Abstract
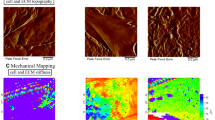
Cell contraction regulates how cells sense their mechanical environment. We sought to identify the set-point of cell contraction, also referred to as tensional homeostasis. In this work, bovine aortic endothelial cells (BAECs), cultured on substrates with different stiffness, were characterized using traction force microscopy (TFM). Numerical models were developed to provide insights into the mechanics of cell–substrate interactions. Cell contraction was modeled as eigenstrain which could induce isometric cell contraction without external forces. The predicted traction stresses matched well with TFM measurements. Furthermore, our numerical model provided cell stress and displacement maps for inspecting the fundamental regulating mechanism of cell mechanosensing. We showed that cell spread area, traction force on a substrate, as well as the average stress of a cell were increased in response to a stiffer substrate. However, the cell average strain, which is cell type-specific, was kept at the same level regardless of the substrate stiffness. This indicated that the cell average strain is the tensional homeostasis that each type of cell tries to maintain. Furthermore, cell contraction in terms of eigenstrain was found to be the same for both BAECs and fibroblast cells in different mechanical environments. This implied a potential mechanical set-point across different cell types. Our results suggest that additional measurements of contractility might be useful for monitoring cell mechanosensing as well as dynamic remodeling of the extracellular matrix (ECM). This work could help to advance the understanding of the cell-ECM relationship, leading to better regenerative strategies.
Similar content being viewed by others
References
Beloussov L, Louchinskaia N, Stein A (2000) Tension-dependent collective cell movements in the early gastrula ectoderm of Xenopus laevis embryos. Dev Genes Evol 210(2):92–104
Brown R et al (1998) Tensional homeostasis in dermal fibroblasts: mechanical responses to mechanical loading in three-dimensional substrates. J Cell Physiol 175(3):323–332
Butler JP et al (2002) Traction fields, moments, and strain energy that cells exert on their surroundings. Am J Physiol Cell Physiol 282(3):C595–C605
Califano JP, Reinhart-King CA (2008) A balance of substrate mechanics and matrix chemistry regulates endothelial cell network assembly. Cell Mol Bioeng 1(2–3):122–132
Califano JP, Reinhart-King CA (2010) Substrate stiffness and cell area predict cellular traction stresses in single cells and cells in contact. Cell Mol Bioeng 3(1):68–75
Chawla K (2012) Biomaterials for tissue engineering and regenerative medicine: treatment of musculoskeletal injury and disease. Mater Sci Eng A 557:45–53
Dembo M, Wang Y-L (1999) Stresses at the cell-to-substrate interface during locomotion of fibroblasts. Biophys J 76(4):2307–2316
Discher DE, Janmey P, Wang Y-L (2005) Tissue cells feel and respond to the stiffness of their substrate. Science 310(5751):1139–1143
Edwards CM, Schwarz US (2011) Force localization in contracting cell layers. Phys Rev Lett 107(12):128101
Engler AJ et al (2004) Surface probe measurements of the elasticity of sectioned tissue, thin gels and polyelectrolyte multilayer films: correlations between substrate stiffness and cell adhesion. Surf Sci 570(1):142–154
Franz CM, Müller DJ (2005) Analyzing focal adhesion structure by atomic force microscopy. J Cell Sci 118(22):5315–5323
Hadjipanayi E, Mudera V, Brown RA (2009) Guiding cell migration in 3D: a collagen matrix with graded directional stiffness. Cell Motil cytoskelet 66(3):121–128
Humphrey JD, Dufresne ER, Schwartz MA (2014) Mechanotransduction and extracellular matrix homeostasis. Nat Rev Mol Cell Biol 15(12):802–812
Jun T-S, Korsunsky AM (2010) Evaluation of residual stresses and strains using the eigenstrain reconstruction method. Int J Solids Struct 47(13):1678–1686
Kumar A et al (2016) Talin tension sensor reveals novel features of focal adhesion force transmission and mechanosensitivity. J Cell Biol 213(3):371–383
Lemmon CA, Romer LH (2010) A predictive model of cell traction forces based on cell geometry. Biophys J 99(9):L78–L80
Lin S et al (2017) Active stiffening of F-actin network dominated by structural transition of actin filaments into bundles. Compos Part B Eng 116:377–381
Marganski WA, Dembo M, Wang Y-L (2003) Measurements of cell-generated deformations on flexible substrata using correlation-based optical flow. Methods Enzymol 361:197–211
Milo R, Phillips R (2015) Cell biology by the numbers. Garland Science, New York
Mizutani T, Haga H, Kawabata K (2004) Cellular stiffness response to external deformation: tensional homeostasis in a single fibroblast. Cell Motil Cytoskelet 59(4):242–248
Munevar S, Wang Y-L, Dembo M (2001) Traction force microscopy of migrating normal and H-ras transformed 3T3 fibroblasts. Biophys J 80(4):1744–1757
Oakes PW et al (2014) Geometry regulates traction stresses in adherent cells. Biophys J 107(4):825–833
Oria R et al (2017) Force loading explains spatial sensing of ligands by cells. Nature 552(7684):219
Pan Y, Zhong Z (2016) Micromechanical modeling of the wood cell wall considering moisture absorption. Compos Part B Eng 91:27–35
Parker KK et al (2002) Directional control of lamellipodia extension by constraining cell shape and orienting cell tractional forces. FASEB J 16(10):1195–1204
Paszek MJ et al (2005) Tensional homeostasis and the malignant phenotype. Cancer Cell 8(3):241–254
Prager-Khoutorsky M et al (2011) Fibroblast polarization is a matrix-rigidity-dependent process controlled by focal adhesion mechanosensing. Nat Cell Biol 13(12):1457–1465
Rape AD, Guo W-H, Wang Y-L (2011) The regulation of traction force in relation to cell shape and focal adhesions. Biomaterials 32(8):2043–2051
Rittweger J et al (2009) Bone loss in the lower leg during 35 days of bed rest is predominantly from the cortical compartment. Bone 44(4):612–618
Solon J et al (2007) Fibroblast adaptation and stiffness matching to soft elastic substrates. Biophys J 93(12):4453–4461
Tan JL et al (2003) Cells lying on a bed of microneedles: an approach to isolate mechanical force. Proc Natl Acad Sci 100(4):1484–1489
Théry M et al (2006) Anisotropy of cell adhesive microenvironment governs cell internal organization and orientation of polarity. Proc Natl Acad Sci 103(52):19771–19776
van Oers RF et al (2014) Mechanical cell-matrix feedback explains pairwise and collective endothelial cell behavior in vitro. PLoS Comput Biol 10(8):e1003774
Vermolen F, Gefen A (2015) Semi-stochastic cell-level computational modelling of cellular forces: application to contractures in burns and cyclic loading. Biomech Model Mechanobiol 14(6):1181–1195
Wang Y-L, Pelham RJ (1998) Preparation of a flexible, porous polyacrylamide substrate for mechanical studies of cultured cells. Methods Enzymol 298:489–496
Wang N et al (2002) Micropatterning tractional forces in living cells. Cell Motil Cytoskelet 52(2):97–106
Yeung T et al (2005) Effects of substrate stiffness on cell morphology, cytoskeletal structure, and adhesion. Cell Motil Cytoskelet 60(1):24–34
Yu H, Mouw JK, Weaver VM (2011) Forcing form and function: biomechanical regulation of tumor evolution. Trends Cell Biol 21(1):47–56
Zhong SP et al (2007) Development of a novel collagen-GAG nanofibrous scaffold via electrospinning. Mater Sci Eng C 27(2):262–266
Zielinski R et al (2013) Finite element analysis of traction force microscopy: influence of cell mechanics, adhesion, and morphology. J Biomech Eng 135(7):071009
Acknowledgements
This work was supported in part by the National Science Foundation CAREER award (CBET-1254095) and the Foundation of Fujian Educational Committee (JAT170422). The authors also thank Mr. Hozhabr Mozafari for constructive comments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Lin, S., Lampi, M.C., Reinhart-King, C.A. et al. Eigenstrain as a mechanical set-point of cells. Biomech Model Mechanobiol 17, 951–959 (2018). https://doi.org/10.1007/s10237-018-1004-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10237-018-1004-0