Abstract
Purpose
Hormonal stimulation prior to IVF influences the ovarian environment and therefore impacts oocytes and subsequent embryo quality. Not every patient has the same response to the same treatment and many fail for unknown reasons. Knowing why a cycle has failed and how the follicles were affected would allow clinicians to adapt the treatment accordingly and improve success rate. This study examines the hypothesis that transcriptomic analysis of follicular cells from failed IVF cycles reveals potential reasons for failure and provides new information on the physiological mechanisms related to IVF failure.
Methods
Follicular cells (granulosa cells) were obtained from IVF patients of four Canadian fertility clinics. Using microarray analysis, patients that did not become pregnant following the IVF cycle were compared to those that did. Functional analysis was performed using ingenuity pathway analysis and qRT-PCR was used to validate the microarray results in a larger cohort of patients.
Results
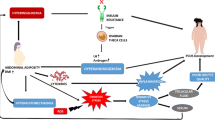
The microarray showed 165 differentially expressed genes (DEGs) in the negative group compared to the pregnancy group. DEGs include many pro-inflammatory cytokines and other factors related to inflammation, suggesting that this process might be altered when IVF fails. Overexpression of several factors, some of which act upstream from vascular endothelial growth factor (VEGF), also indicates increased permeability and vasodilation. Some DEGs were related to abnormal differentiation and increased apoptosis.
Conclusions
Our results suggest that failure to conceive following IVF cycles could be associated with an imbalance between pro-inflammatory and anti-inflammatory mediators. The findings of this study identify potential failure causes and pathways for further investigation. Stimulatory protocols personalized according to patient response could improve the chances of later success.


Similar content being viewed by others
References
Canadian Fertility and Andrology Society. Canadian ART Report. Human Assisted Reproduction 2014 Live Birth Rates for Canada 2014.
The European IVF-Monitoring Consortium for the European Society of Human Reproduction and Embryology, Calhaz-Jorge C, de Geyter C, Kupka MS, de Mouzon J, Erb K, et al. Assisted reproductive technology in Europe, 2012: results generated from European registers by ESHRE. Hum Reprod. 2016;31(8):1638–52. https://doi.org/10.1093/humrep/dew151.
Eppig JJ. Oocyte control of ovarian follicular development and function in mammals. Reproduction. 2001;122(6):829–38.
Matzuk MM, Burns KH, Viveiros MM, Eppig JJ. Intercellular communication in the mammalian ovary: oocytes carry the conversation. Science. 2002;296(5576):2178–80. https://doi.org/10.1126/science.1071965.
Hamel M, Dufort I, Robert C, Gravel C, Leveille M-C, Leader A, et al. Identification of differentially expressed markers in human follicular cells associated with competent oocytes. Hum Reprod. 2008;23(5):1118–27. https://doi.org/10.1093/humrep/den048.
Hamel M, Dufort I, Robert C, Léveillé M-C, Leader A, Sirard M-A. Genomic assessment of follicular marker genes as pregnancy predictors for human IVF. Mol Hum Reprod. 2010;16(2):87–96. https://doi.org/10.1093/molehr/gap079.
Hamel M, Dufort I, Robert C, Léveillé M-C, Leader A, Sirard M-A. Identification of follicular marker genes as pregnancy predictors for human IVF: new evidence for the involvement of luteinization process. Mol Hum Reprod. 2010;16(8):548–56. https://doi.org/10.1093/molehr/gaq051.
Uyar A, Torrealday S, Seli E. Cumulus and granulosa cell markers of oocyte and embryo quality. Fertil Steril. 2013;99(4):979–97. https://doi.org/10.1016/j.fertnstert.2013.01.129.
Burnik Papler T, Vrtacnik Bokal E, Maver A, Lovrecic L. Specific gene expression differences in cumulus cells as potential biomarkers of pregnancy. Reprod BioMed Online. 2015;30:426–33. https://doi.org/10.1016/j.rbmo.2014.12.011.
Gebhardt KM, Feil DK, Dunning KR, Lane M, Russell DL. Human cumulus cell gene expression as a biomarker of pregnancy outcome after single embryo transfer. Fertil Steril. 2011;96(1):47–52 e2. https://doi.org/10.1016/j.fertnstert.2011.04.033.
Iager AE, Kocabas AM, Otu HH, Ruppel P, Langerveld A, Schnarr P, et al. Identification of a novel gene set in human cumulus cells predictive of an oocyte’s pregnancy potential. Fertil Steril. 2013;99(3):745–52 e6. https://doi.org/10.1016/j.fertnstert.2012.10.041.
Wathlet S, Adriaenssens T, Segers I, Verheyen G, Van Landuyt L, Coucke W, et al. Pregnancy prediction in single embryo transfer cycles after ICSI using QPCR: validation in oocytes from the same cohort. PLoS One. 2013;8(4):e54226. https://doi.org/10.1371/journal.pone.0054226.
Anderson RA, Sciorio R, Kinnell H, Bayne RA, Thong KJ, de Sousa PA, et al. Cumulus gene expression as a predictor of human oocyte fertilisation, embryo development and competence to establish a pregnancy. Reproduction. 2009;138(4):629–37. https://doi.org/10.1530/REP-09-0144.
Assidi M, Montag M, Van der Ven K, Sirard MA. Biomarkers of human oocyte developmental competence expressed in cumulus cells before ICSI: a preliminary study. J Assist Reprod Genet. 2011;28(2):173–88. https://doi.org/10.1007/s10815-010-9491-7.
Assou S, Haouzi D, Mahmoud K, Aouacheria A, Guillemin Y, Pantesco V, et al. A non-invasive test for assessing embryo potential by gene expression profiles of human cumulus cells: a proof of concept study. Mol Hum Reprod. 2008;14(12):711–9. https://doi.org/10.1093/molehr/gan067.
Feuerstein P, Puard V, Chevalier C, Teusan R, Cadoret V, Guerif F, et al. Genomic assessment of human cumulus cell marker genes as predictors of oocyte developmental competence: impact of various experimental factors. PLoS One. 2012;7(7):e40449. https://doi.org/10.1371/journal.pone.0040449.
Nivet AL, Leveille MC, Leader A, Sirard MA. Transcriptional characteristics of different sized follicles in relation to embryo transferability: potential role of hepatocyte growth factor signalling. Mol Hum Reprod. 2016;22:475–84. https://doi.org/10.1093/molehr/gaw029.
Blazejczyk M, Miron M, Nadon R. FlexArray: a statistical data analysis software for gene expression microarrays. Montreal, Canada: Genome Quebec 2007.
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002;3(7):RESEARCH0034.
Espey LL. Ovulation as an inflammatory reaction—a hypothesis. Biol Reprod. 1980;22(1):73–106.
Espey LL, Lipner H. Ovulation. In: Knobil E, Neill JD, editors. The physiology of reproduction. 1 ed. New-York, United-States.: Raven Press; 1994. p. 725–80.
Espey LL, Bellinger AS, Healy JA. Ovulation: an inflammatory cascade of gene expression. In: Leung PCK, Adashi EY, editors. The ovary. 2 ed. San Diego, CA, United States: Elsevier Academic Press; 2004. p. 145–165.
Adams J, Liu Z, Ren YA, Wun WS, Zhou W, Kenigsberg S, et al. Enhanced inflammatory transcriptome in the granulosa cells of women with polycystic ovarian syndrome. J Clin Endocrinol Metab. 2016;101(9):3459–68. https://doi.org/10.1210/jc.2015-4275.
Dahm-Kahler P, Runesson E, Lind AK, Brannstrom M. Monocyte chemotactic protein-1 in the follicle of the menstrual and IVF cycle. Mol Hum Reprod. 2006;12(1):1–6. https://doi.org/10.1093/molehr/gah256.
Carletti MZ, Christenson LK. Rapid effects of LH on gene expression in the mural granulosa cells of mouse periovulatory follicles. Reproduction. 2009;137(5):843–55. https://doi.org/10.1530/REP-08-0457.
Kaur S, Archer KJ, Devi MG, Kriplani A, Strauss JF 3rd, Singh R. Differential gene expression in granulosa cells from polycystic ovary syndrome patients with and without insulin resistance: identification of susceptibility gene sets through network analysis. J Clin Endocrinol Metab. 2012;97(10):E2016–21. https://doi.org/10.1210/jc.2011-3441.
Yan SF, Fujita T, Lu J, Okada K, Shan Zou Y, Mackman N, et al. Egr-1, a master switch coordinating upregulation of divergent gene families underlying ischemic stress. Nat Med. 2000;6(12):1355–61. https://doi.org/10.1038/82168.
Beckmann AM, Wilce PA. Egr transcription factors in the nervous system. Neurochem Int. 1997;31(4):477–510 discussion 7-6.
Khachigian LM, Collins T. Early growth response factor 1: a pleiotropic mediator of inducible gene expression. J Mol Med (Berl). 1998;76(9):613–6.
Cavaillon JM. Cytokines in inflammation. C R Seances Soc Biol Fil. 1995;189(4):531–44.
Okada M, Fujita T, Sakaguchi T, Olson KE, Collins T, Stern DM, et al. Extinguishing Egr-1-dependent inflammatory and thrombotic cascades after lung transplantation. FASEB J. 2001;15(14):2757–9. https://doi.org/10.1096/fj.01-0490fje.
Silverman ES, De Sanctis GT, Boyce J, Maclean JA, Jiao A, Green FH, et al. The transcription factor early growth-response factor 1 modulates tumor necrosis factor-alpha, immunoglobulin E, and airway responsiveness in mice. Am J Respir Crit Care Med. 2001;163(3 Pt 1):778–85. https://doi.org/10.1164/ajrccm.163.3.2003123.
Brannstrom M. Potential role of cytokines in ovarian physiology: the case for interleukin-1. In: Leung PCK, Adashi EY, editors. The ovary. 2 ed. San Diego, CA, United States: Elsevier Academic Press; 2004. p. 261–271.
Guirao X, Lowry SF. Biologic control of injury and inflammation: much more than too little or too late. World J Surg. 1996;20(4):437–46.
Cassatella MA, Meda L, Bonora S, Ceska M, Constantin G. Interleukin 10 (IL-10) inhibits the release of proinflammatory cytokines from human polymorphonuclear leukocytes. Evidence for an autocrine role of tumor necrosis factor and IL-1 beta in mediating the production of IL-8 triggered by lipopolysaccharide. J Exp Med. 1993;178(6):2207–11.
Medzhitov R. Inflammation 2010: new adventures of an old flame. Cell. 2010;140(6):771–6. https://doi.org/10.1016/j.cell.2010.03.006.
Muszynski JA, Frazier WJ, Hall MW. Pro-inflammatory and anti-inflammatory mediators in critical illness and injury. In: Wheeler SD, Wong RH, Shanley PT, editors. Pediatric critical care medicine: volume 1: care of the critically ill or injured child. London: Springer London; 2014. p. 231–8.
Haslett C. Granulocyte apoptosis and inflammatory disease. Br Med Bull. 1997;53(3):669–83.
Huynh ML, Fadok VA, Henson PM. Phosphatidylserine-dependent ingestion of apoptotic cells promotes TGF-beta1 secretion and the resolution of inflammation. J Clin Invest. 2002;109(1):41–50. https://doi.org/10.1172/JCI11638.
Voll RE, Herrmann M, Roth EA, Stach C, Kalden JR, Girkontaite I. Immunosuppressive effects of apoptotic cells. Nature. 1997;390(6658):350–1. https://doi.org/10.1038/37022.
McGrath EE, Marriott HM, Lawrie A, Francis SE, Sabroe I, Renshaw SA, et al. TNF-related apoptosis-inducing ligand (TRAIL) regulates inflammatory neutrophil apoptosis and enhances resolution of inflammation. J Leukoc Biol. 2011;90(5):855–65. https://doi.org/10.1189/jlb.0211062.
Renshaw SA, Parmar JS, Singleton V, Rowe SJ, Dockrell DH, Dower SK, et al. Acceleration of human neutrophil apoptosis by TRAIL. J Immunol. 2003;170(2):1027–33.
Bergh PA, Navot D. Ovarian hyperstimulation syndrome: a review of pathophysiology. J Assist Reprod Genet. 1992;9(5):429–38.
McClure N, Healy DL, Rogers PA, Sullivan J, Beaton L, Haning RV Jr, et al. Vascular endothelial growth factor as capillary permeability agent in ovarian hyperstimulation syndrome. Lancet. 1994;344(8917):235–6.
Soares SR, Gomez R, Simon C, Garcia-Velasco JA, Pellicer A. Targeting the vascular endothelial growth factor system to prevent ovarian hyperstimulation syndrome. Hum Reprod Update. 2008;14(4):321–33. https://doi.org/10.1093/humupd/dmn008.
Vlahos NF, Gregoriou O. Prevention and management of ovarian hyperstimulation syndrome. Ann N Y Acad Sci. 2006;1092:247–64. https://doi.org/10.1196/annals.1365.021.
Lee JW, Bae SH, Jeong JW, Kim SH, Kim KW. Hypoxia-inducible factor (HIF-1)alpha: its protein stability and biological functions. Exp Mol Med. 2004;36(1):1–12. https://doi.org/10.1038/emm.2004.1.
Kim W, Moon SO, Sung MJ, Kim SH, Lee S, So JN, et al. Angiogenic role of adrenomedullin through activation of Akt, mitogen-activated protein kinase, and focal adhesion kinase in endothelial cells. FASEB J. 2003;17(13):1937–9. https://doi.org/10.1096/fj.02-1209fje.
Sugo S, Minamino N, Shoji H, Kangawa K, Kitamura K, Eto T, et al. Interleukin-1, tumor necrosis factor and lipopolysaccharide additively stimulate production of adrenomedullin in vascular smooth muscle cells. Biochem Biophys Res Commun. 1995;207(1):25–32. https://doi.org/10.1006/bbrc.1995.1148.
Ando M, Kol S, Kokia E, Ruutiainen-Altman K, Sirois J, Rohan RM, et al. Rat ovarian prostaglandin endoperoxide synthase-1 and -2: periovulatory expression of granulosa cell-based interleukin-1-dependent enzymes. Endocrinology. 1998;139(5):2501–8. https://doi.org/10.1210/endo.139.5.5988.
Levitas E, Chamoun D, Udoff LC, Ando M, Resnick CE, Adashi EY. Periovulatory and interleukin-1 beta-dependent up-regulation of intraovarian vascular endothelial growth factor (VEGF) in the rat: potential role for VEGF in the promotion of periovulatory angiogenesis and vascular permeability. J Soc Gynecol Investig. 2000;7(1):51–60.
Smith-Mungo LI, Kagan HM. Lysyl oxidase: properties, regulation and multiple functions in biology. Matrix Biol. 1998;16(7):387–98.
Baker AM, Bird D, Welti JC, Gourlaouen M, Lang G, Murray GI, et al. Lysyl oxidase plays a critical role in endothelial cell stimulation to drive tumor angiogenesis. Cancer Res. 2013;73(2):583–94. https://doi.org/10.1158/0008-5472.CAN-12-2447.
Mammoto A, Mammoto T, Kanapathipillai M, Wing Yung C, Jiang E, Jiang A, et al. Control of lung vascular permeability and endotoxin-induced pulmonary oedema by changes in extracellular matrix mechanics. Nat Commun. 2013;4:1759. https://doi.org/10.1038/ncomms2774.
Vloeberghs V, Peeraer K, Pexsters A, D'Hooghe T. Ovarian hyperstimulation syndrome and complications of ART. Best Pract Res Clin Obstet Gynaecol. 2009;23(5):691–709. https://doi.org/10.1016/j.bpobgyn.2009.02.006.
Bevilacqua MP, Pober JS, Wheeler ME, Cotran RS, Gimbrone MA Jr. Interleukin-1 activation of vascular endothelium. Effects on procoagulant activity and leukocyte adhesion. Am J Pathol. 1985;121(3):394–403.
Bevilacqua MP, Pober JS, Wheeler ME, Cotran RS, Gimbrone MA Jr. Interleukin 1 acts on cultured human vascular endothelium to increase the adhesion of polymorphonuclear leukocytes, monocytes, and related leukocyte cell lines. J Clin Invest. 1985;76(5):2003–11. https://doi.org/10.1172/JCI112200.
Ley K, Laudanna C, Cybulsky MI, Nourshargh S. Getting to the site of inflammation: the leukocyte adhesion cascade updated. Nat Rev Immunol. 2007;7(9):678–89. https://doi.org/10.1038/nri2156.
Nourshargh S, Alon R. Leukocyte migration into inflamed tissues. Immunity. 2014;41(5):694–707. https://doi.org/10.1016/j.immuni.2014.10.008.
Rizk B, Aboulghar M, Smitz J, Ron-El R. The role of vascular endothelial growth factor and interleukins in the pathogenesis of severe ovarian hyperstimulation syndrome. Hum Reprod Update. 1997;3(3):255–66.
Wong GG, Clark SC. Multiple actions of interleukin 6 within a cytokine network. Immunol Today. 1988;9(5):137–9. https://doi.org/10.1016/0167-5699(88)91200-5.
Hammond ME, Lapointe GR, Feucht PH, Hilt S, Gallegos CA, Gordon CA, et al. IL-8 induces neutrophil chemotaxis predominantly via type I IL-8 receptors. J Immunol. 1995;155(3):1428–33.
Brannstrom M, Pascoe V, Norman RJ, McClure N. Localization of leukocyte subsets in the follicle wall and in the corpus luteum throughout the human menstrual cycle. Fertil Steril. 1994;61(3):488–95.
Norman R, Bonello N, Jasper M, Van der Hoek K. Leukocytes: essential cells in ovarian function and ovulation. Reprod Med Rev. 1997;6(02):97–111. https://doi.org/10.1017/S0962279900001447.
Hock DL, Huhn RD, Kemmann E. Leukocytosis in response to exogenous gonadotrophin stimulation. Hum Reprod. 1997;12(10):2143–6.
Fabregues F, Balasch J, Manau D, Jimenez W, Arroyo V, Creus M, et al. Haematocrit, leukocyte and platelet counts and the severity of the ovarian hyperstimulation syndrome. Hum Reprod. 1998;13(9):2406–10.
Loret de Mola JR, Baumgardner GP, Goldfarb JM, Friedlander MA. Ovarian hyperstimulation syndrome: pre-ovulatory serum concentrations of interleukin-6, interleukin-1 receptor antagonist and tumour necrosis factor-alpha cannot predict its occurrence. Hum Reprod. 1996;11(7):1377–80.
Bertout JA, Mahutte NG, Preston SL, Behrman HR. Reactive oxygen species and ovarian function. In: Leung PCK, Adashi EY, editors. The ovary. 2 ed. San Diego, CA, United States: Elsevier Academic Press; 2004. p. 353–368.
Davisson MT, Bechtel LJ, Akeson EC, Fortna A, Slavov D, Gardiner K. Evolutionary breakpoints on human chromosome 21. Genomics. 2001;78(1–2):99–106. https://doi.org/10.1006/geno.2001.6639.
Nishizaki R, Ota M, Inoko H, Meguro A, Shiota T, Okada E, et al. New susceptibility locus for high myopia is linked to the uromodulin-like 1 (UMODL1) gene region on chromosome 21q22.3. Eye (Lond). 2009;23(1):222–9. https://doi.org/10.1038/eye.2008.152.
Shibuya K, Nagamine K, Okui M, Ohsawa Y, Asakawa S, Minoshima S, et al. Initial characterization of an uromodulin-like 1 gene on human chromosome 21q22.3. Biochem Biophys Res Commun. 2004;319(4):1181–9. https://doi.org/10.1016/j.bbrc.2004.05.094.
Wang W, Tang Y, Ni L, Kim E, Jongwutiwes T, Hourvitz A, et al. Overexpression of uromodulin-like1 accelerates follicle depletion and subsequent ovarian degeneration. Cell Death Dis. 2012;3:e433. https://doi.org/10.1038/cddis.2012.169.
Rodriguez P, Vinuela JE, Alvarez-Fernandez L, Gomez-Marquez J. Prothymosin alpha antisense oligonucleotides induce apoptosis in HL-60 cells. Cell Death Differ. 1999;6(1):3–5. https://doi.org/10.1038/sj.cdd.4400450.
Jiang X, Kim HE, Shu H, Zhao Y, Zhang H, Kofron J, et al. Distinctive roles of PHAP proteins and prothymosin-alpha in a death regulatory pathway. Science. 2003;299(5604):223–6. https://doi.org/10.1126/science.1076807.
Qi X, Wang L, Du F. Novel small molecules relieve prothymosin alpha-mediated inhibition of apoptosome formation by blocking its interaction with Apaf-1. Biochemistry. 2010;49(9):1923–30. https://doi.org/10.1021/bi9022329.
Acknowledgements
The authors thank Isabelle Dufort for her support and assistance in the lab, and Scot Hamilton, Marie-Claude Léveillé, Martin Laforest, and Hanane Lachgar for their involvement in the collection of follicle cell samples.
Authors’ roles
Fortin, C.: conception and design of the study, sample processing, data acquisition, data analysis and interpretation, drafting the article, approval of the final manuscript.
Leader, A.: data acquisition, critical discussion, reading and approval of the final manuscript.
Mahutte, N.: data acquisition, critical discussion, reading and approval of the final manuscript.
Hamilton, S.: data acquisition, critical discussion, reading and approval of the final manuscript.
Léveillé, M.C.: data acquisition, critical discussion, reading and approval of the final manuscript.
Villeneuve, M.: data acquisition, critical discussion, reading and approval of the final manuscript.
Sirard, M.A.: conception and design of the study, data analysis and interpretation, improvement of the draft, reading and approval of the final manuscript.
Funding
This work was supported by the Merck Serono Grant for Fertility Innovation (GFI).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. A GFI grant was received in support of this work. This article does not contain any data or conclusion relating to any commercial product.
Ethical approval
This project was approved by Université Laval REB (2014-102/08-09-2014) for the collection and use of human tissues.
Large-scale data
The data discussed in this publication are accessible through GEO Series accession number GSE87545.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 42 kb)
Rights and permissions
About this article
Cite this article
Fortin, C., Leader, A., Mahutte, N. et al. Gene expression analysis of follicular cells revealed inflammation as a potential IVF failure cause. J Assist Reprod Genet 36, 1195–1210 (2019). https://doi.org/10.1007/s10815-019-01447-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-019-01447-4