Abstract
Main conclusion
Soybean possesses 19 CMF genes which mainly arose from duplication events. Their features and motifs are highly conserved but transcriptional data indicated functional diversity in metabolism and stress responses.
Abstract
CCT [for CONSTANS, CONSTANS-like (CO-like), and timing of CAB expression1 (TOC1)] domain-containing genes play important roles in regulating flowering, plant growth, and grain yield and are also involved in stress responses. The CMF (CCT motif family) genes, included in the CCT family, contain a single CCT domain as the only identifiable domain in their predicted protein sequence and are interesting targets for breeding programs. In this study, we identified 19 putative GmCMF genes, based on the latest soybean (Glycine max) genome annotation. The predicted GmCMF proteins were characterized based on conserved structural features, and a phylogenetic tree was constructed including all CMF proteins from rice and Arabidopsis as representative examples of the monocotyledonous (monocot) and dicotyledonous (dicot) plants, respectively. High similarities in the conserved motifs of the protein sequences and the gene structures were found. In addition, by analyzing the CMF gene family in soybean, we identified seven pairs of genes that originated from segmental chromosomal duplication events attributable to the most recent whole-genome duplication (WGD) event in the Glycine lineage. Expression analysis of GmCMF genes in various tissues and after specific treatments demonstrated tissue and stress-response specific differential expression. Gene expression analysis was complemented by the identification of putative cis-elements present in the promoter regions of the genes through a bioinformatics approach, using the existing soybean reference genome sequence and gene models. Co-functional networks inferred from distinct types of genomics data—including microarrays and RNA-seq samples from soybean—revealed that GmCMF genes might play crucial roles in metabolism and transport processes. The results of this study, the first systematic analysis of the soybean CCT gene family, can serve as a strong foundation for further elucidation of their physiological functions and biological roles.






Similar content being viewed by others
Availability of data and materials
The data generated or analyzed in this study are included in this article and its Additional materials. All sequence information regarding soybean is available at a public database, SoyBase (http://www.soybase.org/). AtCMF, OsCMF proteins and the expression data of GmCMFs were retrieved from Phytozome V12.1 database (https://www.phytozome.jgi.doe.gov/pz/portal.html). Duplication blocks were identified using the PGDD database (http://www.chibba.agtec.uga.edu/duplication/), and co-functional genes were retrieved from SoyNet database (http://www.inetbio.org/soynet/).
Abbreviations
- CCT:
-
CONSTANS, CONSTANS-like (CO-like), and timing of CAB expression1 (TOC1)
- CIA2:
-
CHLOROPLAST IMPORT APPARATUS 2
- CMF:
-
CCT motif family
- CO-like:
-
CONSTANS-LIKE
- Ka/Ks:
-
Non-synonymous/synonymous substitution
- MYA:
-
Millions of years ago
- PRR:
-
PSEUDORESPONSE REGULATOR
- WGD:
-
Whole-genome duplication
References
Alabadi D, Oyama T, Yanovsky MJ, Harmon FG, Mas P, Kay SQ (2001) Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 293:880–883
Altschul S, Gish W, Miller W, Myers E, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208. https://doi.org/10.1093/nar/gkp335
Bowers JE, Chapman BA, Rong J, Paterson AH (2003) Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events. Nature 422:433–438
Brambilla V, Fornara F (2017) Y flowering? Regulation and activity of CONSTANS and CCT-domain proteins in Arabidopsis and crop species. Biochim Biophys Acta 1860:655–660
Brown AV, Hudson KA (2015) Developmental profiling of gene expression in soybean trifoliate leaves and cotyledons. BMC Plant Biol 15:169. https://doi.org/10.1186/s12870-015-0553-y
Cannon SB, Shoemaker RC (2012) Evolutionary and comparative analyses of the soybean genome. Breed Sci 61:437–444. https://doi.org/10.1270/jsbbs.61.437
Cannon SB, Mitra A, Baumgarten A, Young AD, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:10. https://doi.org/10.1186/1471-2229-4-10
Chao J, Yingzhen K, Wang Q, Yuhe S, Daping G, Lv J, Guanshan L (2015) MapGene2Chrom, a tool to draw gene physical map based on Perl and SVG languages. Hereditas (Beijing) 37:91–97
Cheong W-H, Tan Y-C, Yap S-J, Ng K-P (2015) ClicO FS: an interactive web-based service of Circos. Bioinformatics 31:3685–3687. https://doi.org/10.1093/bioinformatics/btv433
Chia TYP, Mueller A, Jung C, Mutasa-Goettgens ES (2008) Sugar beet contains a large CONSTANS-LIKE gene family including a CO homologue that is independent of the early-bolting (B) gene locus. J Exp Bot 59:2735–2748. https://doi.org/10.1093/jxb/ern129
Cho C-W, Lee H-J, Chung E, Kim KM, Heo JE, Kim J-I, Chung J, Ma Y, Fukui K, Lee D-W, Kim D-H, Lee C-S, J-H, (2007) Molecular characterization of the soybean l-asparaginase gene induced by low temperature stress. Mol Cells 23:280–286
Cockram J, Thiel T, Steuernagel B, Stein N, Taudien S, Bailey PC, O’Sullivan DM (2012) Genome dynamics explain the evolution of flowering time CCT domain gene families in the Poaceae. PLoS ONE 7:e45307. https://doi.org/10.1371/journal.pone.0045307
Cusack BP, Wolfe KH (2007) When gene marriages don’t work out: divorce by subfunctionalization. Trends Genet 23:270–272
Denison FC, Paul AL, Zupanska AK, Ferl RJ (2011) 14-3-3 proteins in plant physiology. Semin Cell Dev Biol 22:720–727
Di Rienzo JA, Macchiavelli R, Casanoves F (2017) Modelos lineales generalizados mixtos en InfoStat. Edición electrónica, p 101. ISBN 978-987-42-4985-2
Dubcovsky J, Loukoianov A, Fu D, Valarik M, Sanchez A, Yan L (2006) Effect of photoperiod on the regulation of wheat vernalization genes VRN1 and VRN2. Plant Mol Biol 60:469–480. https://doi.org/10.1007/s11103-005-4814-2
Dunn MJ, Kinney GM, Washington PM, Berman J, Anderson MZ (2018) Functional diversification accompanies gene family expansion of MED2 homologs in Candida albicans. PLoS Genet 14:e1007326
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Farré EM, Liu T (2013) The PRR family of transcriptional regulators reflects the complexity and evolution of plant circadian clocks. Curr Opin Plant Biol 16:621–629
Gasteiger E (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788. https://doi.org/10.1093/nar/gkg563
Gawroński P, Burdiak P, Scharff LB, Mielecki J, Zaborowska M, Waszczak C, Karpiński S (2019) Dual-targeted transcription factors are required for optimal photosynthesis and stress responses in Arabidopsis thaliana. bioRxiv. https://doi.org/10.1101/793968
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186. https://doi.org/10.1093/nar/gkr944
Graham PH, Vance CP (2003) Legumes: importance and constraints to greater use. Plant Physiol 131:872–877
Grant D, Nelson RT, Cannon SB, Shoemaker RC (2009) SoyBase, the USDA-ARS soybean genetics and genomics database. Nucleic Acids Res 38(Suppl. 1):D843-846. https://doi.org/10.1093/nar/gkp798
Griffiths S, Dunford RP, Coupland G, Laurie DA (2003) The evolution of CONSTANS-like gene families in barley, rice, and Arabidopsis. Plant Physiol 131:1855–1867
Hemming MN, Peacock WJ, Dennis ES, Trevaskis B (2008) Low-temperature and daylength cues are integrated to regulate FLOWERING LOCUS T in barley. Plant Physiol 147:355–366
Howe E, Holton K, Nair S, Schlauch D, Sinha R, Quackenbush J (2010) MeV: MultiExperiment viewer. Biomedical informatics for cancer research. Springer US, New York, pp 267–277
Hu B, Jin J, Guo A-Y, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Kanna R, Kronmiller B, Maszle DR, Coupland G, Holm M, Mizuno T, Wu S-H (2009) The Arabidopsis B-box zinc finger family. Plant Cell 21:3416–3420
Kim JA, Kim JS, Hong JK, Lee YH, Choi B-S, Seol Y-J, Jeon CH (2012) Comparative mapping, genomic structure, and expression analysis of eight pseudo-response regulator genes in Brassica rapa. Mol Genet Genom 287:373–388. https://doi.org/10.1007/s00438-012-0682-z
Kim DY, Kwon SI, Choi C, Lee H, Ahn I, Park SR, Bae SC, Lee SC, Hwang DJ (2013a) Expression analysis of rice VQ genes in response to biotic and abiotic stresses. Gene 529:208–214. https://doi.org/10.1016/j.gene.2013.08.023
Kim MY, Kang YJ, Lee T, Lee SH (2013b) Divergence of flowering-related genes in three legume species. Plant Genome 6:4. https://doi.org/10.3835/plantgenome2013.03.0008
Kim E, Hwang S, Lee I (2017) SoyNet: a database of co-functional networks for soybean Glycine max. Nucleic Acids Res 45:D1082–D1089. https://doi.org/10.1093/nar/gkw704
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645. https://doi.org/10.1101/gr.092759.109
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lamesch P, Berardini TZ, Li D, Swarbreck D, Wilks C, Sasidharan R, Muller R, Dreher K, Alexander DL, Garcia-Hernandez M, Karthikeyan AS, Lee CH, Nelson WD, Ploetz L, Singh S, Wensel A, Huala E (2012) The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res 40(Database issue):D1202–D1210
Lang D, Ullrich KK, Murat F, Fuchs J, Jenkins J, Haas FB, Piednoel M, Gundlach H, Van Bel M, Meyberg R, Vives C, Morata J, Symeonidi A, Hiss M, Muchero W et al (2018) The Physcomitrella patens chromosome-scale assembly reveals moss genome structure and evolution. Plant J 93:515–533
Le DT, Nishiyama R, Watanabe Y, Tanaka M, Seki M, Ham LH, Yamaguchi-Shinozaki K, Shinozaki K, Tran L-SP (2012) Differential gene expression in soybean leaf tissues at late developmental stages under drought stress revealed by genome-wide transcriptome analysis. PLoS ONE 7:e48522. https://doi.org/10.1371/journal.pone.0049522.t001
Lee T-H, Tang H, Wang X, Paterson AH (2012) PGDD: a database of gene and genome duplication in plants. Nucleic Acids Res 41:D1152–D1158. https://doi.org/10.1093/nar/gks1104
Lee T, Yang S, Kim E, Ko Y, Hwang S, Shin J, Shim JE, Shim H, Kim H, Kim C, Lee I (2015) AraNet v2: an improved database of co-functional gene networks for the study of Arabidopsis thaliana and 27 other nonmodel plant species. Nucleic Acids Res 43:D996–D1002. https://doi.org/10.1093/nar/gku1053
Lescot M (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327. https://doi.org/10.1093/nar/30.1.325
Letunic I, Bork P (2019) Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47:W256–W259. https://doi.org/10.1093/nar/gkz239
Li WH, Gojobori T, Nei M (1981) Pseudogenes as a paradigm of neutral evolution. Nature 292:237–239. https://doi.org/10.1038/292237a0
Libault M, Farmer A, Joshi T, Takahashi K, Langley RJ, Franklin LD, He J, Xu D, May G, Stacey G (2010) An integrated transcriptome atlas of the crop model Glycine max, and its use in comparative analyses in plants. Plant J 63:86–99. https://doi.org/10.1111/j.1365-313X.2010.04222.x
Liu T, Liu H, Zhang H, Xing Y (2013) Validation and characterization of Ghd7.1, a major quantitative trait locus with pleiotropic effects on spikelets per panicle, plant height, and heading date in rice (Oryza sativa L.). J Integr Plant Biol 55:917–927. https://doi.org/10.1111/jipb.12070
Liu J, Shen J, Xu Y, Li X, Xiao J, Xiong L (2016) Ghd2, a CONSTANS-like gene, confers drought sensitivity through regulation of senescence in rice. J Exp Bot 67:5785–5798
Liu H, Zhou X, Li Q, Wang L, Xing Y (2020) CCT domain-containing genes in cereal crops: flowering time and beyond. Theor Appl Genet 133(5):1385–1396. https://doi.org/10.1007/s00122-020-03554-8
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lu S, Dong L, Fang C et al (2020) Stepwise selection on homeologous PRR genes controlling flowering and maturity during soybean domestication. Nat Genet 52:428–436. https://doi.org/10.1038/s41588-020-0604-7
Ma X, Zhang Y, Turečková V, Xue GP, Fernie AR, Mueller-Roeber B, Balazadeh S (2018) The NAC transcription factor SLNAP2 regulates leaf senescence and fruit yield in tomato. Plant Physiol 177:1286–1302. https://doi.org/10.1104/pp.18.00292
Masaki T, Tsukagoshi H, Mitsui N, Nishii T, Hattori T, Morikami A, Nakamura K (2005) Activation tagging of a gene for a protein with novel class of CCT-domain activates expression of a subset of sugar-inducible genes in Arabidopsis thaliana. Plant J 43:142–152. https://doi.org/10.1111/j.1365-313X.2005.02439.x
Merchant SS, Prochnik SE, Vallon O et al (2007) The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318(5848):245–250
Murphy RL, Klein RR, Morishige DT, Brady JA, Rooney WL, Miller FR, Dugas DV, Klein PE, Mullet JE (2011) Coincident light and clock regulation of pseudoresponse regulator protein 37 (PRR37) controls photoperiodic flowering in sorghum. Proc Natl Acad Sci USA 108:16469–16474
Nakamichi N, Kiba T, Henriques R, Mizuno T, Chua N-H, Sakakibara N (2010) PSEUDO-RESPONSE REGULATORS 9, 7, and 5 are transcriptional repressors in the Arabidopsis circadian clock. Plant Cell 22:594–605
Navarro A, Barton NH (2003) Chromosomal speciation and molecular divergence—accelerated evolution in rearranged chromosomes. Science 300:321–324. https://doi.org/10.1126/science.1080600
One Thousand Plant Transcriptomes Initiative (2019) One thousand plant transcriptomes and the phylogenomics of green plants. Nature 574:679–685. https://doi.org/10.1038/s41586-019-1693-2
Osella AV, Mengarelli DA, Mateos J, Dong S, Yanovsky MJ, Balazadeh S, Valle EM, Zanor MI (2018) FITNESS, a CCT domain-containing protein, deregulates reactive oxygen species levels and leads to fine-tuning trade-offs between reproductive success and defence responses in Arabidopsis. Plant Cell Environ 41:2328–2341. https://doi.org/10.1111/pce.13354
Ouyang S, Zhu W, Hamilton J, Lin H, Campbell M, Childs K, Thibaud-Nissen F, Malek RL, Lee Y, Zheng L, Orvis J, Haas B, Wortman J, Buell CR (2007) The TIGR rice genome annotation resource: improvements and new features. Nucleic Acids Res 35(Database issue):D883–D887
Panchy N, Lehti-Shiu M, Shiu SH (2016) Evolution of gene duplication in plants. Plant Physiol 171:2294–2316. https://doi.org/10.1104/pp.16.00523
Patil G, Valliyodan B, Deshmukh R, Prince S, Nicander B, Zhao M, Sonah H, Song L, Lin L, Chaudhary J, Liu Y, Joshi T, Xu D, Nguyen HT (2015) Soybean (Glycine max) SWEET gene family: insights through comparative genomics, transcriptome profiling and whole genome re-sequence analysis. BMC Genom 16:520. https://doi.org/10.1186/s12864-015-1730-y
Putterill J, Robson F, Lee K, Simon R, Coupland G (1995) The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc finger transcription factors. Cell 80:847–857. https://doi.org/10.1016/0092-8674(95)90288-0
Robson F, Costa MMR, Hepworth SR, Vizir I, Pineiro M, Reeves PH, Putterill J, Coupland G (2002) Functional importance of conserved domains in the flowering-time gene CONSTANS demonstrated by analysis of mutant alleles and transgenic plants. Plant J 28:619–631
Salomé PA, Merchant SS (2019) A series of fortunate events: introducing Chlamydomonas as a reference organism. Plant Cell 31:1682–1707
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T et al (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183. https://doi.org/10.1038/nature08670
Severin AJ, Cannon SB, Graham MM, Grant D, Shoemaker RC (2011) Changes in twelve homoeologous genomic regions in soybean following three rounds of polyploidy. Plant Cell 23:3129–3136. https://doi.org/10.1105/tpc.111.089573
Sieciechowicz KA, Joy KW, Ireland RJ (1988) The metabolism of asparagine in plants. Phytochemistry 27:663–671
Song L, Nguyen N, Deshmukh RK, Patil GB, Prince SJ, Valliyodan B, Mutava R, Pike SM, Gassmann W, Nguyen HT (2016) Soybean TIP gene family analysis and characterization of GmTIP1;5 and GmTIP2;5 water transport activity. Front Plant Sci 7:1564. https://doi.org/10.3389/fpls.2016.01564
Strayer C, Oyama T, Schultz TF, Raman R, Somers DE, Mas P, Panda S, Kreps JA, Kay SA (2000) Cloning of the Arabidopsis clock gene TOC1, an autoregulatory response regulator homolog. Science 289:768–771
Sun CW, Huang YC, Chang HY (2009) CIA2 coordinately up-regulates protein import and synthesis in leaf chloroplasts. Plant Physiol 150:879–888. https://doi.org/10.1104/pp.109.137240
Tiwari SB, Shen Y, Chang HC, Hou Y, Harris A, Ma SF, McPartland M, Hymus GJ, Adam L, Marion C, Belachew A, Repetti PP, Reuber TL, Ratcliffe OJ (2010) The flowering time regulator CONSTANS is recruited to the FLOWERING LOCUS T promoter via a unique cis-element. New Phytol 187:57–66. https://doi.org/10.1111/j.1469-8137.2010.03251.x
Turner A, Beales J, Faure S, Dunford RP, Laurie DA (2005) The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley. Science 310:1031–1034
Van Bel M, Diels T, Vancaester E, Kreft L, Botzki A, Van de Peer Y, Coppens F, Vandepoele K (2018) PLAZA 4.0: an integrative resource for functional, evolutionary and comparative plant genomics. Nucleic Acids Res 46:D1190–D1196. https://doi.org/10.1093/nar/gkx1002
Walling JG, Shoemaker R, Young N, Mudge J, Jackson S (2006) Chromosome-level homeology in paleopolyploid soybean (Glycine max) revealed through integration of genetic and chromosome maps. Genetics 172:1893–1900
Wan Q, Chen S, Shan Z, Yang Z, Chen L, Zhang C, Yuan S, Hao Q, Zhang X, Qiu D, Chen H, Zhou X (2017) Stability evaluation of reference genes for gene expression analysis by RT-qPCR in soybean under different conditions. PLoS ONE 12:e0189405. https://doi.org/10.1371/journal.pone.0189405
Wang Y, Tang H, DeBarry JD, Tan X, Li J, Wang X, Lee T-H, Jin H, Marler B, Guo H, Kissinger JC, Paterson AH (2012) MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res 40:e49–e49. https://doi.org/10.1093/nar/gkr1293
Wang QX, Xie WB, Xing HK, Yan J, Meng XZ, Li XL, Fu XK, Xu JY, Lian XM, Yu SB, Xing YZ, Wang GW (2015) Genetic architecture of natural variation in rice chlorophyll content revealed by a genome-wide association study. Mol Plant 8:946–957
Wang W, Jiang W, Liu J, Li Y, Gai J, Li Y (2017) Genome-wide characterization of the aldehyde dehydrogenase gene superfamily in soybean and its potential role in drought stress response. BMC Genom 18:1–17. https://doi.org/10.1186/s12864-017-3908-y
Weng X, Wang L, Wang J, Hu Y, Du H, Xu C, Xing Y, Li X, Xiao J, Zhang Q (2014) Grain Number, Plant Height, and Heading Date7 is a central regulator of growth, development, and stress response. Plant Physiol 164:735–747. https://doi.org/10.1104/pp.113.231308
Wenkel S, Turck F, Singer K, Gissot L, Le Gourrierec J, Samach A, Coupland G (2006) CONSTANS and the CCAAT box binding complex share a functionally important domain and interact to regulate flowering of Arabidopsis. Plant Cell 18:2971–2984
Woods DP, McKeown MA, Dong Y, Preston JC, Amasino RM (2016) Evolution of VRN2/Ghd7-like genes in vernalization-mediated repression of grass flowering. Plant Physiol 170:2124–2135
Wu F, Price BW, Haider W, Seufferheld G, Nelson R, Hanzawa Y (2014) Functional and evolutionary characterization of the CONSTANS gene family in short-day photoperiodic flowering in soybean. PLoS ONE 9:e85754. https://doi.org/10.1371/journal.pone.0085754
Xu G, Guo C, Shan H, Kong H (2012) Divergence of duplicate genes in exon-intron structure. Proc Natl Acad Sci USA 109:1187–1192. https://doi.org/10.1073/pnas.1109047109
Xue W, Xing Y, Weng X, Zhao Y, Tang W, Wang L, Zhou H, Yu S, Xu C, Li X, Zhang Q (2008) Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40:761–767. https://doi.org/10.1038/ng.143
Yan L, Loukoianov A, Blechl A, Tranquilli G, Ramakrishna W, SanMiguel P, Bennetzen JL, Echenique V, Dubcovsky J (2004) The wheat VRN2 gene is a flowering repressor downregulated by vernalization. Science 303:1640–1644. https://doi.org/10.1126/science.10943
Yang Q, Li Z, Li W, Ku L, Wang C, Ye J, Li K, Yang N, Li Y, Zhong T, Li J, Chen Y, Yan J, Yang X, Xu M (2013) CACTA-like transposable element in ZmCCT attenuated photoperiod sensitivity and accelerated the postdomestication spread of maize. Proc Natl Acad Sci USA 110:16969–16974. https://doi.org/10.1073/pnas.1310949110
Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y, Sasaki T (2000) Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12:2473–2483. https://doi.org/10.1105/tpc.12.12.2473
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinform 13:134. https://doi.org/10.1186/1471-2105-13-134
Ye J, Niu X, Yang Y, Wang S, Xu Q, Yuan X, Yu H, Wang Y, Wang S, Feng Y, Wei X (2018) Divergent Hd1, Ghd7, and DTH7 alleles control heading date and yield potential of Japonica rice in Northeast China. Front Plant Sci 9:35. https://doi.org/10.3389/fpls.2018.00035
Zhang L, Li Q, Dong H, He Q, Liang L, Tan C, Han Z, Yao W, Li G, Zhao H, Xie W, Xing Y (2015) Three CCT domain-containing genes were identified to regulate heading date by candidate gene-based association mapping and transformation in rice. Sci Rep 5:7663. https://doi.org/10.1038/srep07663
Zheng X, Li X, Ge C, Chang J, Shi M, Chen J, Qiao L, Chang Z, Zheng J, Zhang J (2017) Characterization of the CCT family and analysis of gene expression in Aegilops tauschii. PLoS ONE 12(12):e0189333. https://doi.org/10.1371/journal.pone.0189333
Acknowledgements
D. A. M. is recipient of a fellowship of Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET). M. I. Z. is a career member of CONICET.
Funding
This work was supported by Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT, PICT2017-1301).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
425_2020_3537_MOESM1_ESM.pdf
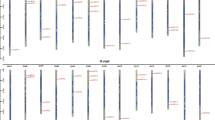
Supplementary file1 Supplementary Fig. S1 Comparative analysis of CMF genes in several plant genomes. a Number of CCT, CMF, and total coding genes in each plant species retrieved from PLAZA4.0 (Van Bel et al. 2018). b Parameters of linear regression analysis for gene number pairs: total coding gene and CMF gene number, total coding gene and CCT gene number, and finally CCT and CMF gene number. Values were calculated with InfoStat software (Rienzo et al. 2017). Supplementary Fig. S2 A maximum likelihood (ML) unrooted phylogenetic tree was constructed using MEGA X with 1000 bootstrap replications and an optimal Jones-Taylor-Thornton (JTT) model using the CMF proteins from soybean, rice, Arabidopsis, Chlamydomonas reinhardtii and Physcomitrella patens subsp. patens. For Arabidopsis and rice the TAIR annotation release 10 of the A. thaliana genome release 9 (Lamesch et al. 2012) and the MSU Release 7.0 of the annotation of the genome of the Nipponbare/ japonica subspecies of O. sativa (Ouyang et al. 2007) were used. A release of the mapped CC-503 MT + genome assembly and version 5.5 annotation (Merchant et al. 2007) and for P. patens subsp. patens the v3.3 annotation of the v3.0 assembly (Lang et al. 2018) were used for C. reinhardtii and P. patens subsp. patens, respectively. Supplementary Fig. S3 Expression levels of GmCMF genes in five tissues: flower, pod, leaves, stem, and root by qRT-PCR (40-dCT). Results are presented as means ± SD of three biological replicates. Different letters indicate significant differences at P < 0.05 according to the LSD ANOVA test (PDF 1284 KB)
Rights and permissions
About this article
Cite this article
Mengarelli, D.A., Zanor, M.I. Genome-wide characterization and analysis of the CCT motif family genes in soybean (Glycine max). Planta 253, 15 (2021). https://doi.org/10.1007/s00425-020-03537-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-020-03537-5