569. doi:10.1074/jbc.M110.136622
Carpentier SC, Witters E, Laukens K, Deckers P, Swennen R, Panis B (2005) Preparation of protein extracts from recalcitrant plant tissues: an evaluation of different methods for two-dimensional gel electrophoresis analysis. Proteomics 5:2497–2507. doi:10.1002/pmic.200401222
Chi F, Yang P, Han F, Jing Y, Shen S (2010) Proteomic analysis of rice seedlings infected by Sinorhizobium meliloti 1021. Proteomics 10:1861–1874. doi:10.1002/pmic.200900694
Clouse SD, Sasse JM (1998) Brassinosteroids: essential regulators of plant growth and development. Annu Rev Plant Phys 49:427–451. doi:10.1146/annurev.arplant.49.1.427
Cloutier P, Coulombe B (2013) Regulation of molecular chaperones through post-translational modifications: decrypting the chaperone code. BBA Gene Regul Mech 1829:443–454. doi:10.1016/j.bbagrm.2013.02.010
Comparot S, Lingiah G, Martin T (2003) Function and specificity of 14-3-3 proteins in the regulation of carbohydrate and nitrogen metabolism. J Exp Bot 54:595–604. doi:10.1093/jxb/erg057
Cordeiro FA, Tadra-Sfeir MZ, Huergo LF, Pedrosa FO, Monteiro RA, Souza EM (2013) Proteomic analysis of Herbaspirillum seropedicae cultivated in the presence of sugar cane extract. J Proteome Res 12:1142–1150. doi:10.1021/pr300746j
David-Assael O, Berezin I, Shoshani-Knaani N, Saul H, Mizrachy-Dagri T, Chen J, Brook E, Shaul O (2006) AtMHX is an auxin and ABA-regulated transporter whose expression pattern suggests a role in metal homeostasis in tissues with photosynthetic potential. Funct Plant Biol 33:661–672. doi:10.1071/fp05295
de Oliveira ALM, Canuto EDL, Urquiaga S, Reis VM, Baldani JI (2006) Yield of micropropagated sugarcane varieties in different soil types following inoculation with diazotrophic bacteria. Plant Soil 284:23–32. doi:10.1007/s11104-006-0025-0
Denison FC, Paul AL, Zupanska AK, Ferl RJ (2011) 14-3-3 proteins in plant physiology. Semin Cell Dev Biol 22:720–727. doi:10.1016/j.semcdb.2011.08.006
Dettmer J, Hong-Hermesdorf A, Stierhof YD, Schumacher K (2006) Vacuolar H+-ATPase activity is required for endocytic and secretory trafficking in Arabidopsis. Plant Cell 18:715–730. doi:10.1105/tpc.105.037978
Dohmann EM, Levesque MP, De Veylder L, Reichardt I, Jurgens G, Schmid M, Schwechheimer C (2008) The Arabidopsis COP9 signalosome is essential for G2 phase progression and genomic stability. Development 135:2013–2022. doi:10.1242/dev.020743
Dos Santos MF, Muniz de Padua VL, Nogueira EM, Hemerly AS, Domont GB (2010) Proteome of Gluconacetobacter diazotrophicus co-cultivated with sugarcane plantlets. J Proteomics 73:917–931. doi:10.1016/j.jprot.2009.12.005
Duca D, Lorv J, Patten CL, Rose D, Glick BR (2014) Indole-3-acetic acid in plant-microbe interactions. Anton Van Lee 106:85–125. doi:10.1007/s10482-013-0095-y
Faurobert M, Mihr C, Bertin N, Pawlowski T, Negroni L, Sommerer N, Causse M (2007) Major proteome variations associated with cherry tomato pericarp development and ripening. Plant Physiol 143:1327–1346. doi:10.1104/pp.106.092817
Gampala SS, Kim TW, He JX et al (2007) An essential role for 14-3-3 proteins in brassinosteroid signal transduction in Arabidopsis. Dev Cell 13:177–189. doi:10.1016/j.devcel.2007.06.009
Hale CN, Wilkie JP (1972) Bacterial leaf stripe of sorghum in New Zealand caused by Pseudomonas rubrisubalbicans. N Z J Agric Res 15:457–463.
Herrmann KM (1995) The shikimate pathway: early steps in the biosynthesis of aromatic compounds. Plant Cell 7:907–919. doi:10.1105/tpc.7.7.907
Hoagland DR, Arnon DI (1950) The water culture method for growing plants without soil. In: California agriculture experimental station circular, College of Agriculture, University of California, Berkeley
James EK, Olivares FL, Baldani JI, Döbereiner J (1997) Herbaspirillum, an endophytic diazotroph colonizing vascular tissue in leaves of Sorghum bicolor L. Moench. J Exp Bot 48:785–797. doi:10.1093/jxb/48.3.785
Kandasamy S, Loganathan K, Muthuraj R, Duraisamy S, Seetharaman S, Thiruvengadam R, Ponnusamy B, Ramasamy S (2009) Understanding the molecular basis of plant growth promotional effect of Pseudomonas fluorescens on rice through protein profiling. Proteome Sci 7:47–56. doi:10.1186/1477-5956-7-47
Kasai K, Kanno T, Akita M, Ikejiri-Kanno Y, Wakasa K, Tozawa Y (2005) Identification of three shikimate kinase genes in rice: characterization of their differential expression during panicle development and of the enzymatic activities of the encoded proteins. Planta 222:438–447. doi:10.1007/s00425-005-1559-8
Klassen G, Pedrosa FO, Souza EM, Funayama S, Rigo LU (1997) Effect of nitrogen compounds on nitrogenase activity in Herbaspirillum seropedicae SMR1. Can J Microbiol 43:887–891
Kluge C, Lahr J, Hanitzsch M, Bolte S, Golldack D, Dietz KJ (2003) New insight into the structure and regulation of the plant vacuolar H+-ATPase. J Bioenerg Biomembr 35:377–388
Lery LM, Hemerly AS, Nogueira EM, Von Kruger WM, Bisch PM (2011) Quantitative proteomic analysis of the interaction between the endophytic plant-growth-promoting bacterium Gluconacetobacter diazotrophicus and sugarcane. Mol Plant Microbe Interact 24:562–576. doi:10.1094/mpmi-08-10-0178
Li J, Xu HH, Liu WC, Zhang XW, Lu YT (2015) Ethylene inhibits root elongation during alkaline stress through AUXIN1 and associated changes in auxin accumulation. Plant Physiol 168:1777–1791. doi:10.1104/pp.15.00523
Ma B, Yin CC, He SJ, Lu X, Zhang WK, Lu TG, Chen SY, Zhang JS (2014) Ethylene-induced inhibition of root growth requires abscisic acid function in rice (Oryza sativa L.) seedlings. PLoS Genet 10:e1004701. doi:10.1371/journal.pgen.1004701
Miche L, Battistoni F, Gemmer S, Belghazi M, Reinhold-Hurek B (2006) Upregulation of jasmonate-inducible defense proteins and differential colonization of roots of Oryza sativa cultivars with the endophyte Azoarcus sp. Mol Plant Microbe Interact 19:502–511. doi:10.1094/mpmi-19-0502
Monteiro RA, Balsanelli E, Wassem R et al (2012) Herbaspirillum-plant interactions: microscopical, histological and molecular aspects. Plant Soil 356:175–196. doi:10.1007/s11104-012-1125-7
Olivares FL, James EK, Baldani JI, Döbereiner J (1997) Infection of mottled stripe disease-susceptible and resistant sugar cane varieties by the endophytic diazotroph Herbaspirillum. New Phytol 135:723–737. doi:10.1046/j.1469-8137.1997.00684.x
Padmanaban S, Lin X, Perera I, Kawamura Y, Sze H (2004) Differential expression of vacuolar H+-ATPase subunit c genes in tissues active in membrane trafficking and their roles in plant growth as revealed by RNAi. Plant Physiol 134:1514–1526. doi:10.1104/pp.103.034025
Pankievicz VC, Camilios-Neto D, Bonato P et al (2016) RNA-seq transcriptional profiling of Herbaspirillum seropedicae colonizing wheat (Triticum aestivum) roots. Plant Mol Biol 90:589–603. doi:10.1007/s11103-016-0430-6
Pedrosa FO, Monteiro RA, Wassem R et al (2011) Genome of Herbaspirillum seropedicae strain SmR1, a specialized diazotrophic endophyte of tropical grasses. PLoS Genet 7:e1002064. doi:10.1371/journal.pgen.1002064
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:e45
Pimentel JP, Olivares F, Pitard RM, Urquiaga S, Akiba F, Dobereiner J (1991) Dinitrogen fixation and infection of grass leaves by Pseudomonas-rubrisubalbicans and Herbaspirillum-seropedicae. Plant Soil 137:61–65. doi:10.1007/bf02187433
Rigden DJ, Lamani E, Mello LV, Littlejohn JE, Jedrzejas MJ (2003) Insights into the catalytic mechanism of cofactor-independent phosphoglycerate mutase from X-ray crystallography, simulated dynamics and molecular modeling. J Mol Biol 328:909–920. doi:10.1016/s0022-2836(03)00350-4
Schmidt MA, Balsanelli E, Faoro H et al (2012) The type III secretion system is necessary for the development of a pathogenic and endophytic interaction between Herbaspirillum rubrisubalbicans and Poaceae. BMC Microbiol 12:98. doi:10.1186/1471-2180-12-98
Schumacher K, Vafeados D, McCarthy M, Sze H, Wilkins T, Chory J (1999) The Arabidopsis det3 mutant reveals a central role for the vacuolar H(+)-ATPase in plant growth and development. Gene Dev 13:3259–3270
Seaver LC, Imlay JA (2001) Alkyl hydroperoxide reductase is the primary scavenger of endogenous hydrogen peroxide in Escherichia coli. J Bacteriol 183:7173–7181. doi:10.1128/JB.183.24.7173-7181.2001
Shaul O (2002) Magnesium transport and function in plants: the tip of the iceberg. Biometals 15:309–323
Straub D, Yang H, Liu Y, Tsap T, Ludewig U (2013) Root ethylene signalling is involved in Miscanthus sinensis growth promotion by the bacterial endophyte Herbaspirillum frisingense GSF30(T). J Exp Bot 64:4603–4615. doi:10.1093/jxb/ert276
Street IH, Aman S, Zubo Y, Ramzan A, Wang X, Shakeel SN, Kieber JJ, Schaller GE (2015) Ethylene inhibits cell proliferation of the Arabidopsis root meristem. Plant Physiol 169:338–350. doi:10.1104/pp.15.00415
Sze H, Li X, Palmgren MG (1999) Energization of plant cell membranes by H+-pumping ATPases. Regulation and biosynthesis. Plant Cell 11:677–690
Tanaka T, Antonio BA, Kikuchi S et al (2008) The rice annotation project database (RAP-DB): 2008 update. Nucleic Acids Res 36:1028–1033. doi:10.1093/nar/gkm978
Valdameri G, Kokot TB, Pedrosa FO, Souza EM (2015) Rapid quantification of rice root-associated bacteria by flow cytometry. Lett Appl Microbiol 60:237–241. doi:10.1111/lam.12351
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3:1–11. doi:10.1186/gb-2002-3-7-research0034
Wang X, Wang D, Wang D, Wang H, Chang L, Yi X, Peng M, Guo A (2012) Systematic comparison of technical details in CBB methods and development of a sensitive GAP stain for comparative proteomic analysis. Electrophoresis 33:296–306. doi:10.1002/elps.201100300
Wang W, Chen LN, Wu H, Zang H, Gao S, Yang Y, Xie S, Gao X (2013) Comparative proteomic analysis of rice seedlings in response to inoculation with Bacillus cereus. Lett Appl Microbiol 56:208–215. doi:10.1111/lam.12035
Wei N, Serino G, Deng XW (2008) The COP9 signalosome: more than a protease. Trends Biochem Sci 33:592–600. doi:10.1016/j.tibs.2008.09.004
Yang CY, Chen YC, Jauh GY, Wang CS (2005) A lily ASR protein involves abscisic acid signaling and confers drought and salt resistance in Arabidopsis. Plant Physiol 139:836–846. doi:10.1104/pp.105.065458
Yang C, Lu X, Ma B, Chen SY, Zhang JS (2015) Ethylene signaling in rice and Arabidopsis: conserved and diverged aspects. Mol Plant 8:495–505. doi:10.1016/j.molp.2015.01.003
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273. doi:10.1146/annurev.arplant.53.091401.143329
Acknowledgements
We thank Roseli Prado, Valter Baura and Marilza Doroti Lamour for technical assistance. We are indebted to William J. Broughton for critical reading of the manuscript and Geraldo Picheth for help with the statistical analysis. The National Institute of Science and Technology of Nitrogen Fixation/CNPq/MCT and Fundação Araucária provided financial support. G.V. thanks PNPD/CAPES for postdoctoral scholarship.
Author information
Authors and Affiliations
Contributions
GV data collection, data analysis and drafting the article. DA, VRM, TBK, CK, LCCBS data collection and analysis. RAM and FOP design of the work and critical revision of the article. EMS conception of the work, data interpretation and critical revision of the article.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Valdameri, G., Alberton, D., Moure, V.R. et al. Herbaspirillum rubrisubalbicans, a mild pathogen impairs growth of rice by augmenting ethylene levels. Plant Mol Biol 94, 625–640 (2017). https://doi.org/10.1007/s11103-017-0629-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-017-0629-1