Abstract
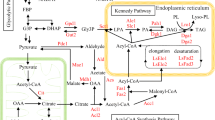
Long chain polyunsaturated fatty acids (LCPUFA) are known to play an important role in human health and nutrition. Considering the limitation of LCPUFA sources, it is necessary to search new avenues for their production. Oleaginous yeasts are an attractive target for harvesting single cell oil, mainly because of the ease of cultivation with cheaper raw material. Lipomyces starkeyi is one such oleaginous yeast, which can accumulate oil to the extent of 60 % of its biomass and where genetic transformation can be achieved. In our earlier work, Δ15 desaturase gene (AEP37840) from flax was transformed into L. starkeyi. In the present work, we report optimization of medium for the production of ω-3 enriched oil from this transformed yeast. A basic medium containing 20 g/l glucose as a carbon source and 10 g/l yeast extract as a nitrogen source was used during fermentation. At regular time intervals, glucose was fed to maintain high C:N ratio (65:10) during fermentation. Under the most favorable conditions, dry biomass and total lipid content were 18 and 7.29 g/l, respectively. Prior to genetic transformation, L. starkeyi contained 56.03 mg/l DHA along with 71.4 mg/l EPA and 42.2 mg/l ALA. Genetic engineering of this yeast resulted in a strain that produced 1080 mg/l DHA (17.4 %) along with 74.28 mg/l EPA and 126.72 mg/l ALA possibly through modification of PUFA biosynthetic pathway. To the best of our knowledge, this is a first report of DHA enrichment and opens up avenues for LCPUFA production through L. starkeyi.



Similar content being viewed by others
References
Abbadi A, Domergue F, Bauer J, Napier J, Welti R, Ulrich Z, Cirpus P, Heinz E (2004) Biosynthesis of very long chain polyunsaturated fatty acids in transgenic oilseeds: constraints on their accumulation. Plant Cell 16:2734–2748
Warude D, Joshi K, Harsulkar A (2006) Polyunsaturated fatty acids: biotechnology. Crit Rev Biotechnol 26:83–93
Bajpai P, Bajpai P (1993) Eicosapentaenoic acid (EPA) production from microorganisms: a review. J Biotechnol 30(2):161–183
Gayathri S, Saket P, Mitra A, Kumar A, Shenbaga J, Rajani G, Sundar N, Sivvaswamy N (2010) Soil microorganisms produce omega-3 fatty acids. Indian J Sci Technol 3(5):499–503
Kurtzman C, Albertyn J, Basehoar-Powers E (2007) Multigene phylogenetic analysis of the Lipomycetaceae and the proposed transfer of Zygozyma species to Lipomyces and Babjevia anomala to Dipodascopsis. FEMS Yeast Res 7:1027–1034
Liu H, Zhao X, Wang F, Jiang X, Zhang S, Ye M, Zhao Z, Zou H (2011) The proteome analysis of oleaginous yeast Lipomyces starkeyi. FEMS Yeast Res 11:42–51
Wild R, Patil S, Popovic M, Zappi M, Dufreche S, Bajpai R (2010) Lipids from Lipomyces starkeyi. Food Technol Biotechnol 48(3):329–335
Kraisintu P, Yongmanitchai W, Limtong S (2010) Selection and optimization for lipid production of newly isolated oleaginous yeast, Rhodosporidium toruloides DMKU3-TK16. Kasetsart J (Nat Sci) 44:436–445
Ratledge C, Wynn JP (2002) The biochemistry and molecular biology of lipid accumulation in oleaginous microorganisms. Adv Appl Microbiol 51:1–51
Zhao X, Kong X, Hua Y, Feng B, Zhao Z (2008) Medium optimization for lipid production through co-fermentation of glucose and xylose by the oleaginous yeast Lipomyces starkeyi. Eur J Lipid Sci Technol 110:405–412
Chen G, Qu S, Wang Q, Bian F, Peng Z, Zhang Y, Ge H, Yu J, Xuan N, Bi Y, He Q (2014) Transgenic expression of delta-6 and delta-15 fatty acid desaturases enhances omega-3 polyunsaturated fatty acid accumulation in Synechocystis sp. PCC6803. Biotechnol Biofuels 7:32
Suutari M, Rintamaki A, Laakso S (1996) The effect of temperature on lipid classes and their fatty acid profiles in Lipomyces starkeyi. JOACS 73:1071–1073
Khadake R, Khonde V, Mhaske V, Ranjekar P, Harsulkar A (2010) Functional and bioinformatics characterization of sequence variants of FAD3 gene from flax. J Sci Food Agric 91(14):2689–2696
Somashekar D, Venkateshwaran G, Sambaiah K, Lokesh B (2002) Effect of culture conditions on lipid and gamma-linolenic acid production by mucoraceous fungi. Process Biochem 38(12):1719–1724
Manku M, Horrobin D, Huang Y, Morse N (1983) Fatty acids in plasma and red cell membranes in normal humans. Lipids 18:906–908
Nuria H, Braulio E, Angel G, Albert M, Jose G (2006) Real-time quantitative PCR (QPCR) and reverse transcription-QPCR for detection and enumeration of total yeasts in wine. Appl Environ Micribiol 27(11):7148–7155
Xue Z, Sharpe P, Hong S, Yadav N, Xie D, Short D, Damude H, Rupert R, Seip J, Wang J, Pollak D, Bostick M, Bosak M, Macool D, Hollerbach D, Zhang H, Arcilla D, Bledsoe S, Croker K, McCord E, Tyreus B, Jackson E, Zhu Q (2013) Production of omega-3 eicosapentaenoic acid by metabolic engineering of Yarrowia lipolytica. Nat Biotechnol 31:734–740
Angerbauer C, Siebenhofer M, Mittelbach M, Guebitz G (2008) Conversion of sewage sludge into lipids by Lipomyces starkeyi for biodiesel production. Bioresour Technol 99:3051–3056
Yousuf A, Sannino F, Addorisio V, Pirozzi D (2010) Microbial conversion of olive oil mill wastewaters into lipids suitable for biodiesel production. J Agric Food Chem 58:8630–8635
Marit R, Kristin A, Fenton Julia E, Kaare M, Terje T (2006) The 35S CaMV plant virus promoter is active in human enterocyte-like cells. Eur Food Res Technol 222:185–193
Reddy A, Thomas T (1996) Expression of a cyanobacterial D6-desaturase gene results in gamma-linolenic acid production in transgenic plants. Nat Biotechnol 14:639–642
Hirt H, Kogl M, Murbacher T, Heberle-Bors E (1990) Evolutionary conservation of transcriptional machinery between yeast and plants as shown by the efficient expression from the CaMV 35S promoter and 35S terminator. Curr Genet 17(6):473–479
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Salunke, D., Manglekar, R., Gadre, R. et al. Production of polyunsaturated fatty acids in recombinant Lipomyces starkeyi through submerged fermentation. Bioprocess Biosyst Eng 38, 1407–1414 (2015). https://doi.org/10.1007/s00449-015-1382-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-015-1382-y