Abstract
Leucine-rich repeat receptor-like protein kinases (LRR RLKs) represent the largest group of Arabidopsis RLKs with approximately 235 members. A minority of these LRR RLKs have been assigned to diverse roles in development, pathogen resistance and hormone perception. Using a reverse genetics approach, a collection of homozygous T-DNA insertion lines for 69 root expressed LRR RLK genes was screened for root developmental defects and altered response after exposure to environmental, hormonal/chemical and abiotic stress. The obtained data demonstrate that LRR RLKs play a role in a wide variety of signal transduction pathways related to hormone and abiotic stress responses. The described collection of T-DNA insertion mutants provides a valuable tool for future research into the function of LRR RLK genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Multicellular organisms sense and respond to both external and internal signals in an intricate and accurate way for survival and coordinate development. A multi-step signal transduction set up, involving receptor protein kinases using phosphorylation status to transduce external messages into the cell, creates the needed complexity for sophisticated response regulation (Wang et al. 2007).
The receptor-like protein kinases (RLKs) are the largest class of Arabidopsis protein kinases forming a monophyletic group that contains both transmembrane and cytoplasmic protein kinases (Shiu and Bleecker 2001a, b, 2003). The Arabidopsis genome contains over 600 RLK genes representing about 2.5% of the protein encoding genes. RLKs can function as (hetero-)dimers, adding to their signaling, sensing and regulatory potential, indicating that Arabidopsis is able to perceive a wide range of signals (Johnson and Ingram 2005). The best studied RLKs are those containing extracellular leucine-rich repeats (LRRs). LRRs are tandem repeats of approximately 24 amino acids with conserved leucines. The group of LRR RLKs is the largest RLK class with over 200 members, divided over 13 subfamilies (LRR I to XIII) classified according to the organization of LRRs in the extracellular domain (Shiu and Bleecker 2001a, 2003).
Over the years an increasing number of RLKs have been assigned functions in development, pathogen resistance and hormone perception (Dievart and Clark 2004; Morillo and Tax 2006). In addition, many RLKs show a transcriptional response upon hormone treatment (Chae et al. 2009). Two of the best characterized LRR RLKs in Arabidopsis are CLAVATA 1 (CLV1) and BRASSINOSTEROID INSENSITIVE 1 (BRI1) (Rieu and Laux 2009; Aker and de Vries 2008). The CLV pathway regulates stem cell proliferation and differentiation in the shoot apical meristem (SAM). CLV2 encodes a LRR receptor-like protein required for the stability of CLV1 that is the receptor for CLV3, a small secreted CLE family peptide ligand (Ogawa et al. 2008). Mutations in any of these three CLV genes cause an ectopic accumulation of stem cells and a progressive enlargement of the shoot meristem. Downstream of CLV1 signaling is the homeobox transcription factor WUSCHEL (WUS) that forms a negative feedback loop with CLV3 to maintain meristem size. (Sablowski 2007). Brassinosteroid (BR) signaling through the BRI1 receptor regulates plant growth and development through a complex signal transduction pathway. Binding of BRs to BRI1, releases its negative regulator BKI1, thereby increasing the affinity for the BRI1-ASSOCIATED KINASE (BAK1/SERK3) co-receptor, and allowing downstream signaling to the phosphorylation sensitive BRI1 EMS SUPPRESSOR1 (BES1) and BRASSINAZOLE RESISTANT1 (BZR1) transcription factors (Vert et al. 2005; Aker and de Vries 2008).
Roots arise from stereotyped embryonic divisions and harbor a clearly discernible stem cell set. The radial organization of the Arabidopsis root is maintained by asymmetric cell divisions of different stem cells and their daughters that are located in the root meristem (Ten Hove and Heidstra 2008). These stem cells surround a small group of cells, the quiescent center (QC), required for their maintenance. The current hypothesis is that the angiosperm root meristem has evolved from the shoot apical meristem (Stahl et al. 2009). In addition, roots respond sensitively to plant growth factors and to environmental signaling (Wolters and Jurgens 2009; Iyer-Pascuzzi et al. 2009). Key regulatory themes in the shoot have been found to be significant in the development and growth regulation of the root, involving phytohormones, transcription factors as well as peptide ligands and their receptors. Several observations suggests that a signaling pathway involving CLV1-like LRR RLKs may function in root stem cell maintenance: (1) The similarity between the two apical meristems (Scheres 2007); (2) the fact that overexpression or application of A-class CLE peptides cause differentiation of shoot and root meristems (Ito et al. 2006; Kinoshita et al. 2007; Whitford et al. 2008); and (3) the fact that WUS and its family member WOX5 can substitute for each other in stem cell maintenance (Sarkar et al. 2007).
To study possible conservation of LRR RLK function in shoot and root meristem maintenance, a collection of homozygous T-DNA insertion lines for root expressed LRR RLKs was generated and investigated for root developmental phenotypes. In addition, these lines were screened for altered response to a series of hormone/chemical and abiotic stress treatments. Despite the absence of new developmental phenotypes under normal growth conditions we implicate several previously characterized as well as uncharacterized LRR RLKs in hormone and abiotic stress responses.
Materials and methods
Plant materials
The T-DNA Express database of the SALK Institute Genomic Analysis Laboratory (SIGnAL; http://www.signal.salk.edu/cgi-bin/tdnaexpress) was employed to identify putative T-DNA insertion mutants. We aimed to select T-DNA insertions within the coding region of the gene to enhance the likelihood of successful disruption of gene function. When unavailable, lines were selected with predicted intron or promoter (1,000 bp promoter and 300 bp 5′ UTR) insertions. Available lines of interest generated by The Salk Institute for Biological Studies (SALK) (Alonso et al. 2003), the German plant genomics research program (GABI) (Rosso et al. 2003) and Syngenta Arabidopsis Insertion Library (SAIL) (Sessions et al. 2002) were obtained from the Arabidopsis Biological Resource Center (ABRC; http://www.abrc.osu.edu/), the Nottingham Arabidopsis Stock Centre (NASC; http://www.arabidopsis.info/) or Syngenta. rlk902 (Tarutani et al. 2004) was kindly donated by Dr. Yoshihito Suzuki (University of Tokyo, Japan), bam1-1 and bam3-2 (Deyoung et al. 2006) by Dr. Steven Clark (University of Michigan, USA) and er-105, erl1-2 and erl2-1 (Shpak et al. 2004b) by Dr. Keiko Torii (University of Washington, USA). All T-DNA mutants used were of Columbia ecotype. The authenticity of T-DNA mutations was verified by PCR. Genomic DNA was isolated from approximately 10 individual plants per T-DNA line. Primers (Table S1 in ‘‘Supplementary material”) were generated using the T-DNA primer design tool (http://www.signal.salk.edu/tdnaprimers.2.html) and used in two separate PCR reactions. Reaction one contained a T-DNA specific and gene specific primer to check for the presence of an insertion whereas reaction two contained two gene specific primers spanning the putative insertion site to check for nondisrupted alleles. Plants were considered homozygous for the T-DNA insertion when only PCR reaction one yielded a product, which was subsequently confirmed in their progeny.
Plant growth conditions and microscopy
All seeds (wild type control and T-DNA insertion mutants) used in the assays were obtained from plants harvested at similar time points. Seeds were gas-sterilized in a desiccator for 2 h with 100 ml of bleach (4% NaClO) mixed with 3 ml of HCl in a beaker or were surface-sterilized in 20% bleach for 20 min. Sterilized seeds were imbibed in sterile water containing 0.1% agarose for 3–4 days at 4°C in the dark. For phenotypic analysis of root growth, seeds were germinated on half strength Murashige and Skoog (MS) salt mixture, 1% sucrose and 0.5 g/l 2-(N-morpholino) ethanesulfonic acid (MES), pH5.8, in 0.8% agar (standard medium). Plates were incubated in a near vertical position at 22°C with a cycle of 16 h light/8 h dark. Roots were analyzed after 4–8 days using Nomarksi optics and confocal microscopy. Starch granules were visualized as described (Willemsen et al. 1998). For confocal microscopy, roots were mounted in propidium iodide (PI; 20 μg/ml in distilled water). Seedlings were transferred to soil for further phenotypic analysis of general growth and development. Soil grown plants were cultured in a growth chamber at 22°C, 70% relative humidity and a cycle of 16 h light/8 h dark. For temperature assays, seedlings were grown on plates in a near vertical position and a cycle of 8 h light/16 h dark and analyzed after 7 days (30°C) or 14 days (15°C), respectively. Root gravitropism was studied by growing seedlings on plates at near vertical position that were rotated 90° after 3 days of growth. After 1–2 days, the bending angle of the root was measured.
Conditional phenotypes assays
20 seeds per T-DNA insertion line and 20 wild type control seeds were plated for each experiment. To minimize plate position effects, seeds were plated in alternate groups, i.e. one top row of 10 wild type followed by 10 mutant seeds, and a bottom second row of 10 mutant followed by 10 wild type seeds. For vertical, light germination assays, seed germination was determined by recording the presence of a radicle. For vertical, light, root length and general root growth assays, uniformity of germination was analyzed after 1–2 days and only seedlings that germinated at the same time were taken along for qualitative root length or general growth measurements. A greater than 20% difference in root length between wt and mutant seedlings was set as the qualitative criterion for scoring a genotype as resistant or sensitive to the particular treatment. To test whether RLK genes are involved in responsiveness to stress, seeds were directly sown on standard medium supplemented with hormone (precursor) 1-aminocyclopropane-1-carboxylic acid (ACC), abscisic acid (ABA), methyl-jasmonic acid (MeJa), 24-epibrassinolide (EBL), 6-benzylamino-purine (6-BAP), indole-3-acetic acid (IAA) or with CLE peptides, respectively, at concentrations listed in Table S2 in ‘‘Supplementary material”. For N-1-naphthylphthalamic acid (NPA) treatment, seedlings were transferred 3 days after germination to standard medium supplemented with NPA. For abiotic stress experiments, seeds were directly sown on standard medium amended with 200 mM NaCl and 400 mM mannitol, respectively. The ratio of mutant germination percentage over wild type germination percentage (Table S4) exhibited a bimodal distribution. Based on this distribution, a greater than 1.7 fold increase in germination percentage relative to wild type was chosen as threshold to be called NaCl or mannitol resistant. This threshold identifies only the T-DNA insertion lines that comprise the distinct upper part of the distribution as resistant. Susceptibility to salt stress was tested by sowing seeds on standard medium and transferring seedlings to standard medium supplemented with 50 mM NaCl after 3 days, followed by a 180° rotation. Root growth was measured after 1–2 days. To test for altered responses to sucrose, seeds were sown on standard medium lacking sucrose and transferred after 3 days to standard medium supplemented with 3.5% sucrose.
Kinase phylogeny and coexpression analysis
From the 69 RLKs analyzed in this study 352 positions were aligned automatically corresponding to the kinase domain using the program ClustalW implemented in the Bioedit Software (Hall 1999). The phylogenetic tree was reconstructed using neighbor-joining method in the MEGA package v4.0 (Tamura et al. 2007) with 500 bootstrap resampling. In order to detect coregulation between the kinases, we selected several microarray experiments showing differential expression for at least a subset of the 69 RLKs analyzed. Processed data for microarray experiments were obtained from ArrayExpress (http://www.ebi.ac.uk/microarray-as/ae/) i.e. E-GEOD-3709 for abiotic stress (one slide per treatment), E-GEOD-5617 for light, E-GEOD-7643 for NaCl and E-GEOD-18975 for IAA (only slides with accession Col-0). Genes were clustered based on the expression profiles to find coexpressed gene clusters. Hierarchical clustering of microarray data was performed in MultiExperiment Viewer (MeV) v4.5.0 (Saeed et al. 2003), using Pearson correlation and Average Linkage Clustering algorithm.
Results
A homozygous T-DNA insertion mutant collection for root expressed LRR RLKs
Starting from the assumption that a conserved CLV-like pathway for Arabidopsis root meristem maintenance exists, we investigated LRR RLK function in root development by taking a reverse genetics approach. We first analyzed the expression patterns in the different tissues of the root meristem of all LRR RLKs from subfamilies II, III, IV, VII, VIII-2, X, XI and XIII using the in silico expression database of the Arabidopsis root (http:\\www.arexdb.org) (Birnbaum et al. 2003; Brady et al. 2007), We also included 6 kinases that belong to the same monophyletic group as the LRR RLKs but originate from different classes (L-lectin, SD-1, URK1) that were identified as putative targets of root expressed transcription factors (RH, unpublished data). 87 RLKs of mostly unknown function (Table S3 in “Supplementary material”) were selected that are expressed in different tissues of the root meristem (Fig. 1). Comparison with the GENEVESTIGATOR database indicated that all kinases were also expressed in other tissues (http://www.genevestigator.com/gv/index.jsp, data not shown).
LRR RLK root expression profiles. Heat map of the expression patterns of the 87 LRR RLK genes in the root based on tissue types and longitudinal sections. The expression indices for each tissue/section were obtained from (Brady et al. 2007) and were visualized in MultiExperiment Viewer (MeV) v4.5.0 (Saeed et al. 2003). Colors indicate lowered (black) or increased (yellow) transcript accumulation relative to the respective controls within a 0 to +3.5 range
We collected putative T-DNA insertion lines for these root expressed RLKs, which were subsequently tested by PCR based genotyping and built a collection of 135 homozygous T-DNA lines representing 69 RLK genes, harboring insertions in: (1) exons (98 lines); (2) introns (11 lines); (3) within 500 nucleotides upstream of the open reading frame (15 lines); and (4) between 500 and 1,000 (11 lines) nucleotides upstream of the open reading frame (Table 1). For 18 RLK genes we failed to generate homozygous T-DNA insertion lines and these were excluded from our study. Together this collection makes up around 30% of the LRR RLKs present in the Arabidopsis genome.
Developmental phenotype analysis
To identify LRR RLKs involved in root growth and development we phenotypically analyzed 4–8-day-old roots of the homozygous T-DNA insertion mutants using both confocal microscopy and nomarski optics. We tested the integrity of the stem cell niche based on morphology and absence of starch accumulation in columella stem cells. Two lines appeared to have a root developmental defect. rlk902, showed a reduced root length and meristem size and was further characterized in a separate study. The SALK_009453c line, homozygous for a T-DNA insertion in At2g31880, segregated in a recessive manner for a short root phenotype. This suggests that a mutation unlinked to the T-DNA is responsible for the observed phenotype. Apart from previously reported phenotypes for a number of LRK mutants (e.g. erecta (reduced size and compact stature), bak1/serk3 (weak bri1 phenotype), pskr1 and brl2/vh1 (premature senescence), bam3 (small stature, short siliques, extreme branching)), we did not observe obvious above ground defects in T-DNA insertion lines for the other LRK genes when compared to wild type plants grown under long day conditions for 4–8 weeks. Together, these results suggest that the LRR RLKs analyzed are functionally redundant for developmental pathways or function only under specific stimuli and/or in other than root tissues.
Susceptibility to CLE peptide treatment
Overexpression and exogenous application of CLV3 and other A-class CLE peptides leads to general loss of meristematic activity, suggesting that these peptides act in controlling shoot and root meristem size, whereas B-class CLE peptides (CLE41-CLE44) suppress the differentiation of xylem cells from stem cell-like procambial cells and promote cell division (Ito et al. 2006; Kinoshita et al. 2007; Whitford et al. 2008). In the shoot meristem, CLV3 is the ligand for the CLV1 receptor (Ogawa et al. 2008). To identify putative receptors involved in the perception of CLE peptides in roots, we treated the LRR RLK mutants with synthetic CLV3 and/or CLE19 peptide. Compromised receptors should not be able to transduce any signal upon ligand binding and mutant plant meristems should be similar to untreated wild type meristems. Only the N585175 line, homozygous for a T-DNA insertion in AIK3, displayed resistance to both CLV3 and CLE19 peptide treatment, albeit in a Mendelian fashion (~ 25% resistance, corresponding to a recessive phenotype). PCR analysis confirmed that the T-DNA insertion in AIK3 did not co-segregate with the observed CLE peptide resistance, indicating the presence of an additional mutation. The results indicate that none of the tested RLKs are involved in CLE perception.
Analysis for conditional phenotypes
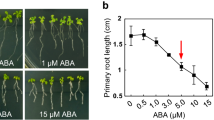
In addition to developmental phenotyping, we undertook a broad-spectrum panel of environmental assays on the T-DNA insertion lines to test the involvement of individual LRR RLKs in the response to these stimuli (see "Materials and methods"). We produced dose response curves for conditions not yet published using the online Arabidopsis Gantlet Project database (http://www.thale.biol.wwu.edu/index.html). Quantifiable traits as root length and germination were used and threshold doses were established to test for sensitivity and saturation doses for insensitivity. No consistent differential responsiveness was observed in comparison to wild type plants for treatments with ACC, ABA, MeJa, EBL, low NaCl, sucrose, low and high temperature or gravitropism (data not shown).
LRR RLK mutants affected in auxin response
Hormonal signaling plays a critical role in almost every aspect of plant development, from embryogenesis to senescence. Although the molecular details of hormone action remain largely unknown, receptors for the major hormones have now been identified (Bishopp et al. 2006; Wolters and Jurgens 2009). The plant hormone auxin plays a key role in many aspects of plant growth and development. Auxin transport is facilitated by auxin influx and efflux carriers, integral plasma membrane proteins that transport auxin molecules into and out of the cell, respectively (Petrášek and Friml 2009). The nuclear localized receptors AFB1, AFB2, AFB3 and TIR1, which are the F-box subunits of the E3-ubiquitin ligase complexes SCF-TIR1 and SCF-AFB bind auxin. This results in the degradation of the Aux/IAA transcriptional repressor proteins by the 26S proteasome and subsequent ARF dependent activation of transcription (Dharmasiri et al. 2005). However, not all auxin-regulated processes can be easily attributed to this type of signaling. Another putative auxin receptor is ABP1 that binds auxin and is implicated in a set of early auxin responses such as rapid activation of ion fluxes at the plasma membrane (Badescu and Napier 2006; Tromas et al. 2009).
To identify LRR RLKs involved in auxin signaling we screened seedling root growth response to the natural auxin IAA and the phytotropin polar auxin transport inhibitor NPA. We tested the T-DNA insertion mutants using concentrations that slightly inhibit root growth in wild type seedlings, i.e. 0.2 μM IAA and 5 μM NPA. For the IAA treatment, 19 T-DNA insertion lines corresponding to 16 RLK genes showed a consistent enhanced root length compared to wild type, indicative of increased resistance to IAA (Table 1). In contrast, 2 T-DNA insertion lines corresponding to BAK1/SERK3 and IRK genes showed a consistent enhanced root growth inhibition, indicative of increased sensitivity to IAA. 23 T-DNA insertion lines corresponding to 22 RLK genes were found to be resistant to NPA treatment (Table 1). Reversely, 3 T-DNA insertion lines, corresponding to BAK1/SERK3 and PSKR1 genes, were more sensitive to NPA.
Resistance to the phytohormone auxin and polar auxin transport inhibitors frequently coincides (Fujita and Syono 1997). These observations are confirmed in our study as we observed an overlap between IAA and NPA resistance observed for T-DNA insertions in 7 genes: At1g67510, At5g43020, At1g53440, At4g03230, PEPR1, BAM1 and ERL2, whereas the T-DNA insertion line N534523 (BAK1/SERK3) showed increased sensitivity for both IAA and NPA. Together, our results suggest that a number of LRR RLKs are involved in auxin signaling and/or response.
LRR RLK mutants affected in cytokinin response
Cytokinins, generally acting antagonistically to auxin (Bishopp et al. 2006), have been shown to play a key role in the regulation of root growth and meristem size (Dello Ioio et al. 2007). Plants respond to cytokinins via a two-component signaling pathway involving the transmembrane histidine kinases AHK2, AHK3 and AHK4/CRE1 (Dello Ioio et al. 2008). These receptors transfer the signal via phosphorelay to the nucleus, thereby activating negative (type-A) and positive (type-B) regulators (ARRs) of the cytokinin response. Type-B ARR transcription factors activate the transcription of cytokinin primary response genes, including type-A ARRs, thereby forming a negative feedback loop to control cytokinin responses (Bishopp et al. 2006).
To explore whether LRR RLKs are involved in cytokinin signaling and/or response we screened seedling root growth response to the synthetic cytokinin 6-BAP. We tested the T-DNA insertion mutants using concentrations that slightly inhibit root growth in wild type seedlings, i.e. 0.1 μM 6-BAP. One homozygous T-DNA insertion line showed a consistent increased (At2g33170) and one showed a consistent reduced (BAK1/SERK3) root length compared to wild type (Table 1). These results suggest that At2g33170 and BAK1/SERK3 mediate cytokinin control on root growth.
Identification of LRR RLKs involved in light signaling and/or response
Light is one of the key external factors controlling seed germination and dormancy (Penfield and King 2009). Perception and response to this stimulus ensures that seedling emergence and growth occur at the most advantageous time. The effect of light on seed germination is mainly conveyed by photoreceptors called phytochromes. Additionally, different hormones favor (gibberellin, ethylene, BR) or repress (ABA) germination.
The LRR RLK T-DNA insertion mutants were screened for light requirement by analyzing their germination potential (measured by radicle emergence) in the absence of light. We scored mutants as light sensitive when they showed germination levels of lower than 25% at 25°C in the dark. High germination levels were observed for all mutant lines and co-plated wild type controls in the light. 11 T-DNA insertion lines showed dark germination levels lower than 25% (Table 1), implicating involvement of the corresponding to 10 RLK genes in light signaling and/or response.
LRR RLK mutants affected in salt and osmotic stress tolerance
Plants vary greatly in their tolerance to abiotic stress such as salt (Xiong and Zhu 2002). Whereas halophytes can complete their life cycle under saline conditions, glycophytes are more sensitive to salt stress although their tolerance varies widely between species and even among varieties. Arabidopsis thaliana is a glycophytic, salt intolerant plant. It is assumed that salt is perceived by specific receptors with RLKs, two component histidine kinases and G-protein-associated receptors implicated in this process (Xiong and Zhu 2001). Recently, the LRR RLK encoding SRLK was implicated in the regulation of the adaptation of Medicago truncatula roots to salt stress (de Lorenzo et al. 2009).
To determine whether any of the Arabidopsis LRR RLKs in our mutant set play a role in the perception of abiotic stress, we have tested the T-DNA insertion lines for their ability to germinate compared to co-plated wild type control seeds on medium containing 200 mM NaCl or 400 mM mannitol. A greater than 1.7 fold increase in germination percentage to wild type was chosen to be called NaCl or mannitol resistant (see "Materials and methods"). 26 T-DNA insertion lines displayed enhanced NaCl tolerance corresponding to 23 RLK genes (Table 1). For mannitol treatment, 37 T-DNA lines corresponding to 31 RLK genes showed enhanced resistance to mannitol (Table 1).
High salinity causes both hyperionic and hyperosmotic stress effects, whereas mannitol induces hyperosmotic stress (Hasegawa et al. 2000). To assess whether the altered response to NaCl treatment was due to altered tolerance towards ionic and/or osmotic stress effects we analyzed the overlap in T-DNA insertion lines with altered NaCl and mannitol responses. 13 T-DNA insertion lines for 13 genes: At3g55550, At2g36570, At5g07620, At5g43020, At5g67200, At5g58150, At4g03230, BAM1, BRL2/VH1, BRL3, ERL1, GSO1 and MRLK1/IMK3 were tolerant to both ionic effects and osmotic pressure, suggesting that these mutants are primarily osmotolerant. Together, our results suggest that these kinases play a role in plant salt and/or osmotic stress tolerance.
Bioinformatic analyses to uncover trends in altered conditional responses
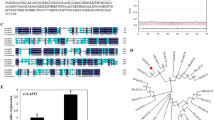
We next investigated whether there was a possible link between the obtained functional data and LRR RLK phylogeny. A neighborhood joining tree of the 69 tested LRR RLKs was constructed using the kinase domain with each treatment depicted on the branches for each of the kinases (Fig. 2). No patterns emerged that connect LRR RLK phylogeny and mutant response.
Combination of RLK phylogeny with functional data. Neighborhood joining tree of the 69 LRR RLKs tested. Subfamilies are indicated. Positive and negative response to tested conditions was color coded on the branches for each of the kinases as indicated. The numbers on the base of the branch indicate bootstrap support out of 500 replicates
We next investigated a possible link between LRR RLK behavior at the transcriptome level with the functional characterization of corresponding mutants. A hierarchical clustering of the 69 studied LRR RLKs was performed based on their behavior in different public microarray experiments using conditional stresses comparable to those described here. We then compared the transcriptomic data with our functional characterization of the mutants and analyzed whether there was an overlap. LRR RLK gene clusters with similar behavior at the transcriptome level upon different stress treatments can be distinguished (Fig. 3). However, comparison of these clustered expression patterns to RLK phylogeny and function in these stress responses did not reveal any significant correlation (Fig. 3).
Cluster analysis of LRR RLKs expression combined with functional data. The 69 studied LRR RLKs are clustered based on their expression behavior during different public microarray experiments upon a range of abiotic stress, light, NaCl and IAA treatment. Each column represents the results from one microarray condition. Colors indicate lowered (green) or increased (red) transcript accumulation relative to the respective controls within a −5.5 to +2.5 range. The results of the conditional tests for mannitol, dark, NaCl and IAA (modified from Table 1) are depicted on the right. R resistant; S sensitive; – similar to wild type
Discussion
Here, we have undertaken a reverse genetics approach concentrating on root meristem expressed LRR RLKs with two objectives: first, to investigate the function of CLV1 paralogs in root meristem maintenance; and second, to gain a broader understanding of the function of LRR RLKs in root growth in general. We generated a collection of 135 homozygous T-DNA insertion mutants for 69 RLK genes that comprised around 30% of the LRR RLKs present in the Arabidopsis genome.
Assuming conservation of the CLV pathway in the root, we expected the kinase involved in root meristem homeostasis to be closely related to CLV1 and/or a member of the LRR RLK class. However, in our screen we did not obtain CLV1 paralogs involved in root development nor in CLE signalling. Recently, different types of receptors putatively involved in CLE signaling were implicated to play a role in regulating root growth. Mutations in CRN/SOL2, a membrane bound receptor kinase lacking an extracellular domain, and the CLV2 receptor-like protein can both prevent CLE induced consumption of the root meristem (Muller et al. 2008; Miwa et al. 2008). Other recent work assigned ACR4, a receptor of the Crinkly4 class, in controlling distal stem cell proliferation in the root meristem (De Smet et al. 2008). CLE40 is the putative ligand of ACR4 and together they regulate WOX5 expression, thus resembling the activity of the CLV3-CLV1-WUS shoot module (Stahl et al. 2009; Stahl and Simon 2009). Although these studies demonstrated recruitment of receptors other than the LRR class in controlling root meristem maintenance, they certainly do not exclude that LRR RLKs operate in this process.
The degree of specificity and redundancy among RLKs has been a matter of debate. Lack of identification of biological functions for RLK genes can be explained by functional redundancy that complicates studies employing reverse genetic strategies. Two emerging themes are that receptor kinases are part of a cellular network of regulatory proteins that includes physical interactions with other RLKs, and that multiple receptor kinases are involved in similar or overlapping processes. Double and triple mutants have been found that display phenotypes supporting this hypothesis, e.g. synergistic actions of ER, ERL1 and ERL2 controls organ growth and cell proliferation whereas BAM1, redundantly with BAM2 and BAM3, balances cell division and differentiation in the shoot meristem (Shpak et al. 2004a; Deyoung et al. 2006). In addition, receptors can potentially participate in different receptor complexes and this explains why some of these receptors play roles in diverse processes. ER is the best example as a pleiotropic regulator of developmental, physiological and processes as well as a modulator of responses to environmental stimuli (van Zanten et al. 2009). Strategies employing RNA interference to knockdown the expression of several RLKs simultaneously should help in overcoming functional redundancy among RLK genes. clv1 null alleles show a weak phenotype and all intermediate and strong alleles appeared dominant-negative most likely interfering with the signaling function (Dievart et al. 2003). Similar observations have been made for bak1 and har1 mutants (Dievart and Clark 2003). Generating dominant negative mutations for RLKs e.g. by removing their kinase domain could possibly lead to a better understanding of their function.
To gain a broader understanding of the function of LRR RLKs in root growth in general, we screened the T-DNA lines for altered response to environmental, hormonal/chemical and abiotic stress. Of the 69 mutant LRR RLKs tested 16 are involved in response to one type of treatment whereas 36 are involved in response to two or more types of treatment. 24 of the 69 LRR RLKs have been assigned a name of which many have been implicated in various biological programs. Three characterized LRR RLKs mutants were known to respond to the conditions tested in this study, and this was confirmed by us, demonstrating the validity of our screen: The elg mutant allele of BAK1/SERK3 was reported hypersensitive to IAA treatment (Whippo and Hangarter 2005); a T-DNA line for IRK was found to be more sensitive to IAA treatment. Although the function of IRK is elusive, its expression is increased by auxin treatment (Kanamoto et al. 2002). Seemingly contrasting our results, the vh1 mutant was previously reported hypersensitive to low concentrations of the synthetic auxin analog 2,4-dichlorophenoxyacetic acid (2,4-D; 25 and 50 mM), while responding as wild type at higher concentrations (Ceserani et al. 2009). The observed discrepancy may be due to the type of mutation or to the use of different auxin molecules.
In addition, we identified several novel conditional phenotypes linked with mutations in LRR RLK genes. T-DNA lines for BAK1/SERK3 showed an increased sensitivity for IAA and NPA treatment as well as an increased sensitivity to 6-BAP. These results are in line with the known interdependency of brassinosteroid (BR) and auxin signaling in Arabidopsis (Nemhauser et al. 2004). The antagonistic interaction between auxin and cytokinins is known but no relationship has been reported between BRs and cytokinin so far. Our studies provide a link for crosstalk between these three pathways.
In this study, novel phenotypes were found for 52 RLK knockouts. 21 of these concerned RLKs with previously characterized phenotypes but 31 provide functions for hitherto uncharacterized RLKs. We could not detect a clear relationship between conditional phenotypes and phylogeny. This suggests that these transmembrane receptor kinases, despite a fairly similar domain organization, can readily acquire different functions compared to their closest paralogs during evolution. We showed that there are LRR RLK gene clusters with similar behavior at the transcriptome level upon different stress treatments. However, RLK clusters did not correlate with the functional characterization of the mutants. Similarly, a large scale analysis of the transcriptional response of the 604-member RLK gene family to a range of known environmental and developmental stimuli demonstrated a broad response of these kinases to multiple treatments (Chae et al. 2009). Our observations that many T-DNA insertion lines respond to more than one treatment supports the existence of extensive cross talk and signal integration among different signaling pathways. With respect to hormones, for which receptors are identified, resistance or sensitivity may indicate a function of receptor signaling in secondary signaling events. Our study represents a preliminary view of processes in which the studied kinases may be involved. Additionally, the generated collection of LRR RLK T-DNA insertion mutants can be easily applied for the analysis of other developmental aspects, function in defense and additional stress conditions and thus forms a valuable resource for future investigations into the biological role of LRR RLKs.
References
Aker J, de Vries SC (2008) Plasma membrane receptor complexes. Plant Physiol 147:1560–1564
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, Gadrinab C, Heller C, Jeske A, Koesema E, Meyers CC, Parker H, Prednis L, Ansari Y, Choy N, Deen H, Geralt M, Hazari N, Hom E, Karnes M, Mulholland C, Ndubaku R, Schmidt I, Guzman P, Guilar-Henonin L, Schmid M, Weigel D, Carter DE, Marchand T, Risseeuw E, Brogden D, Zeko A, Crosby WL, Berry CC, Ecker JR (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Badescu GO, Napier RM (2006) Receptors for auxin: will it all end in TIRs? Trends Plant Sci 11:217–223
Birnbaum K, Shasha DE, Wang JY, Jung JW, Lambert GM, Galbraith DW, Benfey PN (2003) A gene expression map of the Arabidopsis root. Science 302:1956–1960
Bishopp A, Mahonen AP, Helariutta Y (2006) Signs of change: hormone receptors that regulate plant development. Development 133:1857–1869
Brady SM, Orlando DA, Lee JY, Wang JY, Koch J, Dinneny JR, Mace D, Ohler U, Benfey PN (2007) A high-resolution root spatiotemporal map reveals dominant expression patterns. Science 318:801–806
Ceserani T, Trofka A, Gandotra N, Nelson T (2009) VH1/BRL2 receptor-like kinase interacts with vascular-specific adaptor proteins VIT and VIK to influence leaf venation. Plant J 57:1000–1014
Chae L, Sudat S, Dudoit S, Zhu T, Luan S (2009) Diverse transcriptional programs associated with environmental stress and hormones in the Arabidopsis receptor-like kinase gene family. Mol Plant 2:84–107
de Lorenzo L, Merchan F, Laporte P, Thompson R, Clarke J, Sousa C, Crespi M (2009) A novel plant leucine-rich repeat receptor kinase regulates the response of Medicago truncatula roots to salt stress. Plant Cell 21:668–680
De Smet I, Vassileva V, De RB, Levesque MP, Grunewald W, Van DD, Van NG, Naudts M, Van IG, De CR, Wang JY, Meuli N, Vanneste S, Friml J, Hilson P, Jurgens G, Ingram GC, Inze D, Benfey PN, Beeckman T (2008) Receptor-like kinase ACR4 restricts formative cell divisions in the Arabidopsis root. Science 322:594–597
Dello Ioio R, Linhares FS, Scacchi E, Casamitjana-Martinez E, Heidstra R, Costantino P, Sabatini S (2007) Cytokinins determine Arabidopsis root-meristem size by controlling cell differentiation. Curr Biol 17:678–682
Dello Ioio R, Linhares FS, Sabatini S (2008) Emerging role of cytokinin as a regulator of cellular differentiation. Curr Opin Plant Biol 11:23–27
Deyoung BJ, Bickle KL, Schrage KJ, Muskett P, Patel K, Clark SE (2006) The CLAVATA1-related BAM1, BAM2 and BAM3 receptor kinase-like proteins are required for meristem function in Arabidopsis. Plant J 45:1–16
Dharmasiri N, Dharmasiri S, Weijers D, Lechner E, Yamada M, Hobbie L, Ehrismann JS, Jurgens G, Estelle M (2005) Plant development is regulated by a family of auxin receptor F box proteins. Dev Cell 9:109–119
Dievart A, Clark SE (2003) Using mutant alleles to determine the structure and function of leucine-rich repeat receptor-like kinases. Curr Opin Plant Biol 6:507–516
Dievart A, Clark SE (2004) LRR-containing receptors regulating plant development and defense. Development 131:251–261
Dievart A, Dalal M, Tax FE, Lacey AD, Huttly A, Li J, Clark SE (2003) CLAVATA1 dominant-negative alleles reveal functional overlap between multiple receptor kinases that regulate meristem and organ development. Plant Cell 15:1198–1211
Fujita H, Syono K (1997) PIS1, a negative regulator of the action of auxin transport inhibitors in Arabidopsis thaliana. Plant J 12:583–595
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Ann Rev Plant Physiol Plant Mol Biol 51:463–499
Ito Y, Nakanomyo I, Motose H, Iwamoto K, Sawa S, Dohmae N, Fukuda H (2006) Dodeca-CLE peptides as suppressors of plant stem cell differentiation. Science 313:842–845
Iyer-Pascuzzi A, Simpson J, Herrera-Estrella L, Benfey PN (2009) Functional genomics of root growth and development in Arabidopsis. Curr Opin Plant Biol 12:165–171
Johnson KL, Ingram GC (2005) Sending the right signals: regulating receptor kinase activity. Curr Opin Plant Biol 8:648–656
Kanamoto H, Hattan J, Takemura M, Yokota A, Kohchi T (2002) Molecular cloning and characterization of a gene coding for a putative receptor-like protein kinase with a Leucine-rich repeat expressed in inflorescence and root apices from Arabidopsis. Plant Biotech 19:113–120
Kinoshita A, Nakamura Y, Sasaki E, Kyozuka J, Fukuda H, Sawa S (2007) Gain-of-function phenotypes of chemically synthetic CLAVATA3/ESR-related (CLE) peptides in Arabidopsis thaliana and Oryza sativa. Plant Cell Physiol 48:1821–1825
Miwa H, Betsuyaku S, Iwamoto K, Kinoshita A, Fukuda H, Sawa S (2008) The receptor-like kinase SOL2 mediates CLE signaling in Arabidopsis. Plant Cell Physiol 49:1752–1757
Morillo SA, Tax FE (2006) Functional analysis of receptor-like kinases in monocots and dicots. Curr Opin in Plant Biol 9:460–469
Muller R, Bleckmann A, Simon R (2008) The receptor kinase CORYNE of Arabidopsis transmits the stem cell-limiting signal CLAVATA3 independently of CLAVATA1. Plant Cell 20:934–946
Nemhauser JL, Mockler TC, Chory J (2004) Interdependency of brassinosteroid and auxin signaling in Arabidopsis. PLoS Biol 2:e258
Ogawa M, Shinohara H, Sakagami Y, Matsubayashi Y (2008) Arabidopsis CLV3 peptide directly binds CLV1 ectodomain. Science 319:294
Penfield S, King J (2009) Towards a systems biology approach to understanding seed dormancy and germination. Proc R Soc B Biol Sci 276:3561–3569
Petrášek J, Friml J (2009) Auxin transport routes in plant development. Development 136:2675–2688
Rieu I, Laux T (2009) Signaling pathways maintaining stem cells at the plant shoot apex. Sem Cell Dev Biol 20:1083–1088
Rosso MG, Li Y, Strizhov N, Reiss B, Dekker K, Weisshaar B (2003) An Arabidopsis thaliana T-DNA mutagenized population (GABI-Kat) for flanking sequence tag-based reverse genetics. Plant Mol Biol 53:247–259
Sablowski R (2007) Flowering and determinacy in Arabidopsis. J Exp Bot 58:899–907
Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, Braisted J, Klapa M, Currier T, Thiagarajan M, Sturn A, Snuffin M, Rezantsev A, Popov D, Ryltsov A, Kostukovich E, Borisovsky I, Liu Z, Vinsavich A, Trush V, Quackenbush J (2003) TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34:374–378
Sarkar AK, Luijten M, Miyashima S, Lenhard M, Hashimoto T, Nakajima K, Scheres B, Heidstra R, Laux T (2007) Conserved factors regulate signalling in Arabidopsis thaliana shoot and root stem cell organizers. Nature 446:811–814
Scheres B (2007) Stem-cell niches: nursery rhymes across kingdoms. Nat Rev Mol Cell Biol 8:345–354
Sessions A, Burke E, Presting G, Aux G, McElver J, Patton D, Dietrich B, Ho P, Bacwaden J, Ko C, Clarke JD, Cotton D, Bullis D, Snell J, Miguel T, Hutchison D, Kimmerly B, Mitzel T, Katagiri F, Glazebrook J, Law M, Goff SA (2002) A high-throughput Arabidopsis reverse genetics system. Plant Cell 14:2985–2994
Shiu SH, Bleecker AB (2001a) Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc Natl Acad Sci USA 98:10763–10768
Shiu SH, Bleecker AB (2001b) Plant receptor-like kinase gene family: diversity, function, and signaling. Sci STKE 2001:RE22
Shiu SH, Bleecker AB (2003) Expansion of the receptor-like kinase/Pelle gene family and receptor-like proteins in Arabidopsis. Plant Physiol 132:530–543
Shpak ED, Berthiaume CT, Hill EJ, Torii KU (2004a) Synergistic interaction of three ERECTA-family receptor-like kinases controls Arabidopsis organ growth and flower development by promoting cell proliferation. Development 131:1491–1501
Shpak ED, Berthiaume CT, Hill EJ, Torii KU (2004b) Synergistic interaction of three ERECTA-family receptor-like kinases controls Arabidopsis organ growth and flower development by promoting cell proliferation. Development 131:1491–1501
Stahl Y, Simon R (2009) Is the Arabidopsis root niche protected by sequestration of the CLE40 signal by its putative receptor ACR4? Plant Signal Behav 4:634–635
Stahl Y, Wink RH, Ingram GC, Simon R (2009) A signaling module controlling the stem cell niche in Arabidopsis root meristems. Curr Biol 19:909–914
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tarutani Y, Morimoto T, Sasaki A, Yasuda M, Nakashita H, Yoshida S, Yamaguchi I, Suzuki Y (2004) Molecular characterization of two highly homologous receptor-like kinase genes, RLK902 and RKL1, in Arabidopsis thaliana. Biosci Biotechnol Biochem 68:1935–1941
Ten Hove CA, Heidstra R (2008) Who begets whom? Plant cell fate determination by asymmetric cell division. Curr Opin Plant Biol 11:34–41
Tromas A, Braun N, Muller P, Khodus T, Paponov IA, Palme K, Ljung K, Lee JY, Benfey P, Murray JA, Scheres B, Perrot-Rechenmann C (2009) The AUXIN BINDING PROTEIN 1 is required for differential auxin responses mediating root growth. PLoS One 4:e6648
van Zanten M, Snoek LB, Proveniers MCG, Peeters AJM (2009) The many functions of ERECTA. Trends Plant Sci 14:214–218
Vert Gg, Nemhauser JL, Geldner N, Hong F, Chory J (2005) Molecular mechanisms of steroid hormone signaling in plants. Annual Rev of Cell and Dev Biol 21:177–201
Wang H, Chevalier D, Larue C, Ki CS, Walker JC (2007) The protein phosphatases and protein kinases of Arabidopsis thaliana. In ‘the Arabidopsis book’. The American Society of Plant Biologists, pp 1–38
Whippo CW, Hangarter RP (2005) A brassinosteroid-hypersensitive mutant of BAK1 indicates that a convergence of photomorphogenic and hormonal signaling modulates phototropism. Plant Physiol 139:448–457
Whitford R, Fernandez A, De GR, Ortega E, Hilson P (2008) Plant CLE peptides from two distinct functional classes synergistically induce division of vascular cells. Proc Natl Acad Sci USA 105:18625–18630
Willemsen V, Wolkenfelt H, de Vrieze G, Weisbeek P, Scheres B (1998) The HOBBIT gene is required for formation of the root meristem in the Arabidopsis embryo. Development 125:521–531
Wolters H, Jurgens G (2009) Survival of the flexible: hormonal growth control and adaptation in plant development. Nat Rev Genet 10:305–317
Xiong L, Zhu JK (2001) Abiotic stress signal transduction in plants: molecular and genetic perspectives. Physiol Plant 112:152–166
Xiong L, Zhu J (2002) Salt tolerance. In ‘The Arabidopsis Book’. The American Society of Plant Biologists, pp 1–22
Acknowledgments
We are grateful to ABRC, NASC, Syngenta, Dr. Yoshihito Suzuki, Dr. Steven Clark and Dr. Keiko Torii for providing plant material and to Hiroo Fukuda (University of Tokyo, Japan) for providing CLV3 peptide. CAtH and VMAJ were funded by Centre for BioSystems Genomics (CBSG) and Horizon grants to RH, respectively, GFSP was funded by the Netherlands Consortium for Systems Biology (NCSB), originating from the Netherlands Genomics Initiative/Netherlands Organization for Scientific Research (NGI/NWO).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
ten Hove, C.A., Bochdanovits, Z., Jansweijer, V.M.A. et al. Probing the roles of LRR RLK genes in Arabidopsis thaliana roots using a custom T-DNA insertion set. Plant Mol Biol 76, 69–83 (2011). https://doi.org/10.1007/s11103-011-9769-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-011-9769-x