Abstract
Introduction
EGFR mutations occur most frequently in patients with lung adenocarcinoma in East Asia. However, the prognostic and therapeutic impact of co-mutational status of EGFR and tumor suppressor genes is not fully understood. This study aims to provide a deeper understanding of lung adenocarcinoma patients with co-mutation of EGFR and tumor suppressor genes.
Methods
From November 2009 to May 2016, 675 patients with lung adenocarcinoma who underwent complete surgery were included in this study. Samples were collected and pathologically examined. Whole-exome sequencing was performed on 197 samples, while direct sequencing of major driver genes, including EGFR, KRAS, ERBB2 and BRAF and Ion-torrent targeted sequencing of tumor suppressor genes, including TP53, KEAP1, MGA, NF1, RB1, SMARCA4 and STK11, were performed on 478 samples. Tumor mutational burden was calculated and survival analyses were performed.
Results
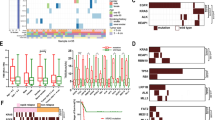
The frequency of EGFR and TP53 mutation was 409 (60.6%) and 215 (31.9%), respectively. Co-mutation of EGFR and TP53 occured in 151 patients (22.4%), while co-mutation of EGFR and at least one tumor suppressor gene occured in 184 patients (27.3%). Compared with patients with only EGFR mutations, patients with co-mutations of EGFR and TP53 had a higher tumor mutational burden (p = 0.007) and worse recurrence-free survival (p = 0.010), while patients with co-mutations of EGFR and at least one tumor suppressor gene had a higher tumor mutational burden (p = 0.007), worse recurrence-free survival (p = 0.016) and worse overall survival (p = 0.018).
Conclusions
Lung adenocarcinoma patients harboring EGFR and co-mutational tumor suppressor genes should be regarded as a unique subgroup.



Similar content being viewed by others
References
Arbour KC, Jordan E, Kim HR et al (2018) Effects of co-occurring genomic alterations on outcomes in patients with KRAS-mutant non-small cell lung cancer. Clin Cancer Res 24(2):334–340
Borghaei H, Paz-Ares L, Horn L et al (2015) Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N Engl J Med 373:1627–1639
Brahmer J, Reckamp KL, Baas P et al (2015) Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N Engl J Med 373:123–135
Campbell JD, Alexandrov A, Kim J et al (2016) Distinct patterns of somatic genome alterations in lung adenocarcinomas and squamous cell carcinomas. Nat Genet 48(6):607–616
Cancer Genome Atlas Research Network (2012) Comprehensive genomic characterization of squamous cell lung cancers. Nature 489(7417):519–525
Cancer Genome Atlas Research Network (2014) Comprehensive molecular profiling of lung adenocarcinoma. Nature 511:543–550
Christopoulos P, Kirchner M, Bozorgmehr F et al (2019) Identification of a highly lethal V3+ TP53+ subset in ALK+ lung adenocarcinoma. Int J Cancer 144(1):190–199
Cibulskis K, Lawrence MS, Carter SL et al (2013) Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 31(3):213–219
DePristo MA, Banks E, Poplin R et al (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43(5):491–498
Dong ZY, Zhong WZ, Zhang XC et al (2017) Potential predictive value of TP53 and KRAS mutation status for response to PD-1 blockade immunotherapy in lung adenocarcinoma. Clin Cancer Res 23(12):3012–3024
Fehrenbacher L, Spira A, Ballinger M et al (2016) Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicenter, open-label, phase 2 randomised controlled trial. Lancet 378:1837–1846
Garon EB, Rizvi NA, Hui R et al (2015) Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med 372:2018–2028
Hellmann MD, Ciuleanu TE, Pluzanski A et al (2018) Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. N Engl J Med 378:2093–2104
Hellyer JA, Stehr H, Das M et al (2019) Impact of KEAP1/NFE2L2/CUL3 mutations on duration of response to EGFR tyrosine kinase inhibitors in EGFR mutated non-small cell lung cancer. Lung Cancer 134:42–45
Izar B, Sequist L, Lee M et al (2013) The impact of EGFR mutation status on outcomes in patients with resected stage I non-small cell lung cancers. Ann Thorac Surg 96(3):962–968
Janne PA, Yang JC, Kim DW et al (2015) AZD9291 in EGFR inhibitor-resistant non-small-cell lung cancer. N Engl J Med 372:1689–1699
Jorge SE, Lucena-Araujo AR, Yasuda H et al (2018) EGFR Exon 20 insertion mutations display sensitivity to Hsp90 inhibition in preclinical models and lung adenocarcinomas. Clin Cancer Res 24:6548–6555
Kohno T, Ichikawa H, Totoki Y et al (2012) KIF5B-RET fusions in lung adenocarcinoma. Nat Med 18:375–377
La Fleur L, Falk-Sorqvist E, Smeds P et al (2019) Mutation patterns in a population-based non-small cell lung cancer cohort and prognostic impact of concomitant mutations in KRAS and TP53 or STK11. Lung Cancer 130:50–58
Lee CK, Wu YL, Ding PN et al (2015) Impact of specific epidermal growth factor receptor (EGFR) mutations and clinical characteristics on outcomes after treatment with EGFR tyrosine kinase inhibitors versus chemotherapy in EGFR-mutant lung cancer: a meta-analysis. J Clin Oncol 33:1958–1965
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv:1303.3997v2
Lynch TJ, Bell DW, Sordella R et al (2004) Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 350:2129–2139
Ma X, Le Teuff G, Lacas B et al (2016) Prognostic and predictive effect of TP53 mutations in patients with non-small cell lung cancer from adjuvant cisplatin-based therapy randomized trials: a LACE-Bio pooled analysis. J Thorac Oncol 11(6):850–861
Ramos AH, Lichtenstein L, Gupta M et al (2015) Oncotator: cancer variant annotation tool. Hum Mutat 36(4):E2423–2429
Reck M, Rodriguez-Abreu D, Robinson AG et al (2016) Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N Engl J Med 375:1823–1833
Rittmeyer A, Barlesi F, Waterkamp D et al (2017) Atezolizumab versus docetaxel in patients with previously treated non-small-cell lung cancer (OAK): a phase 3, open-label, multicentre randomised controlled trial. Lancet 389:255–265
Rizvi NA, Hellmann MD, Snyder A et al (2015) Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348:124–128
Saunders CT, Wong WS, Swamy S et al (2012) Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28(14):1811–1817
Shepherd FA, Lacas B, Le Teuff G et al (2017) Pooled analysis of the prognostic and predictive effects of TP53 comutation status combined with KRAS or EGFR mutation in early-stage resected non-small-cell lung cancer in four trials of adjuvant chemotherapy. J Clin Oncol 35(18):2018–2027
Siegel RL, Miller KD, Jemal A (2019) Cancer statistics, 2019. CA Cancer J Clin 69:7–34
Travis WD, Brambilla E, Noguchi M et al (2011) International association for the study of lung cancer/American Thoracic Society/European respiratory society international multidisciplinary classification of lung adenocarcinoma. J Thorac Oncol 6(2):244–285
VanderLaan PA, Rangachari D, Mockus SM et al (2017) Mutations in TP53, PIK3CA, PTEN and other genes in EGFR mutated lung cancers: correlation with clinical outcomes. Lung Cancer 106:17–21
Vogelstein B, Lane D, Levine AJ (2000) Surfing the p53 network. Nature 408(6810):307–310
Wang R, Zhang Y, Pan Y et al (2015) Comprehensive investigation of oncogenic driver mutations in Chinese non-small cell lung cancer patients. Oncotarget 6:34300–34308
Wu K, Zhang X, Li F et al (2015) Frequent alterations in cytoskeleton remodelling genes in primary and metastatic lung adenocarcinomas. Nat Commun 6:10131
Yang JC, Sequist LV, Geater SL et al (2015) Clinical activity of afatinib in patients with advanced non-small-cell lung cancer harbouring uncommon EGFR mutations: a combined post-hoc analysis of LUX-Lung 2, LUX-Lung 3, and LUX-Lung 6. Lancet Oncol 16:830–838
Yasuda H, Kobayashi S, Costa DB (2012) EGFR exon 20 insertion mutations in non-small-cell lung cancer: preclinical data and clinical implications. Lancet Oncol 13:e23–31
Yasuda H, Park E, Yun CH et al (2013) Structural, biochemical, and clinical characterization of epidermal growth factor receptor (EGFR) exon 20 insertion mutations in lung cancer. Sci Transl Med 5:216ra177
Funding
This work was supported by the National Natural Science Foundation of China (Grant numbers: 81572253 and 81930073); Shanghai Shen Kang Hospital Development Center City Hospital Emerging Cutting-edge Technology Joint Research Project (Grant number: SHDC12017102); Shanghai Municipal Health Commission Key Discipline Project (Grant numbers: 2017ZZ02025 and 2017ZZ01019).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
These authors declare no conflicts of interest.
Ethical approval
This study was approved by the Committee for Ethical Review of Research (Fudan University Shanghai Cancer Center IRB# 090977-1).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.

432_2020_3237_MOESM2_ESM.jpg
Supplementary figure 1. Pie charts showing the co-mutational composition of each driver gene with every tumor suppressor gene in lung adenocarcinoma. Composition of each driver gene is shown in patients with mutations in a) at least one tumor suppressor gene, b) TP53, c) MGA, d) NF1, e) RB1, f) SMARCA4, g) STK11 and h) KEAP1. (JPG 2108 kb)
Rights and permissions
About this article
Cite this article
Zhao, Y., Pan, Y., Cheng, C. et al. EGFR-mutant lung adenocarcinoma harboring co-mutational tumor suppressor genes predicts poor prognosis. J Cancer Res Clin Oncol 146, 1781–1789 (2020). https://doi.org/10.1007/s00432-020-03237-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-020-03237-3