Abstract
Mitochondria are organelles with double-membrane structure of inner and outer membrane, which provides main energy support for cell growth and metabolism. Reactive oxygen species (ROS) mainly comes from mitochondrial and can cause irreversible damage to cells under oxidative stress. Thus, mitochondrial homeostasis is the basis for maintaining the normal physiological function of cells and mitophagy plays a pivotal role in the maintenance of mitochondrial homeostasis. At present, to enhance the sensitivity of cancer cells to radiotherapy and chemotherapy by regulating mitochondria has increasingly become a hot spot of cancer therapy. It is particularly important to study the effect of ionizing radiation (IR) on mitochondria and the role of mitophagy in the radiosensitivity of cancer cells. Most of the existing reviews have focused on mitophagy-related molecules or pathways and the impact of mitophagy on diseases. In this review, we mainly focus on discussing the relationship between mitophagy and radiosensitivity of cancer cells around mitochondria and IR.
Similar content being viewed by others
Introduction
Mitochondria are double-membrane organelles present in most eukaryotic cells, and mitochondria are the main sites of intracellular oxidative phosphorylation and formation of adenosine triphosphate (ATP) (Herst et al. 2017; Pfanner et al. 2019). It can be divided into four functional areas: outer mitochondrial membrane (OMM), membrane space, inner mitochondrial membrane (IMM) and matrix from the outside to the inside. As shown in Fig. 1, the IMM possesses a several-fold larger surface than the OMM, resulting in an invagination of so-called cristae membranes that harbor the oxidative phosphorylation system, including the respiratory complexes I–IV and the F1FO-ATP synthase for ATP. Therefore, major metabolic pathways of mitochondria concern the energy metabolism, such as the tricarboxylic acid cycle (TCA) also known as citric acid cycle or Krebs cycle, and the metabolism of amino acids, lipids and nucleotides (Pfanner et al. 2019). Besides, mitochondria are involved in processes, such as heme biosynthesis, calcium buffering, apoptosis, and innate immune surveillance (Franco-Iborra et al. 2018; Ni et al. 2015). The metabolism of mitochondria mentioned above produces a large amount of ROS, includes the superoxide anion (O2−), hydrogen peroxide (H2O2), hydroxyl radical (·OH), and singlet oxygen (1O2) (Han et al. 2018).
Recent studies showed that ROS levels correlate with the radiosensitivity of cells. There are two major sources for ROS production in cells: mitochondria, which generate ROS as a by-product of respiration; NADPH oxidase (NOX), which actively produces superoxide across the membranes of neutrophils and phagosomes (Li et al. 2015). On the other hand, intracellular ROS remains at a relatively low level and is precisely controlled by endogenous free radical scavengers, such as superoxide dismutase, catalase, and the glutathione peroxidase and thioredoxin reductase systems (Venardos et al. 2007; Finkel and Holbrook 2000). Antioxidants capable of scavenging excessive ROS may help maintain oxidative homeostasis and prevent related damages. Various factors, such as toxins, oxidants and IR, can increase ROS level sharply and make the anti-oxidation defense systems in a weak position, leading to disruption of the balance between generation and elimination of ROS and consequently progression of diseases and even eventual death (Dalle-Donne et al. 2001; Yu et al. 2006; Motoori et al. 2001; Ayaka et al. 2012).
Mitochondrial homeostasis is particularly important for ROS level and maintained by an intricate balance between fission, fusion, mitochondrial biogenesis, and mitophagy (Chan 2012; Hoitzing et al. 2015). Mitophagy as a kind of selective autophagy is one of the main pathways for mitochondrial quality and quantity control. The word autophagy is derived from the Greek roots “auto” (self) and “phagy” (eating) (Li et al. 2020). As the name implies, mitophagy is an intracellular degradation pathway in eukaryotes, which depends on lysosomes, resulting in some damaged or excess aging mitochondria being swallowed and degraded. Previous studies have shown that ROS is closely related to autophagy (Yang et al. 2008; Pyo et al. 2008; Dadakhujaev et al. 2008; Chen et al. 2008a, b). And mitophagy also can be further stimulated by oxidative stress. Upon stress or injury, mitophagy prevents the accumulation of damaged mitochondria and the increased level of ROS leading to oxidative stress and cell death (Ashrafi and Schwarz 2013).
In summary, we hold the opinion that there is also a relationship between mitophagy and the radiosensitivity of cells, but there is no related review of this issues at present. Next, we will discuss the effects of IR on cancer cells and mitochondria, and the effects of mitophagy on the radiosensitivity of cancer cells.
Effects of IR on cancer cells
ROS and DNA
The killing effects of IR on cells are divided into direct and indirect effects. IR could cause irreparable single-strand or double-strand breaks of DNA, resulting in cell death, which belong to the direct effects of IR. Indirect effects refer to the reaction of IR with water molecules in cancer cells to form oxygen free radicals, causing oxidative stress, which is more damaging to cells. Among them, oxidative stress is larger and more serious to cells. Oxidative stress is defined as the imbalance between oxidants-ROS, Reactive Nitrogen Species (RNS) and antioxidants. Under oxidative stress, excessive accumulation of ROS could further aggravate oxidative damage using mitochondria as primary targets and destroy cellular proteins, lipids and DNA, resulting in fatal cell damage, which in turn involves a variety of pathology, such as aging, cancer, metabolic syndrome, neurodegenerative diseases, cardiovascular disease, diabetes, and so on (Ichimiya et al. 2020; Rodolfo et al. 2018; Drake et al. 2017).
Radiosensitivity
The radiosensitivity of tumor cells is related to many factors, such as type of tumor cells, presence of cancer stem cells, type of radiation, and tumor microenvironment. More and more studies have shown that mitochondria also play an important role in the radiosensitivity of cells, energy metabolism, mitochondrial apoptosis, mitophagy, regulation of redox homeostasis, and calcium influx all play an indispensable role in this process (Lynam-Lennon et al. 2014; Yuehua et al. 2018; Yang et al. 2021). Mitochondria are the main source of energy for cell growth and proliferation. Various external stimuli lead to mitochondrial damage and loss of oxidative phosphorylation function, which will affect cell metabolic activities and eventually lead to cell apoptosis or death. This is also the main mechanism of action of many chemotherapy drugs at present. Therefore, mitochondria are increasingly becoming an important target for studying the effect of enhancing tumor killing.
Effects of IR on mitochondria
Elevated ROS levels
As mentioned above, ROS is a key medium of mitochondrial damage induced by IR, and one of the main sources of ROS is the respiratory chain of IMM (Addabbo et al. 2009; Zorov et al. 2014; Turrens 2003). ROS production of mitochondria will increase in cells after being irradiated. Hosoki et al. revealed that the cellular level of ROS increased in HeLa S3 cells during post-irradiation (Ayaka et al. 2012). Motoori et al. used di-hydro-rhodamine 123 to detect mitochondrial ROS and radiation incubation using flow cytometry with a fluorescent probe showed a radiation-induced elevation in mitochondrial ROS in human hepatoma cells (Motoori et al. 2001). Zhang et al. showed that small airway epithelial exposed to 10α particles through the cytoplasm resulted in an increase in fluorescence intensity of superoxide production at 2 h post irradiation at a level that was three times that of control (Zhang et al. 2013).
Mitochondrial membrane potential (ΔΨm) depolarization
The IMM is distributed with many proton pumps, whose function is to pump intra-matricial protons (H+) into the outer side of the inner membrane, thus forming ΔΨm across the inner membrane to maintain the normal function of mitochondria. When mitochondria are exposed to IR, it can lead to obstacles in the electron transport process of the respiratory chain and affect the formation of H+ transmembrane gradient, which will lead to a decrease in the ΔΨm of the original external positive and internal negative, that is, ΔΨm depolarization. Mitochondrial depolarization signals cause autophagy-related proteins, such as PINK, to stabilize the OMM and accumulate, initiating the process of mitophagy (Ashrafi and Schwarz 2013).
Mitochondrial DNA (mtDNA) damage
Human mtDNA is a circular double-stranded molecule with a full length of 16,569 bp and it can be divided into heavy chain and light chain.
MtDNA accounts for only 1–2% of the total human DNA, it has no intron and contains 37 genes encoding 13 polypeptides, 2 ribosomal RNAs (rRNAs), and 22 transfer RNAs (tRNAs) (Mishra and Chan 2014; Anderson et al. 1981). There is also the only two non-coding regions, namely D-loop and the replication start point of the light chain, the former is the main regulatory region of mtDNA replication and transcription. The part of nuclear DNA that is involved in protein coding accounts for only 1% of the total human DNA, so scholars generally believe that damage of mtDNA is more likely to lead to disease than nuclear DNA (Birney et al. 2007). Specifically, it is as follows: (1) mtDNA is bare and lacks histone protection; (2) The high fat/DNA value in mitochondria makes lipophilic carcinogens preferentially aggregate on mtDNA; (3) mtDNA is in a state of continuous synthesis throughout the cell cycle, which is susceptible to external interference and poor stability; (4) The high concentration of oxygen in mitochondria is easy to produce oxygen free radicals and hydrogen peroxide, and due to the scarce synthesize glutathione, mtDNA is susceptible to oxidative damage; (5) High frequency of replication mismatch; (6) With the not-so-perfect mechanism for repairing DNA damage (West and Shadel 2017; Yoshida et al. 2012; Clayton et al. 1974; Robert et al. 1975; Croteau et al. 1999; Larsen et al. 2005; Heddi et al. 1999; Penta et al. 2001). To sum up, the damage of mtDNA will be more obvious when cells are exposed to IR. Many scholars also have confirmed that ROS can cause more extensive and lasting damage to mtDNA, and mtDNA depletion can also in turn affect cellular oxidative stress (Yakes and Houten 1997; Azzam et al. 2012; Kim et al. 2008; Zhou et al. 2011; Kobashigawa et al. 2011; Garza-Lombó et al. 2020).
Mitochondria swelling
Mild mitochondrial swelling under physiologic conditions regulates metabolism and function of mitochondria, whereas excessive swelling causes mitochondrial dysfunction (Makarov et al. 2020). The abnormal accumulation of ROS could upregulate inositol triphosphate, resulting in the opening of inositol triphosphate receptor-calcium ion channels on the endoplasmic reticulum membranes, calcium ions are released from the endoplasmic reticulum. The high level of calcium ion/calmodulin kinase (CaMK) in the cytoplasm can open the L-type voltage-gated calcium ion channel of the plasma membrane and the inflow of extracellular calcium ions, which further induces the opening of the mitochondrial bilayer membrane permeable pores, making the mitochondria swell and rupture (Orrenius et al. 2015).
Mitophagy
As we can see from the above, ROS plays a particularly important role in the effects of IR on cells and mitochondria, and mitochondria are the main sites to product ROS. In addition to ROS, other IR-induced alterations in mitochondria can also initiate a series process, such as mitochondrial elimination, fission and fusion, and mitochondrial biogenesis, to maintain mitochondria homeostasis. Whole mitochondrial elimination is accomplished by a selective form of autophagy: mitophagy. The activation of mitophagy is closely related to the redox of cells, such as hypoxic environment and IR, can put cells under oxidative stress and induce the occurrence of mitophagy. Next, we made a review of mitophagy-related molecular receptors and pathways.
Mitophagy-related molecular receptors (shown in Fig. 1)
Various metabolic activities in cells are transmitted by signaling pathways and molecules, and in recent years, great progress has been made in the study of mitophagy mechanism. Microtubule associated protein 1 light chain 3 (LC3) is the most used autophagy marker. LC3 gene is the mammalian homologous gene of autophagy-related protein8 (ATG8), which encodes three proteins: LC3A, LC3B and LC3C. During autophagy, LC3B cleaves to soluble protein LC3B-I, which combines with phosphatidylethanolamine to form LC3B-II. LC3B-II can accumulate in large quantities on the surface of neonatal autophagosomes, which is one of the reliable markers of autophagy (Zhihong et al. 2013). When mitophagy starts, LC3B-II binds to the proteins on mitochondria, causing autophagosomes to wrap damaged mitochondria to form mitochondrial autophagosomes, which is a key step in the occurrence of mitophagy. There are a variety of mitophagy receptors on the mitochondrial membrane, which are listed in Table 1. When cells are irradiated to cause mitochondrial injury or depolarization, such as the most common oxidative stress, changes in the mitochondria themselves will stimulate the changes of autophagic proteins, initiating mitophagy.
SQSTM1-like receptors
Mitochondrial receptor proteins act as junctions between autophagosomes and mitochondria, one of which is called SQSTM1-like receptors (SLRs). In addition to interacting with LC3, these proteins have domains that bind to ubiquitin, through which autophagosomes and mitochondria are connected. Currently, SLRs include SQSTM1, CALCOCO2/NDP52, OPTN, NBR1, and CALCOCO3/TAX1BP1.
SQSTM1/p62 assists selective macro-autophagy and acts as a molecular sentinel in the auto-phagosome membrane for the recognition, sequestration, and degradation of intracellular wastes. In addition, p62 is defined as the odd-jobber protein, able to orchestrate autophagy, coordinate stem cell differentiation, quench inflammation and actively participate in tumor stroma recovery through immune cells recruitment and anti-metastatic activity (Sabbieti et al. 2022).
CALCOCO family proteins are the newly found selective autophagy receptors, which include CALCOCO1, CALCOCO2/NDP52, and CALCOCO3/TAX1BP1 (Chen et al. 2022). NDP52 is a member of the nucleus point family and is distributed in both the cytoplasm and nucleus. The amino end is the SKICH30 domain, the central region is the curly spiral domain, and the carboxyl end is composed of one LIM domain and two zinc finger structures (Morriswood et al. 2007). It is the primary receptor for PINK1/Parkin-mediated mitophagy. The ubiquitin kinase PINK1 recruits NDP52 and Optineurin to mitochondria to directly activate mitophagy independent of Parkin (Michael et al. 2015). In addition, NDP52 is also involved in xenophagy and immune response (Sharma et al. 2018; Leymarie et al. 2017). TAX1BP1 is a selective macro-autophagy/autophagy receptor that plays a central role in host defense against pathogens and regulation of the innate immune system. TAX1BP1 can also perform a variety of auxiliary functions, affecting the biogenesis and maturation of auto-phagosomes (Chen et al. 2022). Other study has shown that TAX1BP1 also plays a role in the negative regulation of cell growth and apoptosis (Verstrepen et al. 2011).
OPTN, like NDP52, it functions as a pink1 downstream protein and contains several structural domains, including two coiled-coil domains, a leucine zipper domain, an LC3-interacting region (LIR), a ubiquitin-binding domain and a zinc finger domain. OPTN-mediated autophagy-dysfunction is closely related to a variety of diseases, such as Neurodegenerative diseases, Neurodegenerative diseases, Cancer, and Nephropathy. Other study has shown the HACE1–optineurin axis promotes tumor suppression in an ubiquitin-dependent fashion (Liu et al. 2014). And OPTN play distinctive roles in different diseases, depending on its primary molecular function in each respective condition (Qiu et al. 2022).
NBR1 was discovered as a selective autophagy receptor due to its interaction with and similarity in domain organization to p62 and direct binding to ATG8 proteins and ubiquitin (Kirkin 2009). NBR1 and p62 share an N-terminal PB1 domain, the ZZ zinc finger domain, LIR motif, and C-terminal UBA domains. In addition, NBR1 contains the four tryptophan domains involved in protein–protein interactions, two coiled-coil domains, and an amphipathic helix domain not found in p62 (Rasmussen et al. 2022). Besides, it also plays an important role in cancer metastasis and immune evasion (Yamamoto et al. 2020; Marsh and Debnath 2020).
In addition to SLRs, some mitochondrial membrane proteins can also bind directly to LC3 without relying on ubiquitin. The common feature of these proteins is the conserved key region, which could directly bind to ATG8/LC3 or other proteins in the family. The following proteins are classified and described according to the location in mitochondria:
OMM proteins
FUNDC1 is a 155-amino acid macromolecular protein with 3 transmembrane domains, in addition, the N-terminal domain exposed to the cytoplasm contains a domain LIR that interacts with LC3 (Qiu et al. 2022). Under normal circumstances, phosphorylated FUNDC1 is present in the OMM, and when in a hypoxic environment, unc-51-like kinase 1 (ULK1) phosphorylated Ser17 on FUNDC1, and phosphoglycerate mutase family member 5 (PGAM5) dephosphorylates Ser13, thereby promoting FUNDC1 LIR interacts with LC3 to initiate mitophagy occurring. Other study has shown that FUNDC1 is also involved in regulating mitochondrial fission and fusion to help mitochondrial quality control (Chen et al. 2016a, b).
NIX, also known as BNIP3L, is a member of the B cell lymphoma-2 (Bcl-2) family and has 56% homology with BNIP3. It is a mitochondrial autophagy receptor located in the outer membrane of the mitochondria. During the maturation of mammalian erythrocytes, Nix-mediated mitochondrial autophagy plays an important role in the removal of mitochondria (Ashrafi and Schwarz 2013). The related mechanism may be that NIX acts as an autophagy receptor to recruit autophagy-related molecules to initiate autophagy (Zhang and Ney 2009). In addition, NIX is similar to BNIP3 in function, and its C-terminal transmembrane domain induces apoptosis by interacting with Bcl-2 and BCL-XL (Diwan et al. 2007; Zhang and Ney 2009). NIX protein can not only induce apoptosis but also mediate mitochondrial autophagy (Dorn 2010).
BNIP3 and NIX are both members of the Bcl-2 family containing only the BH3 domain subfamily (Lampert et al. 2019). BNIP3 and NIX were found to be a class of pro-apoptotic proteins (Fei et al. 2004), which are highly expressed in cancer cells and cardiomyocytes under hypoxic conditions and are closely related to cell death. Then these two proteins were found that both of them also play an important role in promoting mitophagy of cell survival. BNIP3 induces the release of Beclin-1 by competitively binding Bcl-2 with the autophagy core protein Beclin-1, and then activates autophagy. In addition, there is a conservative LIR that recognizes auto-phagosome LC3 at the N-terminal of BNIP3, which can promote mitophagy. Several studies have shown that elevated mitochondrial ROS increases the expression of BNIP3/NIX to trigger mitophagy (Chourasia et al. 2015; Li et al. 2015; Hu et al. 2016).
Bcl2L13 is also a member of the Bcl-2 protein family and is in the OMM. It has four conserved BH structures and one transmembrane structure of the Bcl-2 family (Meng et al. 2021). Bcl2L13 is a newly reported mitophagy receptor, which is a homologous protein of mitophagy-related gene ATG32, which mediates the clearance of damaged mitochondria (Murakawa et al. 2015.). In addition, like all members of the Bcl-2 protein family, Bcl2L13 has the function of regulating apoptosis (Meng et al. 2021).
FKBP8 is a member of the FK506-binding protein family and the only member of the family with a transmembrane domain, which enables it to be located on the membrane, such as mitochondria and endoplasmic reticulum (Misaka et al. 2018; Shirane et al. 2003). In addition, FKBP8 has four domains: peptidyl-prolyl cis–trans isomerase domain, TPR sequence, calmodulin-binding site and glutamate-rich region. FKBP8 can regulate mitophagy by combining LIR with autophagosome marker protein LC3 (Bhujabal et al. 2017). Different from other autophagy receptors, FKBP8 can transfer from mitochondria to endoplasmic reticulum during mitophagy, to avoid being degraded by auto-phagosomes together with damaged mitochondria. The mechanism of this escape remains to be studied. In addition, FKBP8 can regulate apoptosis through peptidyl-prolyl cis–trans isomerase domain, TPR sequence, calmodulin-binding site, and apoptosis regulator Bcl-2 (Edlich and Lücke 2011).
AMBRA1 is composed of 1300 amino acids with a molecular weight of about 130 kDa. When mitochondria are damaged, the depolarization of mitochondrial membrane potentials significantly increases the interaction between Parkin and Ambra1. Ambra1 are recruited around the depolarized mitochondria in a Parkin-dependent manner, activating the PI3KIII complex and promoting its clearance through autophagy. In addition, Ambra1-LC3 interaction can transport damaged mitochondria to autophagosomes, which is essential for amplifying Parkin-mediated mitochondrial clearance. Ambra1 can mediate mitophagy in human neuroblastoma SH-SY5Y cells, thus inhibiting oxidative stress and apoptosis induced by 6-hydroxydopamine and rotenone (Di Rita et al. 2018). Many studies have shown that Ambra1-mediated autophagy may be an important mechanism of drug resistance. At the same time, Ambra1 may also play a role in tumor inhibition by regulating other signal pathways (Liu et al. 2017; Sun et al. 2019, 2018).
NLRX1 (nucleotide-binding domain and leucine-rich-repeat-containing protein X1) belongs to the NLR family of intracellular sensors that regulate major cellular pathways including cell death and inflammation (Imbeault et al. 2014). In addition, study has shown that it is also related to cellar apoptosis (Soares et al. 2014).
IMM proteins
PHB2 belongs to the anti-proliferative protein family PHBs, which mainly encodes two protein members, PHB1 and PHB2 in the human genome. PHB1 and PHB2 can be combined as heterodimers to form a rosette palisade protein complex anchored in the IMM, which is involved in maintaining the stability of mitochondrial structure, regulating mitochondrial dynamics and mitochondrial crest morphology, regulating mitochondrial differentiation and development, and maintaining the stability of mitochondrial genome (Merkwirth and Langer 2009). PHB2 contains transmembrane domain (amino acid 1–36), central prohibitin domain (amino acid 36–201) and overlapping coiled helix domain (amino acid 188–264) for mitochondrial localization. In addition, PHB2 also contains estrogen receptor binding domain. It is mainly located in mitochondria, but also in cytoplasm, nucleus and plasma membrane (Kuramori et al. 2009). Prohibitin-2 can stabilize PINK1 on the OMM to mediate the PINK1-parkin pathway and promote mitophagy (Yan et al. 2020). When the OMM is damaged and the IMM is exposed, it can directly bind to LC3 to assist in the timely phagocytosis and degradation of damaged mitochondria by lysosomes (Wei et al. 2017).
Cardiolipin (CL), is a kind of mitochondrial-specific phospholipid, which is mainly distributed on the IMM. It is composed of two phosphoric acid molecules and three glycerol molecules, and then connected with four fatty acid molecules. CL can produce negative curvature elastic stress in lipid bilayer membrane, so it plays a key role in maintaining a certain curvature of mitochondrial inner membrane crest (Ikon and Ryan 2017). Chu et al. confirmed that CL can be transported from the IMM to the OMM after mitochondrial injury, which directly interacts with LC3 and mediates mitophagy (deArriba et al. 2013; Fernandez et al. 2002; Chu et al. 2013). In addition, cardiolipin–cytochrome C complex can also regulate apoptosis (McMillin and Dowhan 2002).
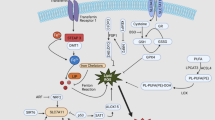
Mitophagy-related signaling pathways (shown in Fig. 2)
Process and related pathways of mitophagy. Note: (1) mitochondrial damage leads to mitochondrial depolarization and induces the activation of mitophagy-related proteins; (2) autophagosomes wrap damaged mitochondria to form mitochondrial autophagosomes; (3) lysosome fusion to form mitochondrial autophagy lysosomes; (4) degradation and recycling of related substances; (a) PINK1-parkin pathway; (b) Parkin-independent pathway; (c) direct interaction of LC3 with IMM/OMM proteins
PINK1-parkin pathway
PINK1-parkin is currently one of the most classic signaling pathways of mitophagy. PINK1 is a 581 amino acid protein with serine/threonine protein kinase activity, consisting of an N-terminal mitochondrial targeting motif that contains a transmembrane domain (110 amino acids long), a highly conserved kinase domain with three insertions in the N lobe, and a C-terminal autoregulatory sequence (Kumar et al. 2017). PINK1 is synthesized in the cytoplasm and is very low in normal mitochondria, but it could accumulate in the OMM when mitochondria are under depolarizing state, which could act as molecular receptors for damaged mitochondria. Then it phosphorylates Parkin and ubiquitin to recruit them to the damaged mitochondria, leading to the ubiquitination of mitochondrial proteins and the initiation of mitophagy. Parkin is a downstream protein of PINK1, an E3 ubiquitin ligase composed of 465 amino acids encoded by the PRKN gene, which mainly mediates substrate ubiquitination, regulates protein degradation and signal transduction. PINK1 acts as a key sensor of mitochondrial damage, whereas Parkin amplifies this damage signal by facilitating the formation of ubiquitin chains, which recruit more Parkin to the damaged mitochondria. Phosphorylated parkin cascades amplify ubiquitination signals and initiate mitophagy. Lin et al. found that PINK1-parkin pathway of mitophagy protects cells via decreasing mitochondrial ROS (Lin et al. 2019).
Additionally, parkin-dependent ubiquitination of OMM proteins is necessary for mitophagy, it probably acts as a priming event that allows the OMM localized NIPSNAP1 and NIPSNAP2 to recruit the autophagy receptors. Several studies have shown that NIPSNAP1 and NIPSNAP2 will locate in the OMM when mitochondria are depolarized by external stimulation, sending out “eat me” signals to recruit ATG8-family proteins and ubiquitin-dependent SLRs to mediate more powerful mitophagy (Abudu et al. 2019; Princely Abudu et al. 2019).
Parkin-independent pathway
The depolarization of the mitochondrial membrane makes PINK1 stable in the OMM. PINK1 can directly exert its own kinase activity without parkin, phosphorylate and activate ubiquitin proteins, and then ubiquitinate the proteins related to autophagy. Polyubiquitin chain plays the role of promoting mitophagy (Ordureau et al. 2014).
Other pathway
In addition, there are some pathways that are independent of parkin and ubiquitin, and directly mediate the process of mitophagy through mitochondrial membrane proteins. The OMM and IMM proteins mentioned in “OMM proteins” and “IMM proteins” above can start the process of mitophagy binding to LC3 on autophagy lysosomes.
Relationship between mitophagy and radiosensitivity of cancer cells
According to the current research on mitophagy in cellular radiosensitivity, as shown in Fig. 1, there are mainly the following two perspectives:
On the one hand, mitophagy can enhance the radio-resistance of cancer cells. That is, mitophagy play a protective role in irradiated tumor cells. The existence of mitophagy can identify and clear the damaged mitochondria early, the decrease of the number of mitochondria inhibits the process of oxidative phosphorylation and compensatively induces glycolysis, and glycolysis increases the contents of lactic acid, pyruvate, and ketone bodies in cells. These metabolites are secreted by tumor matrix and reused by cancer cells to meet their metabolic needs and maintain intracellular homeostasis. Besides, the rapid renewal of mitochondria leads to the presence of a large number of newly generated mitochondria (Wu et al. 2022). Several research results support the above point of view. Chen et al. demonstrated through in vivo and in vitro experiments that the LACTB2 protein rendered nasopharyngeal carcinoma resistant to radiation therapy, and PINK1/Parkin-mediated mitophagy induced a healthier mitochondrial network and contributed to radio-resistance of nasopharyngeal carcinoma (Chen et al. 2021). Yang et al. have shown that mitophagy can resist oxidative stress caused by IR, and inhibition of mitophagy can increase the accumulation of ROS and induce cancer cell death (Yang et al. 2021). Wang et al. found that protein disulfide isomerase can inhibit radiotherapy-induced cell death by regulating mitophagy signaling, increasing cellular radio-resistance. Cancer stem cells population has higher mitophagy level, which could promote tumorigenesis and cell survival in various tumor types by allowing the removal of abnormal mitochondria (Baghban et al. 2020; Takeda et al. 2019). Fan et al. also proved that early oxidative stress enhanced mitophagy to protect cells, but mitophagy was reduced and apoptosis was increased once the cells had irreversible damage after long-term oxidative exposure (Fan et al. 2019). Wu et al. found that the level of mitophagy was elevated in radio-resistant A549R cells, and inhibition of mitophagy can increase the radiosensitivity of A549R cells (Wu et al. 2022), they believe that it is related to ROS and DNA damage.
On the other hand, increasing radiosensitivity of cancer cells. This view is mainly related to intracellular ROS and DNA damage. DNA damage is closely related to tumor occurrence and development. Ren et al. found that mitophagy, as an upstream signal, increases ionizing radiation-induced DNA damage by downregulating or overexpressing key mitophagy proteins Parkin and BNIP3 (Ren et al. 2023). H2O in irradiated cells can be dissociated to ROS including H2O2 and \({\text{O}}_{2}^{ \cdot - }\), then a cascade reaction will tend to occur. Furthermore, mitochondria are also the sources of ROS. Strong external stimulation causes excessive mitochondrial damage, serious imbalance of cellular oxidative phosphorylation, excessive accumulation of ROS, inducing excess mitochondria are removed and causing cell death. Yu and Chen et al. irradiated Hela and MCF-7 cells and found that radiation can induce autophagy, resulting in increased intracellular ROS levels and increased mitochondrial damage, mitophagy here increases the sensitivity of cells to IR (Yu et al. 2021; Chen et al. 2016a, b). In addition, insufficient mitophagy can lead to the accumulation of harmful substances and apoptosis or death of cells.
Conclusion
This review summarized the receptors and pathways related to mitophagy, and mainly discussed the relationship between mitophagy and radiosensitivity of cancer cells around mitochondria and IR. In summary, IR as an external stimulus can affect the occurrence and activity of mitophagy of cancer cells. Mitophagy plays a “Double-edged sword” role in the radiosensitivity of cancer cells, and the time–effect relationship and dose–effect relationship need to be further explored.
Data availability
The datasets generated during and/or analysed during the current study are available in the PubMed: https://pubmed.ncbi.nlm.nih.gov/.
Abbreviations
- ROS:
-
Reactive oxygen species
- IR:
-
Ionizing radiation
- ATP:
-
Adenosine triphosphate
- OMM:
-
Outer mitochondrial membrane
- IMM:
-
Inner mitochondrial membrane
- TCA:
-
Tricarboxylic acid cycle
- NOX:
-
NADPH oxidase
- RNS:
-
Reactive nitrogen species
- ΔΨm:
-
Mitochondrial membrane potential
- mtDNA:
-
Mitochondrial DNA
- rRNAs:
-
Ribosomal RNAs
- tRNAs:
-
Transfer RNAs
- CaMK:
-
Calcium ion/calmodulin kinase
- FC:
-
Flow cytometry
- LC3:
-
Light chain 3
- ATG:
-
Autophagy-related protein
- SLRs:
-
SQSTM1-like receptors
- LIR:
-
LC3-interacting region
- ULK1:
-
Unc-51-like kinase 1
- PGAM5:
-
Phosphoglycerate mutase family member 5
- Bcl-2:
-
B-cell lymphoma-2
- CL:
-
Cardiolipin
References
Abudu YP, Pankiv S, Mathai BJ, Lamark T, Johansen T, Simonsen A (2019) NIPSNAP1 and NIPSNAP2 act as “eat me” signals to allow sustained recruitment of autophagy receptors during mitophagy. Autophagy 15:1845–1847
Addabbo F, Montagnani M, Goligorsky MS (2009) Mitochondria and reactive oxygen species. Hypertension 53:885–892
Anderson S, Bankier AT, Barrell BG, de Bruijn MH, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, Sanger F, Schreier PH, Smith AJ, Staden R, Young IG (1981) Sequence and organization of the human mitochondrial genome. Nature 290:457–465
Ashrafi G, Schwarz TL (2013) The pathways of mitophagy for quality control and clearance of mitochondria. Cell Death Differ 20:31–42
Ayaka H, Shin-Ichiro Y, Qing-Li Z, Zheng-Li W, Ichiro T, Yoshiaki T, Li-Li W, Shiga H, Takaharu N, Akira T (2012) Mitochondria-targeted superoxide dismutase (SOD2) regulates radiation resistance and radiation stress response in HeLa cells. J Radiat Res 53:58–71
Azzam EI, Jay-Gerin JP, Pain D (2012) Ionizing radiation-induced metabolic oxidative stress and prolonged cell injury. Cancer Lett 327:48–60
Baghban R, Roshangar L, Jahanban-Esfahlan R, Seidi K, Zare P (2020) Tumor microenvironment complexity and therapeutic implications at a glance. Cell Commun Signal 18:1–19
Bhujabal Z, Birgisdottir ÅB, Sjøttem E, Brenne HB, Øvervatn A, Habisov S, Kirkin V, Lamark T, Johansen T (2017) FKBP8 recruits LC3A to mediate Parkin-independent mitophagy. EMBO Rep 18:947–961
Birney E, Stamatoyannopoulos JA, Dutta A, Guigó R, Gingeras TR, Margulies EH, Weng Z, Snyder M, Dermitzakis ET (2007) Identification and analysis of functional elements in 1% of the human genome by ENCODE pilot project. Nature 447:799–816
Chan DC (2012) Fusion and fission: interlinked processes critical for mitochondrial health. Annu Rev Genet 46:265–287
Chen F, Wang CC, Kim E, Harrison LE (2008a) Hyperthermia in combination with oxidative stress induces autophagic cell death in HT-29 colon cancer cells. Cell Biol Int 32:715–723
Chen YQ, Mcmillan-Ward E, Kong JM, Israels SJ, Gibson SB (2008b) Oxidative stress induces autophagic cell death independent of apoptosis in transformed and cancer cells. Cell Death Differ 15:171
Chen M, Chen Z, Wang Y, Tan Z, Zhu C, Li Y, Han Z, Chen L, Gao R, Liu L, Chen Q (2016a) Mitophagy receptor FUNDC1 regulates mitochondrial dynamics and mitophagy. Autophagy 12:689–702
Chen Z, Wang B, Yu F, Chen Q, Tian Y, Ma S, Liu X (2016b) The roles of mitochondria in radiation-induced autophagic cell death in cervical cancer cells. Tumour Biol J Int Soc Oncodev Biol Med 37:4083–4091
Chen Q, Zheng W, Zhu L, Liu H, Song Y, Hu S, Bai Y, Pan Y, Zhang J, Guan J, Shao C (2021) LACTB2 renders radio resistance by activating PINK1/Parkin-dependent mitophagy in nasopharyngeal carcinoma. Cancer Lett 518:127–139
Chen W, Ouyang X, Chen L, Li L (2022) Multiple functions of CALCOCO family proteins in selective autophagy. J Cell Physiol 237:3505–3516
Chourasia AH, Tracy K, Frankenberger C, Boland ML, Sharifi MN, Drake LE, Sachleben JR, Asara JM, Locasale JW, Karczmar GS, Macleod KF (2015) Mitophagy defects arising from BNip3 loss promote mammary tumor progression to metastasis. EMBO Rep 16:1145–1163
Chu CT, Ji J, Dagda RK, Jiang JF, Tyurina YY, Kapralov AA, Tyurin VA, Yanamala N, Shrivastava IH, Mohammadyani D, Wang KZQ, Zhu J, Klein-Seetharaman J, Balasubramanian K, Amoscato AA, Borisenko G, Huang Z, Gusdon AM, Cheikhi A, Steer EK, Wang R, Baty C, Watkins S, Bahar I, Bayir H, Kagan VE (2013) Cardiolipin externalization to the outer mitochondrial membrane acts as an elimination signal for mitophagy in neuronal cells. Nat Cell Biol 15:1197–1205
Clayton DA, Doda JN, Friedberg EC (1974) The absence of a pyrimidine dimer repair mechanism in mammalian mitochondria. Proc Natl Acad Sci USA 71:2777–2781
Croteau DL, Stierum RH, Bohr VA (1999) Mitochondrial DNA repair pathways. Mutat Res DNA Repair 434:137–148
Dadakhujaev S, Noh HS, Jung EJ, Hah YS, Chang JK, Kim DR (2008) The reduced catalase expression in TrkA-induced cells leads to autophagic cell death via ROS accumulation. Exp Cell Res 314:3094–3106
Dalle-Donne I, Rossi R, Milzani A, Di Simplicio P, Colombo R (2001) The actin cytoskeleton response to oxidants: from small heat shock protein phosphorylation to changes in the redox state of actin itself. Free Radic Biol Med 31:1624–1632
de Arriba G, Calvino M, Benito S, Parra T (2013) Cyclosporine A-induced apoptosis in renal tubular cells is related to oxidative damage and mitochondrial fission. Toxicol Lett 218(1):30–38
Di Rita A, D’Acunzo P, Simula L, Campello S, Strappazzon F, Cecconi F (2018) AMBRA1-mediated mitophagy counteracts oxidative stress and apoptosis induced by neurotoxicity in human neuroblastoma SH-SY5Y cells. Front Cell Neurosci 12:92
Diwan A, Koesters AG, Odley AM, Pushkaran S, Baines CP, Spike BT, Daria D, Jegga AG, Geiger H, Aronow BJ, Molkentin JD, Macleod KF, Kalfa TA, Dorn GW (2007) Unrestrained erythroblast development in Nix−/− mice reveals a mechanism for apoptotic modulation of erythropoiesis. Proc Natl Acad Sci USA 104:6794–6799
Dorn GW (2010) Mitochondrial pruning by Nix and BNip3: an essential function for cardiac-expressed death factors. J Cardiovasc Transl Res 3:374–383
Drake LE, Springer MZ, Poole LP, Kim CJ, Macleod KF (2017) Expanding perspectives on the significance of mitophagy in cancer. Semin Cancer Biol 47:110–124
Edlich F, Lücke C (2011) From cell death to viral replication: the diverse functions of the membrane-associated FKBP38. Curr Opin Pharmacol 11:348–353
Fan P, Yu XY, Xie XH, Chen CH, Wang YT (2019) Mitophagy is a protective response against oxidative damage in bone marrow mesenchymal stem cells. Life Sci 229:36–45
Fei P, Wang W, Kim SH, Wang S, Burns TF, Sax JK, Buzzai M, Dicker DT, McKenna WG, Bernhard EJ, El-Deiry WS (2004) Bnip3L is induced by p53 under hypoxia, and its knockdown promotes tumor growth. Cancer Cell 6:597–609
Fernandez MG, Troiano L, Moretti L, Nasi M, Cossarizza A (2002) Early Changes in Intramitochondrial Cardiolipin Distribution during Apoptosis. Cell Growth Differ Mol Biol J Am Assoc Can Res 13:449–455
Finkel T, Holbrook NJ (2000) Oxidants, oxidative stress and the biology of ageing. Nature 408:239–247
Franco-Iborra S, Vila M, Perier C (2018) Mitochondrial quality control in neurodegenerative diseases: focus on Parkinson’s disease and Huntington’s disease. Front Neurosci 12:342
Garza-Lombó C, Pappa A, Panayiotidis MI, Franco R (2020) Redox homeostasis, oxidative stress and mitophagy. Mitochondrion 51:105–117
Han Y, Ishibashi S, Iglesias-Gonzalez J, Chen Y, Love NR, Amaya E (2018) Ca-induced mitochondrial ROS regulate the early embryonic cell cycle. Cell Rep 22:218–231
Heddi A, Stepien G, Benke PJ, Wallace DC (1999) Coordinate induction of energy gene expression in tissues of mitochondrial disease patients. J Biol Chem 274:22968
Herst PM, Rowe MR, Carson GM, Berridge MV (2017) Functional mitochondria in health and disease. Front Endocrinol 8:296
Hoitzing H, Johnston IG, Jones NS (2015) What is the function of mitochondrial networks? A theoretical assessment of hypotheses and proposal for future research. BioEssays 37:687–700
Hu L, Wang H, Huang L, Zhao Y, Wang J (2016) The protective roles of ROS-mediated mitophagy on I seeds radiation induced cell death in HCT116 cells. Oxid Med Cell Longev 2016:9460462
Ichimiya T, Yamakawa T, Hirano T, Yokoyama Y, Nakase H (2020) Autophagy and autophagy-related diseases: a review. Int J Mol Sci 21:1–21
Ikon N, Ryan RO (2017) Cardiolipin and mitochondrial cristae organization. Biochem Biophys Acta 1859:1156–1163
Imbeault E, Mahvelati TM, Braun R, Gris P, Gris D (2014) Nlrx1 regulates neuronal cell death. Mol Brain 7:90
Kim EM, Yang HS, Kang SW, Ho JN, Lee SB, Um HD (2008) Amplification of the γ-irradiation-induced cell death pathway by reactive oxygen species in human U937 cells. Cell Signal 20:916–924
Kirkin V (2009) A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol Cell 33:505–516
Kobashigawa S, Suzuki K, Yamashita S (2011) Ionizing radiation accelerates Drp1-dependent mitochondrial fission, which involves delayed mitochondrial reactive oxygen species production in normal human fibroblast-like cells. Biochem Biophys Res Commun 414:795–800
Kumar A, Tamjar J, Waddell AD, Woodroof HI, Aalten D (2017) ’Structure of PINK1 and mechanisms of Parkinson’s disease-associated mutations. Elife 6:e29985
Kuramori C, Azuma M, Kume K, Kaneko Y, Inoue A, Yamaguchi Y, Kabe Y, Hosoya T, Kizaki M, Suematsu M (2009) Capsaicin binds to prohibitin 2 and displaces it from the mitochondria to the nucleus. Biochem Biophys Res Commun 379:519–525
Lampert MA, Orogo AM, Najor RH, Hammerling BC, Leon LJ, Wang BJ, Kim T, Sussman MA, Gustafsson ÅB (2019) BNIP3L/NIX and FUNDC1-mediated mitophagy is required for mitochondrial network remodeling during cardiac progenitor cell differentiation. Autophagy 15:1182–1198
Lansman RA, Clayton DA (1975) Selective nicking of mammalian mitochondrial DNA in vivo: photosensitization by incorporation of 5-bromodeoxyuridine. J Mol Biol 99:761–776
Larsen NB, Rasmussen M, Rasmussen LJ (2005) Nuclear and mitochondrial DNA repair: similar pathways? Mitochondrion 5:89–108
Leymarie O, Meyer L, Tafforeau L, Lotteau V, Costa BD, Delmas B, Chevalier C, Le Goffic R (2017) Influenza virus protein PB1-F2 interacts with CALCOCO2 (NDP52) to modulate innate immune response. J Gen Virol 98:1196–1208
Li L, Tan J, Miao Y, Lei P, Zhang Q (2015) ROS and autophagy: interactions and molecular regulatory mechanisms. Cell Mol Neurobiol 35:615–621
Li W, He P, Huang Y, Li YF, Feng D (2020) Selective autophagy of intracellular organelles: recent research advances. Theranostics 11:222–256
Lin Q, Li S, Jiang N, Shao X, Zhang M, Jin H, Zhang Z, Shen J, Zhou Y, Zhou W (2019) PINK1-parkin pathway of mitophagy protects against contrast-induced acute kidney injury via decreasing mitochondrial ROS and NLRP3 inflammasome activation. Redox Biol 26:101254
Liu Z, Chen P, Gao H, Gu Y, Yang J, Peng H, Xu X, Wang H, Yang M, Liu X, Fan L, Chen S, Zhou J, Sun Y, Ruan K, Cheng S, Komatsu M, White E, Li L, Ji H, Finley D, Hu R (2014) Ubiquitylation of autophagy receptor Optineurin by HACE1 activates selective autophagy for tumor suppression. Cancer Cell 26:106–120
Liu J, Chen Z, Guo J, Wang L, Liu X (2017) Ambra1 induces autophagy and desensitizes human prostate cancer cells to cisplatin. Biosci Rep. https://doi.org/10.1042/BSR20170770
Lynam-Lennon N, Maher SG, Maguire A, Phelan J, Muldoon C, Reynolds JV, O’Sullivan J (2014) Altered mitochondrial function and energy metabolism is associated with a radioresistant phenotype in oesophageal adenocarcinoma. PLoS ONE 9:e100738
Makarov VI, Khmelinskii I, Khuchua Z, Javadov S (2020) In silico simulation of reversible and irreversible swelling of mitochondria: the role of membrane rigidity. Mitochondrion 50:71–81
Marsh T, Debnath J (2020) Autophagy suppresses breast cancer metastasis by degrading NBR1. Autophagy 16:1164–1165
McMillin JB, Dowhan W (2002) Cardiolipin and apoptosis. Biochem Biophys Acta 1585:97–107
Meng F, Sun N, Liu D, Jia J, Xiao J, Dai H (2021) BCL2L13: physiological and pathological meanings. Cell Mol Life Sci CMLS 78:2419–2428
Merkwirth C, Langer T (2009) Prohibitin function within mitochondria: essential roles for cell proliferation and cristae morphogenesis. Biochim Biophys Acta (BBA) Mol Cell Res 1793:27–32
Michael L, Sliter DA, Kane LA, Sarraf SA, Wang C (2015) The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524(7565):309–314
Misaka T, Murakawa T, Nishida K, Omori Y, Otsu K (2018) FKBP8 protects the heart from hemodynamic stress by preventing the accumulation of misfolded proteins and endoplasmic reticulum-associated apoptosis in mice. J Mol Cell Cardiol 114:93–104
Mishra P, Chan DC (2014) ’Mitochondrial dynamics and inheritance during cell division, development and disease. Nat Rev Mol Cell Biol 15:634–646
Morriswood B, Ryzhakov G, Puri C, Arden SD, Roberts R, Dendrou C, Kendrick-Jones J, Buss F (2007) T6BP and NDP52 are myosin VI binding partners with potential roles in cytokine signalling and cell adhesion. J Cell Sci 120:2574
Motoori S, Majima HJ, Ebara M, Kato H, Hirai F, Kakinuma S, Yamaguchi C, Ozawa T, Nagano T, Tsujii H (2001) Overexpression of mitochondrial manganese superoxide dismutase protects against radiation-induced cell death in the human hepatocellular carcinoma cell line HLE. Can Res 61:5382–5388
Murakawa T, Yamaguchi O, Hashimoto A, Hikoso S, Takeda T, Oka T, Yasui H, Ueda H, Akazawa Y, Nakayama H (2015) Bcl-2-like protein 13 is a mammalian Atg32 homologue that mediates mitophagy and mitochondrial fragmentation. Nat Commun 6:7527
Ni HM, Williams JA, Ding WX (2015) Mitochondrial dynamics and mitochondrial quality control. Redox Biol 4:6–13
Ordureau A, Sarraf S, Duda D, Heo JM, Jedrychowski M, Sviderskiy V, Olszewski J, Koerber J, Xie T, Beausoleil S (2014) Quantitative proteomics reveal a feedforward mechanism for mitochondrial PARKIN translocation and ubiquitin chain synthesis. Mol Cell 56:462–562
Orrenius S, Gogvadze V, Zhivotovsky B (2015) Calcium and mitochondria in the regulation of cell death. Biochem Biophys Res Commun 460(1):72–81
Penta JS, Johnson FM, Wachsman JT, Copeland WC (2001) Mitochondrial DNA in human malignancy. Mutat Res 488:119–133
Pfanner N, Warscheid B, Wiedemann N (2019) Mitochondrial proteins: from biogenesis to functional networks. Nat Rev Mol Cell Biol 20:267–284
Princely Abudu Y, Pankiv S, Mathai BJ, HåkonLystad A, Bindesbøll C, Brenne HB, Yoke Wui Ng M, Thiede B, Yamamoto A, MutugiNthiga T, Lamark T, Esguerra CV, Johansen T, Simonsen A (2019) NIPSNAP1 and NIPSNAP2 act as “eat me” signals for mitophagy. Dev Cell 49:509–25.e12
Pyo JO, Nah J, Kim HJ, Lee HJ, Heo J, Lee H, Jung YK (2008) Compensatory activation of ERK1/2 in Atg5-deficient mouse embryo fibroblasts suppresses oxidative stress-induced cell death. Autophagy 4:315–321
Qiu Y, Wang J, Li H, Yang B, Wang J, He Q, Weng Q (2022) Emerging views of OPTN (optineurin) function in the autophagic process associated with disease. Autophagy 18:73–85
Rasmussen NL, Kournoutis A, Lamark T, Johansen T (2022) NBR1: the archetypal selective autophagy receptor. J Cell Biol 221:e202208092
Ren Y, Yang P, Li C, Wang WA, Zhang T, Li J, Li H, Dong C, Meng W, Zhou H (2023) Ionizing radiation triggers mitophagy to enhance DNA damage in cancer cells. Cell Death Discov 9:267
Rodolfo C, Campello S, Cecconi F (2018) Mitophagy in neurodegenerative diseases. Neurochem Int 117:156–166
Sabbieti MG, Marchegiani A, Sufianov AA, Gabai VL, Shneider A, Agas D (2022) P62/SQSTM1 beyond autophagy: physiological role and therapeutic applications in laboratory and domestic animals. Life (basel, Switzerland) 12:539
Sharma V, Verma S, Seranova E, Sarkar S, Kumar D (2018) Selective autophagy and xenophagy in infection and disease. Front Cell Dev Biol 6:147
Shirane M, Nakayama KI (2003) Inherent calcineurin inhibitor FKBP38 targets Bcl-2 to mitochondria and inhibits apoptosis. Nat Cell Biol 5:28–37
Soares F, Tattoli I, Rahman MA, Robertson SJ, Belcheva A, Liu D, Streutker C, Winer S, Winer DA, Martin A, Philpott DJ, Arnoult D, Girardin SE (2014) The mitochondrial protein NLRX1 controls the balance between extrinsic and intrinsic apoptosis. J Biol Chem 289:19317–19330
Sun W-L, Wang L, Luo J, Zhu H-W, Cai Z-W (2018) Ambra1 modulates the sensitivity of breast cancer cells to epirubicin by regulating autophagy via ATG12. Cancer Sci 109:3129–3138
Sun WL, Wang L, Luo J, Zhu HW, Cai ZW (2019) Ambra1 inhibits paclitaxel-induced apoptosis in breast cancer cells by modulating the Bim/mitochondrial pathway. Neoplasma 66:377–385
Takeda M, Koseki J, Takahashi H, Miyoshi N, Nishida N, Nishimura J, Hata T, Matsuda C, Mizushima T, Yamamoto H, Ishii H, Doki Y, Mori M, Haraguchi N (2019) Disruption of endolysosomal RAB5/7 efficiently eliminates colorectal cancer stem cells. Can Res 79:1426–1437
Turrens JF (2003) Mitochondrial formation of reactive oxygen species. J Physiol 552:3335–3344
Venardos KM, Perkins A, Headrick J, Kaye DM (2007) Myocardial ischemia-reperfusion injury, antioxidant enzyme systems, and selenium: a review. Curr Med Chem 14:1539–1549
Verstrepen L, Verhelst K, Carpentier I, Beyaert R (2011) TAX1BP1, a ubiquitin-binding adaptor protein in innate immunity and beyond. Trends Biochem Sci 36:347–354
Wei Y, Chiang WC, Sumpter R, Mishra P, Levine B (2017) Prohibitin 2 is an inner mitochondrial membrane mitophagy receptor. Cell 168:224–238.e10. https://doi.org/10.1016/j.cell.2016.11.042
West AP, Shadel GS (2017) Mitochondrial DNA in innate immune responses and inflammatory pathology. Nat Rev Immunol 17(6):363–375
Wu S, Li Z, Li H, Liao K (2022) Dihydroartemisinin reduces irradiation-induced mitophagy and radioresistance in lung cancer A549 cells via CIRBP inhibition. Life (basel, Switzerland) 12:1129
Yakes FM, Houten BV (1997) Mitochondrial DNA damage is more extensive and persists longer than nuclear DNA damage in human cells following oxidative stress. Proc Natl Acad Sci USA 94(2):514–519
Yamamoto K, Venida A, Yano J, Biancur DE, Kakiuchi M, Gupta S, Sohn ASW, Mukhopadhyay S, Lin EY, Parker SJ, Banh RS, Paulo JA, Wen KW, Debnath J, Kim GE, Mancias JD, Fearon DT, Perera RM, Kimmelman AC (2020) Autophagy promotes immune evasion of pancreatic cancer by degrading MHC-I. Nature 581:100–105
Yan C, Gong L, Chen L, Xu M, Abou-Hamdan H, Tang M, Désaubry L, Song Z (2020) PHB2 (prohibitin 2) promotes PINK1-PRKN/Parkin-dependent mitophagy by the PARL-PGAM5-PINK1 axis. Autophagy 16:419–434
Yang J, Wu LJ, Tashino S, Onodera S, Ikejima T (2008) Reactive oxygen species and nitric oxide regulate mitochondria-dependent apoptosis and autophagy in evodiamine-treated human cervix carcinoma HeLa cells. Free Radic Res 42:492–504
Yang P, Luo X, Li J, Zhang T, Gao X, Hua J, Li Y, Ding N, He J, Zhang Y, Wei W, Wang J, Zhou H (2021) Ionizing radiation upregulates glutamine metabolism and induces cell death via accumulation of reactive oxygen species. Oxid Med Cell Longev 2021:5826932
Yoshida T, Goto S, Kawakatsu M, Urata Y, Li TS (2012) Mitochondrial dysfunction, a probable cause of persistent oxidative stress after exposure to ionizing radiation. Free Radic Res 46:147–153
Yu L, Wan F, Dutta S, Welsh S, Liu Z, Freundt E, Baehrecke EH, Lenardo M (2006) Autophagic programmed cell death by selective catalase degradation. Proc Natl Acad Sci USA 103:4952–4957
Yu L, Yang X, Li X, Qin L, Xu W, Cui H, Jia Z, He Q, Wang Z (2021) Pink1/PARK2/mROS-dependent mitophagy initiates the sensitization of cancer cells to radiation. Oxid Med Cell Longev 2021:5595652
Yuehua W, Chen Lulu Xu, Hui XC, Yunfeng Z, Fuxiang Z (2018) Mitochondrial dysfunctions regulated radio resistance through mitochondria-to-nucleus retrograde signaling pathway of NF-κB/PI3K/AKT2/mTOR. Radiat Res 190:204–215
Zhang J, Ney PA (2009) Role of BNIP3 and NIX in cell death, autophagy, and mitophagy. Cell Death Differ 16:939–946
Zhang B, Davidson MM, Zhou H, Wang C, Walker WF, Hei TK (2013) Cytoplasmic irradiation results in mitochondrial dysfunction and DRP1-dependent mitochondrial fission. Can Res 73:6700–6710
Zhihong C, Li Y, Zhang C, Yi H, Wu C (2013) Downregulation of Beclin1 and impairment of autophagy in a small population of colorectal cancer. Digest Dis Sci 58(10):2887–2894
Zhou X, Li N, Wang Y, Wang Y, Zhang X, Zhang H (2011) Effects of X-irradiation on mitochondrial DNA damage and its supercoiling formation change. Mitochondrion 11:886–892
Zorov DB, Juhaszova M, Sollott SJ (2014) Mitochondrial reactive oxygen species (ROS) and ROS-induced ROS release. Physiol Rev 94:909–950
Acknowledgements
Not applicable.
Funding
Shandong Provincial Medical and Health Science and Technology Development Program, Fund No. 202209030786.
Author information
Authors and Affiliations
Contributions
WQ: writing and editing; LCX: monitoring and review. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose. The authors declare no competing interests.
Ethics approval
This is a systematic review. The Research Ethics Committee has confirmed that no ethical approval is required.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Q., Liu, C. Mitophagy plays a “double-edged sword” role in the radiosensitivity of cancer cells. J Cancer Res Clin Oncol 150, 14 (2024). https://doi.org/10.1007/s00432-023-05515-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00432-023-05515-2