Abstract
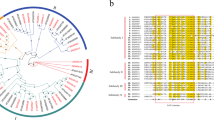
Prunus mume is an ornamental flower and fruit tree in Rosaceae. We investigated the GRAS gene family to improve the breeding and cultivation of P. mume and other Rosaceae fruit trees. The GRAS gene family encodes transcriptional regulators that have diverse functions in plant growth and development, such as gibberellin and phytochrome A signal transduction, root radial patterning, and axillary meristem formation and gametogenesis in the P. mume genome. Despite the important roles of these genes in plant growth regulation, no findings on the GRAS genes of P. mume have been reported. In this study, we discerned phylogenetic relationships of P. mume GRAS genes, and their locations, structures in the genome and expression levels of different tissues. Out of 46 identified GRAS genes, 45 were located on the 8 P. mume chromosomes. Phylogenetic results showed that these genes could be classified into 11 groups. We found that Group X was P. mume-specific, and three genes of Group IX clustered with the rice-specific gene Os4. We speculated that these genes existed before the divergence of dicotyledons and monocotyledons and were lost in Arabidopsis. Tissue expression analysis indicated that 13 genes showed high expression levels in roots, stems, leaves, flowers and fruits, and were related to plant growth and development. Functional analysis of 24 GRAS genes and an orthologous relationship analysis indicated that many functioned during plant growth and flower and fruit development. Our bioinformatics analysis provides valuable information to improve the economic, agronomic and ecological benefits of P. mume and other Rosaceae fruit trees.






Similar content being viewed by others
References
Bolle C (2004) The role of GRAS proteins in plant signal transduction and development. Planta 218:683–692
Bolle C, Koncz C, Chua N (2000) PAT1, a new member of the GRAS family, is involved in phytochrome A signal transduction. Gene Dev 14:1269–1278
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:10
Chen JY (1996) Chinese Mei flowers (in Chinese). Hainan Publishing House, Haikou
Day RB, Shibuya N, Minami E (2003) Identification and characterization of two new members of the GRAS gene family in rice responsive to N-acetylchitooligosaccharide elicitor. BBA Gene Stuct Exp 1625:261–268
Di Laurenzio L, Wysocka-Diller J, Malamy JE, Pysh L, Helariutta Y et al (1996) The SCARECROW gene regulates an asymmetric cell division that is essential for generating the radial organization of the Arabidopsis root. Cell 86:423–433
Dill A, Sun T (2001) Synergistic derepression of gibberellin signaling by removing RGA and GAI function in Arabidopsis thaliana. Genetics 159:777–785
Finn RD, Mistry J, Tate J, Coggill P, Heger A et al (2010) The Pfam protein families database. Nucleic Acids Res 38:D211–D222
Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucleic Acids Res 39:W29–W37
Foster T, Kirk C, Jones WT, Allan AC, Espley R et al (2007) Characterisation of the DELLA subfamily in apple (Malus x domestica Borkh.). Tree Genet Genomes 3:187–197
Fu X, Richards DE, Ait-ali T, Hynes LW, Ougham H et al (2002) Gibberellin-mediated proteasome-dependent degradation of the barley DELLA protein SLN1 repressor. Plant Cell 14:3191–3200
Greb T, Clarenz O, Schäfer E, Müller D, Herrero R et al (2003) Molecular analysis of the LATERAL SUPPRESSOR gene in Arabidopsis reveals a conserved control mechanism for axillary meristem formation. Gene Dev 17:1175–1187
Guo A, Zhu Q, Chen X, Luo J (2007) GSDS: a gene structure display server. Hereditas 29:1023
Helariutta Y, Fukaki H, Wysocka-Diller J, Nakajima K, Jung J et al (2000) The SHORT-ROOT Gene Controls Radial Patterning of the Arabidopsis Root through Radial Signaling. Cell 101:555–567
Hou X, Lee LYC, Xia K, Yan Y, Yu H (2010) DELLAs modulate jasmonate signaling via competitive binding to JAZs. Dev Cell 19:884–894
Ikeda A, Ueguchi-Tanaka M, Sonoda Y, Kitano H, Koshioka M et al (2001) Slender rice, a constitutive gibberellin response mutant, is caused by a null mutation of the SLR1 gene, an ortholog of the height-regulating gene GAI/RGA/RHT/D8. Plant Cell 13:999–1010
Itoh H, Ueguchi-Tanaka M, Sato Y, Ashikari M, Matsuoka M (2002) The gibberellin signaling pathway is regulated by the appearance and disappearance of SLENDER RICE1 in nuclei. Plant Cell 14:57–70
Jaillon O, Aury J, Noel B, Policriti A, Clepet C et al (2007) The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449:463–467
Koizumi Koji, Gallagher LK (2013) Identification of SHRUBBY, a SHORT-ROOT and SCARECROW interacting protein that controls root growth and radial patterning. Development 140:1292–1300
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA et al (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Lee S, Cheng H, King KE, Wang W, He Y et al (2002) Gibberellin regulates Arabidopsis seed germination via RGL2, a GAI/RGA-like gene whose expression is up-regulated following imbibition. Gene Dev 16:646–658
Lee M, Kim B, Song S, Heo J, Yu N et al (2008) Large-scale analysis of the GRAS gene family in Arabidopsis thaliana. Plant Mol Biol 67:659–670
Leister D (2004) Tandem and segmental gene duplication and recombination in the evolution of plant disease resistance genes. Trends Genet 20:116–122
Li X, Qian Q, Fu Z, Wang Y, Xiong G et al (2003) Control of tillering in rice. Nature 422:618–621
Li W, Wu J, Weng S, Zhang Y, Zhang D et al (2010) Identification and characterization of dwarf 62, a loss-of-function mutation in DLT/OsGRAS-32 affecting gibberellin metabolism in rice. Planta 232:1383–1396
Liu RH, Meng JL (2003) MapDraw: a microsoft excel macro for drawing genetic linkage maps based on given genetic linkage data. Hereditas 25:317–321
Llave C, Xie Z, Kasschau KD, Carrington JC (2002) Cleavage of Scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA. Science 297:2053–2056
Morohashi K, Minami M, Takase H, Hotta Y, Hiratsuka K (2003) Isolation and characterization of a novel GRAS gene that regulates meiosis-associated gene expression. J Biol Chem 278:20865–20873
Peng J, Carol P, Richards DE, King KE, Cowling RJ et al (1997) The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses. Gene Dev 11:3194–3205
Peng J, Richards DE, Hartley NM, Murphy GP, Devos KM et al (1999) ‘Green revolution’genes encode mutant gibberellin response modulators. Nature 400:256–261
Piskurewicz U, Lopez-Molina L (2009) The GA-signaling repressor RGL3 represses testa rupture in response to changes in GA and ABA levels. Plant Signal Behav 4:63–65
Pysh LD, Wysocka Diller JW, Camilleri C, Bouchez D, Benfey PN (1999) The GRAS gene family in Arabidopsis: sequence characterization and basic expression analysis of the SCARECROW-LIKE genes. Plant J 18:111–119
Sabatini S, Heidstra R, Wildwater M, Scheres B (2003) SCARECROW is involved in positioning the stem cell niche in the Arabidopsis root meristem. Gene Dev 17:354–358
Schumacher K, Schmitt T, Rossberg M, Schmitz G, Theres K (1999) The Lateral suppressor (Ls) gene of tomato encodes a new member of the VHIID protein family. Proc Natl Acad Sci USA 96:290–295
Shi W (2012) Research on the transcriptome of Prunus mume though RNA-Seq (in Chinese). Beijing Foristory University, Beijing
Silverstone AL, Ciampaglio CN, Sun T (1998) The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellin signal transduction pathway. Plant Cell 10:155–169
Song X, Liu T, Duan W, Ma Q, Ren J et al (2014) Genome-wide analysis of the GRAS gene family in Chinese cabbage (Brassica rapa ssp.pekinensis). Genomics 103:135–146
Stuurman J, Jäggi F, Kuhlemeier C (2002) Shoot meristem maintenance is controlled by a GRAS-gene mediated signal from differentiating cells. Gene Dev 16:2213–2218
Swarbreck D, Wilks C, Lamesch P, Berardini TZ, Garcia-Hernandez M et al (2008) The Arabidopsis information resource (TAIR): gene structure and function annotation. Nucleic Acids Res 36:D1009–D1014
Tamura K, Peterson D, Peterson N, Stecher G, Nei M et al (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tang H, Bowers JE, Wang X, Ming R, Alam M et al (2008a) Synteny and collinearity in plant genomes. Science 320:486–488
Tang H, Wang X, Bowers JE, Ming R, Alam M et al (2008b) Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Res 18:1944–1954
Tian C, Wan P, Sun S, Li J, Chen M (2004) Genome-wide analysis of the GRAS gene family in rice and Arabidopsis. Plant Mol Biol 54:519–532
Torres-Galea P, Huang L, Chua N, Bolle C (2006) The GRAS protein SCL13 is a positive regulator of phytochrome-dependent red light signaling, but can also modulate phytochrome A responses. Mol Genet Genomics 276:13–30
Torres-Galea P, Hirtreiter B, Bolle C (2013) Two GRAS Proteins, SCARECROW-LIKE21 and PHYTOCHROME A SIGNAL TRANSDUCTION1, function cooperatively in phytochrome A signal transduction. Plant Physiol 161:291–304
Velasco R, Zharkikh A, Affourtit J, Dhingra A, Cestaro A et al (2010) The genome of the domesticated apple (Malus × domestica Borkh.). Nat Genet 42:833–839
Wang T, Pan H, Wang J, Yang W, Cheng T et al (2014) Identification and profiling of novel and conserved microRNAs during the flower opening process in Prunus mume via deep sequencing. Mol Genet Genomics 289:169–183
Wen C, Chang C (2002) Arabidopsis RGL1 encodes a negative regulator of gibberellin responses. Plant Cell 14:87–100
Wiśniewska A, Pietraszewska-Bogiel A, Zuzga S, Tagashira N, Łotocka B et al (2013) Molecular characterization of SCARECROW (CsSCR) gene expressed during somatic embryo development and in root of cucumber (Cucumis sativus L.). Acta Physiol Plant 35:1483–1495
Wysocka-Diller JW, Helariutta Y, Fukaki H, Malamy JE, Benfey PN (2000) Molecular analysis of SCARECROW function reveals a radial patterning mechanism common to root and shoot. Development 127:595–603
Zhang Q, Chen W, Sun L, Zhao F, Huang B et al (2012) The genome of Prunus mume. Nat Commun 3:1318
Acknowledgments
This study was supported by Ministry of Science and Technology (Grant No. 2011AA100207 and 2013AA102607) and the State Forestry Administration of China (Grant No. 201004012).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lu, J., Wang, T., Xu, Z. et al. Genome-wide analysis of the GRAS gene family in Prunus mume . Mol Genet Genomics 290, 303–317 (2015). https://doi.org/10.1007/s00438-014-0918-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-014-0918-1