Abstract
‘Early responsive to dehydration’ (ERD) genes are a group of plant genes having functional roles in plant stress tolerance and development. In this study, we have isolated and characterized a Brassica juncea ‘ERD’ gene (BjERD4) which encodes a novel RNA binding protein. The expression pattern of ERD4 analyzed under different stress conditions showed that transcript levels were increased with dehydration, sodium chloride, low temperature, heat, abscisic acid and salicylic acid treatments. The BjERD4 was found to be localized in the chloroplasts as revealed by Confocal microscopy studies. To study the function, transgenic Arabidopsis plants were generated and analyzed for various morphological and physiological parameters. The overexpressing transgenic lines showed significant increase in number of leaves with more leaf area and larger siliques as compared to wild type plants, whereas RNAi:ERD4 transgenic lines showed reduced leaf number, leaf area, dwarf phenotype and delayed seed germination. Transgenic Arabidopsis plants overexpressing BjERD4 gene also exhibited enhanced tolerance to dehydration and salt stresses, while the knockdown lines were susceptible as compared to wild type plants under similar stress conditions. It was observed that BjERD4 protein could bind RNA as evidenced by the gel-shift assay. The overall results of transcript analysis, RNA gel-shift assay, and transgenic expression, for the first time, show that the BjERD4 is involved in abiotic stress tolerance besides offering new clues about the possible roles of BjERD4 in plant growth and development.






Similar content being viewed by others
References
Abel S, Theologis A (2010) Odyssey of auxin. Cold Spring Harb Perspect Biol 2:a004572
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Alves MS, Fietto LG (2013) Functional diversity of early responsive to dehydration (ERD) genes in soybean. In: Board JE (ed) A comprehensive survey of international soybean research—genetics, physiology, agronomy and nitrogen relationships. InTech, Rijeka, pp 475–487. doi:10.5772/53728
Ansari MI, Lin TP (2010) Molecular analysis of dehydration in plants. Int Res J Plant Sci 12:21–25
Arnon DI (1949) Copper enzymes in isolated chloroplasts, polyphenoxidase in Beta vulgaris. Plant Physiol 24:1–15
Bohnert HJ, Gong Q, Li P, Ma S (2006) Unraveling abiotic stress tolerance mechanisms, getting genomics going. Curr Opin Plant Biol 9:180–188
Cabello JV, Lodeyro AF, Zurbriggen MD (2014) Novel perspectives for the engineering of abiotic stress tolerance in plants. Curr Opin Plant Biol 26:62–70
Camargo SR, Cancado GMA, Ulian EC, Menossi M (2007) Identification of genes responsive to the application of ethanol on sugarcane leaves. Plant Cell Rep 26:2119–2128
Chi YH, Moon JC, Park JH et al (2008) Abnormal chloroplast development and growth inhibition in rice thioredoxin m knock-down plants. Plant Physiol 148:808–817
Du Y, Wang P, Chen J, Song CP (2008) Comprehensive functional analysis of the catalase gene family in Arabidopsis thaliana. J Integr Plant Biol 50:1318–1326
Gutierrez RA, Green PJ, Keegstra K, Ohlrogge JB (2004) Phylogenetic profiling of the Arabidopsis thaliana proteome, What proteins distinguish plants from other organisms? Genome Biol 5(8):R53
Hodgson RA, Raison JK (1991) Superoxide production by thylakoids during chilling and its implication in the susceptibility of plants to chilling-induced photoinhibition. Planta 183:222–228
Jabs T, Dietrich RA, Dangl JL (1996) Initiation of runaway cell death in an Arabidopsis mutant by extracellular superoxide. Science 273:1853–1856
Kim SY, Nam KH (2010) Physiological roles of ERD10 in abiotic stresses and seed germination of Arabidopsis. Plant Cell Rep 29(2):203–209
Kiyosue T, Yamaguchi-Shinozaki K, Shinozaki K (1993) Characterization of cDNA for a dehydration-inducible gene that encodes a Clp A, B-like protein in Arabidopsis thaliana L. Biochem Biophys Res Commun 196:1214–1220
Kiyosue T, Yamaguchi-Shinozaki K, Shinozaki K (1994a) Cloning of cDNAs for genes that are early-responsive to dehydration stress (ERDs) in Arabidopsis thaliana L., identification of three ERDs as HSP cognate genes. Plant Mol Biol 25:791–798
Kiyosue T, Yamaguchi-Shinozaki K, Shinozaki K (1994b) Characterization of two cDNAs (ERD10 and ERD14) corresponding to genes that respond rapidly to dehydration stress in Arabidopsis thaliana. Plant Cell Physiol 106:35–225
Kiyosue T, Abe H, Yamaguchi-Shinozaki K, Shinozaki K (1998) ERD6, a cDNA clone for an early dehydration-induced gene of Arabidopsis, encodes a putative sugar transporter. Biochim Biophys Acta 1370:187–191
Kovacs D, Kalmar E, Torok Z, Tompa P (2008) Chaperon activity of ERD10 and ERD14, two disordered stress-related plant proteins. Plant Physiol 147:381–390
Lee J, Lee H, Kim J, Lee S, Kim DH, Kim S, Hwang I (2011) Both the hydrophobicity and a positively charged region flanking the C-terminal region of the transmembrane domain of signal-anchored proteins play critical roles in determining their targeting specificity to the endoplasmic reticulum or endosymbiotic organelles in Arabidopsis cells. Plant Cell 23:1588–1607
Li Y, Yuan F, Wen Z, Li Y, Wang F, Zhu T, Zhuo W, Jin X, Wang Y, Zhao H, Pei Z-M, Han S (2015) Genome-wide survey and expression analysis of the OSCA gene family in rice. BMC Plant Biol 15:261. doi:10.1186/s12870-015-0653-8
Liu Y, Li H, Shi Y, Song Y, Wang T, Li Y (2009) A maize early responsive to dehydration gene, ZmERD4, provides enhanced drought and salt tolerance in Arabidopsis. Plant Mol Biol Rep 27:542–548
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408
Mano Y, Nemoto K (2012) The pathway of auxin biosynthesis in plants. J Exp Bot 63:2853–2872
Mantri N, Patade V, Suprasanna P, Ford R, Pang E (2012) Abiotic stress responses in plants, present and future. In: Ahmad P, Prasad MNV (eds) Abiotic stress responses in plants, metabolism, productivity and sustainability. Springer, New York, pp 1–19
McKersie BD, Murnaghan J, Jones KS, Bowley SR (2000) Iron-superoxide dismutase expression in transgenic alfalfa increases winter survival without a detectable increase in photosynthetic oxidative stress tolerance. Plant Physiol 122:1427–1437
McQualter RB, Dookun-Saumtally A (2007) Expression profiling of abiotic-stress inducible genes in sugarcane. In: XXVI Congress, International Society of Sugar Cane Technologists, pp 878–888
Mitschke J, Fuss J, Blum T, Höglund A, Reski R et al (2009) Prediction of dual protein targeting to plant organelles. New Phytol 183:224–235
Mittler R, Vanderauwera S, Gollery M, Van Breusegem F (2004) Reactive oxygen gene network of plants. Trends Plant Sci 9:490–498
Nakashima K, Satoh R, Kiyosue T, Yamaguchi-Shinozaki K, Shinozaki K (1998) A gene encoding proline dehydrogenase is not only induced by proline and hypoosmolarity, but is also developmentally regulated in the reproductive organs of Arabidopsis. Plant Physiol 118:12–33
Nickelsen J (2003) Chloroplast RNA-binding proteins. Curr Genet 43:392–399
Oono Y, Seki M, Nanjo T, Narusaka M, Fujita M, Satoh R, Satou M, Sakurai T, Ishida J, Akiyama K, Iida K, Maruyama K, Satoh S, Yamaguchi-Shinozaki K, Shinozaki K (2003) Monitoring expression profiles of Arabidopsis gene expression during rehydration process after dehydration using ca 7000 full-length cDNA microarray. Plant J 34:868–887
Pérez-Clemente RM, Vives V, Zandalinas SI, López-Climent MF, Muñoz V, Gómez-Cadenas A (2013) Biotechnological approaches to study plant responses to stress. BioMed Res Int 654120. doi:10.1155/2013/654120
Rabbani MA, Maruyama K, Abe H, Khan MA, Katsura K, Ito Y, Yamaguchi-Shinozaki K (2003) Monitoring expression profiles of rice genes under cold, drought, and high-salinity stresses and abscisic acid application using cDNA microarray and RNA gel-blot analyses. Plant Physiol 133(4):1755–1767
Rai AN, Suprasanna P, D’Souza SF, Kumar V (2012) Membrane topology and predicted RNA-binding function of the ‘early responsive to dehydration (ERD4)’ plant protein. PLoS ONE 7(3):e32658
Shepard JF, Totten RD (1975) Isolation and regeneration of tobacco mesophyll cell protoplasts under low osmotic conditions. Plant Physiol 55:689–694
Shinozaki K, Yamaguchi-Shinozaki K (1997) Gene expression and signal transduction in water-stress response. Plant Physiol 115:327–334
Shinozaki K, Yamaguchi-Shinozaki K (2000) Molecular responses to dehydration and low temperature, differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol 3:217–223
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58:221–227
Soitamo AJ, Piippo M, Allahverdiyeva Y, Battchikova Aro EM (2008) Light has a specific role in modulating Arabidopsis gene expression at low temperature. BMC Plant Biol 8:13. doi:10.1186/1471-2229-8-13
Taji T, Seki M, Yamaguchi-Shinozaki K, Kamada H, Giraudat J, Shinozaki K (1999) Mapping of 25 drought-inducible genes, RD and ERD, in Arabidopsis thaliana. Plant Cell Physiol 40:119–123
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4, Molecular evolutionary genetics analysis (MEGA) software version 40. Mol Biol Evol 24:1596–1599
Thordal-Christensen H, Zhang Z, Wei Y, Collinge DB (1997) Subcellular localization of H2O2 in plants, H2O2 accumulation in papillae and hypersensitive response during barley-powdery mildew interaction. Plant J 11:1187–1194
Tran LS, Mochida K (2010) Functional genomics of soybean for improvement of productivity in adverse conditions. Funct Integr Genomics 10(4):447–462
Tuteja N (2007) Abscisic acid and abiotic stress signaling. Plant Signal Behav 2(3):135–138
Tzin V, Galili G (2010) The biosynthetic pathways for shikimate and aromatic amino acids in Arabidopsis thaliana. Arabidopsis Book 8:e0132
Umezawa T, Fujita M, Fujita Y, Yamaguchi-Shinozaki K, Shinozaki K (2006) Engineering drought tolerance in plants: discovering and tailoring genes to unlock the future. Curr Opin Biotechnol 17:113–122
Xing Y, Jia W, Zhang J (2007) AtMEK1 mediates stress-induced gene expression of CAT1 catalase by triggering H2O2 production in Arabidopsis. J Exp Bot 58:2969–2981
Xiong L, Lee BH, Ishitani M, Lee H, Zhang C, Zhu JK (2001) FIERY1 encoding an inositol polyphosphate 1 phosphatase is a negative regulator of abscisic acid and stress signaling in Arabidopsis. Genes Dev 15:1971–1984
Xu T, Han JH, Kang H (2013) Structural features important for the RNA chaperone activity of zinc finger-containing glycine-rich RNA-binding proteins from wheat (Triticum avestivum) and rice (Oryza sativa). Phytochemistry 94:28–35
Xu T, Sy ND, Lee HJ, Kwak KJ, Gu L, Kim JI, Kang H (2014) Functional characterization of a chloroplast-targeted RNA-binding protein CRP1 in Arabidopsis thaliana under abiotic stress conditions. J Plant Biol 57(6):349–356
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Yokotani N, Higuchi M, Kondou Y, Ichikawa T, Iwabuchi M, Hirochika H, Matsui M, Oda K (2011) A novel chloroplast protein, CEST induces tolerance to multiple environmental stresses and reduces photooxidative damage in transgenic Arabidopsis. J Exp Bot 62:557–569
Yuan F, Yang H, Xue Y, Kong D, Ye R, Li C, Zhang J, Theprungsirikul L, Shrift T, Krichilsky B et al (2014) OSCA1 mediates osmotic-stress-evoked Ca2+ increases vital for osmosensing in Arabidopsis. Nature 514(7522):367–371
Zhang XL, Zhang ZJ, Chen J, Chen Q, Wang XC, Huang RF (2005) Expressing TERF1 in tobacco enhances drought tolerance and abscisic acid sensitivity during seedling development. Planta 222(3):494–501
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Acknowledgments
This study was funded through the departmental funds provided by BARC.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors of this manuscript hereby state that there are no conflicts of interest.
Research involving human participants and/or animals
This research does not involve human participants and/or animals.
Additional information
This article has been retracted by the authors due to the fact that panels in Figure 3C are duplicated for different transgenic lines. All authors have agreed to the retraction of this paper and deeply regret the duplication. The authors maintain that the duplication does not affect the interpretation or conclusions of their study. They will seek to publish the results in a new manuscript with a corrected version of the figure to corroborate the findings of this work. The authors deeply regret the inconvenience caused by the retraction
An erratum to this article is available at http://dx.doi.org/10.1007/s11103-016-0574-4.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2015_423_MOESM1_ESM.ppt
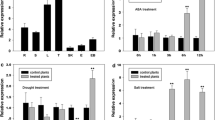
Sup. Figure 1: Expression analysis of ERD4 in leaves: Expression analysis of ERD4 gene using qRT PCR analysis in leaves under salinity (NaCl, 200 mM) (A), drought (PEG, 20 %) (B) at different time periods (temporal expression) and other abiotic stresses after 1 h (C). Tubulin was used as an internal control. The vertical column indicates the relative ERD4 transcript level. The mean and SE from three independent experiments are shown. * indicates significant differences in comparison with the control at P < 0.05 (PPT 182 kb)
About this article
Cite this article
Rai, A.N., Tamirisa, S., Rao, K.V. et al. RETRACTED ARTICLE: Brassica RNA binding protein ERD4 is involved in conferring salt, drought tolerance and enhancing plant growth in Arabidopsis. Plant Mol Biol 90, 375–387 (2016). https://doi.org/10.1007/s11103-015-0423-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-015-0423-x