Abstract
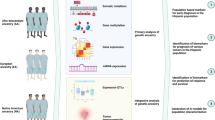
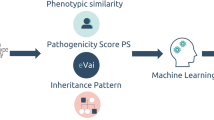
In 20–30% of patients suspected of a familial colorectal cancer (CRC) syndrome, no underlying genetic cause is detected. Recent advances in whole exome sequencing have generated evidence for new CRC-susceptibility genes including POLE, POLD1 and NTHL1¸ but many patients remain unexplained. Whole exome sequencing was performed on DNA from nine patients from five different families with familial clusters of CRC in which traditional genetic testing failed to yield a diagnosis. Variants were filtered by minor allele frequencies, followed by prioritization based on in silico prediction tools, and the presence in cancer susceptibility genes or genes in cancer-associated pathways. Effects of frameshift variants on protein structure were modeled using I-Tasser. One known pathogenic variant in POLD1 was detected (p.S478N), together with variants in 17 candidate genes not previously associated with CRC. Additional in silico analysis using SIFT, PROVEAN and PolyPhen on the 14 missense variants indicated a possible damaging effect in nine of 14 variants. Modeling of the insertions/deletions showed a damaging effect of two variants in NOTCH2 and CYP1B1. One family was explained by a mutation in a known familial CRC gene. In the remaining four families, the most promising candidates found are a frameshift NOTCH2 and a missense RAB25 variant. This study provides potential novel candidate variants in unexplained familial CRC patients, however, functional validation is imperative to confirm the role of these variants in CRC tumorigenesis. Additionally, while whole exome sequencing enables detection of variants throughout the exome, other causes explaining the familial phenotype such as multiple single nucleotide polymorphisms accumulating to a polygenic risk or epigenetic events, might be missed with this approach.




Similar content being viewed by others
References
Valle L (2017) Recent discoveries in the genetics of familial colorectal cancer and polyposis. Clin Gastroenterol Hepatol 15(6):809–819
Pearlman R, Frankel WL, Swanson B, Zhao W, Yilmaz A, Miller K, Bacher J, Bigley C, Nelsen L, Goodfellow PJ et al (2017) Prevalence and spectrum of germline cancer susceptibility gene mutations among patients with early-onset colorectal cancer. JAMA Oncol 3(4):464–471
Vasen HF (2005) Clinical description of the Lynch syndrome [hereditary nonpolyposis colorectal cancer (HNPCC)]. Fam Cancer 4(3):219–225
Boland CR, Goel A (2010) Microsatellite instability in colorectal cancer. Gastroenterology 138(6):2073–2087.e2073
Lindor NM, Rabe K, Petersen GM, Haile R, Casey G, Baron J, Gallinger S, Bapat B, Aronson M, Hopper J et al (2005) Lower cancer incidence in Amsterdam-I criteria families without mismatch repair deficiency: familial colorectal cancer type X. JAMA 293(16):1979–1985
Shiovitz S, Copeland WK, Passarelli MN, Burnett-Hartman AN, Grady WM, Potter JD, Gallinger S, Buchanan DD, Rosty C, Win AK et al (2014) Characterisation of familial colorectal cancer type X, Lynch syndrome, and non-familial colorectal cancer. Br J Cancer 111(3):598–602
FC Da Silva, Wernhoff P, Dominguez-Barrera C, Dominguez-Valentin M (2016) Update on hereditary colorectal cancer. Anticancer Res 36(9):4399–4405
Wei C, Peng B, Han Y, Chen WV, Rother J, Tomlinson GE, Boland CR, Chaussabel D, Frazier ML, Amos CI (2015) Mutations of HNRNPA0 and WIF1 predispose members of a large family to multiple cancers. Fam Cancer 14(2):297–306
Francisco I, Albuquerque C, Lage P, Belo H, Vitoriano I, Filipe B, Claro I, Ferreira S, Rodrigues P, Chaves P et al (2011) Familial colorectal cancer type X syndrome: two distinct molecular entities? Fam Cancer 10(4):623–631
Palles C, Cazier JB, Howarth KM, Domingo E, Jones AM, Broderick P, Kemp Z, Spain SL, Guarino E, Salguero I et al (2013) Germline mutations affecting the proofreading domains of POLE and POLD1 predispose to colorectal adenomas and carcinomas. Nat Genet 45(2):136–144
Briggs S, Tomlinson I (2013) Germline and somatic polymerase epsilon and delta mutations define a new class of hypermutated colorectal and endometrial cancers. J Pathol 230(2):148–153
Shinbrot E, Henninger EE, Weinhold N, Covington KR, Goksenin AY, Schultz N, Chao H, Doddapaneni H, Muzny DM, Gibbs RA et al (2014) Exonuclease mutations in DNA polymerase epsilon reveal replication strand specific mutation patterns and human origins of replication. Genome Res 24(11):1740–1750
Weren RD, Ligtenberg MJ, Kets CM, de Voer RM, Verwiel ET, Spruijt L, van Zelst-Stams WA, Jongmans MC, Gilissen C, Hehir-Kwa JY et al (2015) A germline homozygous mutation in the base-excision repair gene NTHL1 causes adenomatous polyposis and colorectal cancer. Nat Genet 47(6):668–671
Kuiper RP, Hoogerbrugge N (2015) NTHL1 defines novel cancer syndrome. Oncotarget 6(33):34069–34070
Stoffel EM, Koeppe E, Everett J, Ulintz P, Kiel M, Osborne J, Williams L, Hanson K, Gruber SB, Rozek LS (2018) Germline genetic features of young individuals with colorectal cancer. Gastroenterology 154(4):897–905.e891
Dominguez-Valentin M, Nakken S, Tubeuf H, Vodak D, Ekstrøm PO, Nissen AM, Morak M, Holinski-Feder E, Martins A, Møller P et al (2018) Identification of genetic variants for clinical management of familial colorectal tumors. BMC Med Genet 19:26
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38(16):e164–e164
Choi Y, Sims GE, Murphy S, Miller JR, Chan AP (2012) Predicting the functional effect of amino acid substitutions and indels. PLoS ONE 7(10):e46688
Dakal TC, Kala D, Dhiman G, Yadav V, Krokhotin A, Dokholyan NV (2017) Predicting the functional consequences of non-synonymous single nucleotide polymorphisms in IL8 gene. Sci Rep 7(1):6525
Ng PC, Henikoff S (2003) SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res 31(13):3812–3814
Ramensky V, Bork P, Sunyaev S (2002) Human non-synonymous SNPs: server and survey. Nucleic Acids Res 30(17):3894–3900
Ashkenazy H, Erez E, Martz E, Pupko T, Ben-Tal N (2010) ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res 38(Web Server issue):W529–W533
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5(4):725–738
Dakal TC, Kumar R, Ramotar D (2017) Structural modeling of human organic cation transporters. Comput Biol Chem 68:153–163
Simpson MA, Irving MD, Asilmaz E, Gray MJ, Dafou D, Elmslie FV, Mansour S, Holder SE, Brain CE, Burton BK et al (2011) Mutations in NOTCH2 cause Hajdu-Cheney syndrome, a disorder of severe and progressive bone loss. Nat Genet 43(4):303–305
Guo F-J, Zhang W-J, Li Y-L, Liu Y, Li Y-H, Huang J, Wang J-J, Xie P-L, Li G-C (2010) Expression and functional characterization of platelet-derived growth factor receptor-like gene. World J Gastroenterol: WJG 16(12):1465–1472
Fujiwara Y, Ohata H, Kuroki T, Koyama K, Tsuchiya E, Monden M, Nakamura Y (1995) Isolation of a candidate tumor suppressor gene on chromosome 8p21.3-p22 that is homologous to an extracellular domain of the PDGF receptor beta gene. Oncogene 10(5):891–895
Montagner S, Leoni C, Emming S, Chiara GD, Balestrieri C, Barozzi I, Piccolo V, Togher S, Ko M, Rao A et al (2016) TET2 Regulates mast cell differentiation and proliferation through catalytic and non-catalytic activities. Cell Rep 15(7):1566–1579
Nazha A, Meggendorfer M, Nadarajah N, Kneen KE, Radivoyevitch T, Przychodzen B, Makishima H, Patel BJ, Sanikommu SR, Hobson S et al (2015) TET2 alterations in myeloid malignancies, impact on clinical characteristics, outcome, and disease predisposition. Blood 126(23):1645–1645
Kanchi KL, Johnson KJ, Lu C, McLellan MD, Leiserson MDM, Wendl MC, Zhang Q, Koboldt DC, Xie M, Kandoth C et al (2014) Integrated analysis of germline and somatic variants in ovarian cancer. Nat Commun 5:3156
Huang Y, Wang G, Liang Z, Yang Y, Cui L, Liu C-Y (2016) Loss of nuclear localization of TET2 in colorectal cancer. Clin Epigenet 8:9
McDaniell R, Warthen DM, Sanchez-Lara PA, Pai A, Krantz ID, Piccoli DA, Spinner NB (2006) NOTCH2 mutations cause Alagille syndrome, a heterogeneous disorder of the notch signaling pathway. Am J Hum Genet 79(1):169–173
Wang NJ, Sanborn Z, Arnett KL, Bayston LJ, Liao W, Proby CM, Leigh IM, Collisson EA, Gordon PB, Jakkula L et al (2011) Loss-of-function mutations in Notch receptors in cutaneous and lung squamous cell carcinoma. Proc Natl Acad Sci 108(43):17761–17766
Goldenring JR, Nam KT (2011) Rab25 as a tumour suppressor in colon carcinogenesis. Br J Cancer 104(1):33–36
Nam KT, Lee HJ, Smith JJ, Lapierre LA, Kamath VP, Chen X, Aronow BJ, Yeatman TJ, Bhartur SG, Calhoun BC et al (2010) Loss of Rab25 promotes the development of intestinal neoplasia in mice and is associated with human colorectal adenocarcinomas. J Clin Investig 120(3):840–849
Amornphimoltham P, Rechache K, Thompson J, Masedunskas A, Leelahavanichkul K, Patel V, Molinolo A, Gutkind JS, Weigert R (2013) Rab25 regulates invasion and metastasis in head and neck cancer. Clin Cancer Res 19(6):1375–1388
Cheng JM, Volk L, Janaki DK, Vyakaranam S, Ran S, Rao KA (2010) Tumor suppressor function of Rab25 in triple-negative breast cancer. Int J Cancer 126(12):2799–2812
DeRycke MS, Gunawardena SR, Middha S, Asmann YW, Schaid DJ, McDonnell SK, Riska SM, Eckloff BW, Cunningham JM, Fridley BL et al (2013) Identification of novel variants in colorectal cancer families by high-throughput exome sequencing. Cancer Epidemiol Biomarkers Prev 22(7):1239–1251
Pearlman R, Frankel WL, Swanson B et al (2017) Prevalence and spectrum of germline cancer susceptibility gene mutations among patients with early-onset colorectal cancer. JAMA Oncol 3(4):464–471
Funding
The present work was supported by the CA72851, CA184792, CA187956 and CA202797 grants from the National Cancer Institute, National Institute of Health; grants from the Sammons Cancer Center and Baylor Foundation, as well as funds from the Baylor Scott & White Research Institute, Dallas, TX, USA to AG. This work was also supported by grant CA160911 from the NIH to PG.
Author information
Authors and Affiliations
Contributions
AMLJ: conceived and designed the study, collected the data, performed the analysis, drafting the manuscript, PG and TCD: Performed analysis, drafting the manuscript, critical reviewing of the manuscript, TPS: critical reviewing of the manuscript, CRB: critical reviewing of the manuscript, funding acquisition AG: Conceived and designed the study, critical reviewing of the manuscript, funding acquisition, supervision
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts to disclose.
Electronic Supplementary Material
Below is the link to the electronic supplementary material
Supplementary Figure 1:
Pedigree of family F. Squares represent males, circles represent females and diamonds is undisclosed gender. Phenotype is shown as tumor type followed by age of onset. Patients presented with colorectal cancer (CRC; fully filled symbol) or polyps (pol, right top corner). Arrows indicate patient sequenced with whole exome sequencing.*DNA available for co-segregation study. Supplementary Figure 2: Enriched terms of familial and sporadic CRC TCGA cohort.Enriched terms comparing the familial cohort (this study) with the sporadic CRC TCGA data are represented as scatterplots in a two dimensional space to summarize GO terms'semantic similarities (using REVIGO). Sematic space X and Y are used to be able to show distance between dots, but have no intrinsic meaning. The size of the dots indicates the frequency of the GO terms in the underlying GOA database. Distance of the dots from each other indicate the relatedness of the GO terms. Color of the dots indicates the p-value for false discovery rates. P-value is the enrichment p-value computed according to the mHG or HG model. This p-value is not corrected for multiple testing of 15413 GO terms. The most significant changes in the familiar cohort are seen in cell-cell adhesion pathways—Supplementary material 2 (PPTX 239 kb)
Rights and permissions
About this article
Cite this article
Jansen, A.M.L., Ghosh, P., Dakal, T.C. et al. Novel candidates in early-onset familial colorectal cancer. Familial Cancer 19, 1–10 (2020). https://doi.org/10.1007/s10689-019-00145-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10689-019-00145-5