Abstract
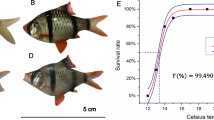
Larimichthys polyactis is one of the most economically important marine fish species that have become newly cultured in China in recent years. The gene expression changes that L. polyactis experiences in cold toleranceis still unknown, limiting the expansion of its cultivation, fast growth, and high yield. To investigate the molecular mechanism behind L. polyactis’s cold tolerance and to provide a resource for conducting genetic research on L. polyactis, transcriptome sequencing using RNA-seq was performed on individuals that survived cold stress at 4 °C (cold tolerant, CT), and individuals that barely survived 4 °C (cold sensitive, CS), which was considered as the control. A number of 387,607,550 clean reads were obtained from the transcriptomes, and comparative transcriptomic analysis identified 141 differently expressed genes (DEGs), of which 67 were up-regulated and 74 were down-regulated in CT compared to CS under cold stress. Furthermore, ten differently expressed genes were selected from the RNA-Seq analysis to be further validated by real-time PCR. Functional network analysis indicated that L. polyactis adapted to cold stress by employing a series of mechanisms to minimize damages caused by exposure to cold temperatures. The molecular mechanisms identified through RNA-Seq included Extracellular matrix (ECM) receptor interaction, glycerolipid metabolism, regulation of autophagy and focal adhesion pathway as playing vital roles in cold tolerance in L. polyactis. This study may help elucidate how L. polyactis tolerates cold, which is of value for breeding cold-tolerant L. polyactis stocks for cultivation.



Similar content being viewed by others
References
Barrett RD, Paccard A, Healy TM, Bergek S, Schulte PM, Schluter D, Rogers SM (2011) Rapid evolution of cold tolerance in stickleback. Proc Biol Sci 278:233–238
Chen RY, Lou B, Zhan W, Xu DD, Chen L, Liu F, Wang LG, Sun C (2016) Broodstock cultivation and spawning induction techniques in small yellow croaker Pseudosciaena polyactis. Fish Sci 35:250–254 (In Chinese with English abstract)
Chou MY, Hsiao CD, Chen SC, Chen IW, Liu ST, Hwang PP (2008) Effects of hypothermia on gene expression in zebrafish gills: upregulation in differentiation and function of ionocytes as compensatory responses. J Exp Biol 211:3077–3084
Cossins AR, Crawford DL (2005) Fish as models for environmental genomics. Nat Rev Genet 6:324–333
De Wit P, Pespeni MH, Ladner JT, Barshis DJ, Seneca F, Jaris H, Therkildsen NO, Morikawa M, Palumbi SR (2012) The simple fool’s guide to population genomics via RNA-Seq: an introduction to high-throughput sequencing data analysis. Mol Ecol Res 12:1058–1067
Donaldson MR, Cooke SJ, Patterson DA, Macdonald JS (2008) Cold shock and fish. J Fish Biol 73:1491–1530
FishBase (2014) fishbase. http://www.fishbase.org
Froese R, Pauly D (2003) Global capture production for Larimichthys polyactis. In: FAO fishery statistic 2003. FAO, Rome, Italy
Gracey AY, Fraser EJ, Li W, Fang Y, Taylor RR, Rogers J, Brass A, Cossins AR (2004) Coping with cold: an integrative, multitissue analysis of the transcriptome of a poikilothermic vertebrate. Proc Natl Acad Sci 101(48):16970–16975
He C, Klionsky DJ (2009) Regulation mechanisms and signaling pathways of autophagy. Annu Rev Genet 43(1):67–93
He J, Qiang J, Yang H, Xu P, Zhu ZX, Yang RQ (2015) Changes in the fatty acid composition and regulation of antioxidant enzymes and physiology of juvenile genetically improved farmed tilapia Oreochromis niloticus (L.), subjected to short-term low temperature stress. J Therm Biol 53:90–97
Hsieh SL, Hu CY, Hsu YT, Hsieh TJ (2007) Influence of dietary lipids on the fatty acid composition and stearoyl-CoA desaturase expression in hybrid tilapia (Oreochromis niloticus × O. aureus) under cold shock. Comp Biochem Physiol Part B 147:438–444
Hu J, You F, Wang Q, Weng S, Liu H, Wang L, Zhang P, Tan X (2014) Transcriptional responses of olive flounder (Paralichthys olivaceus) to low temperature. PLoS One 9:e108582
Hu P, Liu ML, Zhang D, Wang JF, Niu HB, Liu YM, Wu ZC, Han BS, Zhai WY, Shen Y, Chen LB (2015) Global identification of the genetic networks and cis-regulatory elements of the cold response in zebrafish. Nucleic Acids Res 43:9198–9213
Jin XS (2004) Long-term changes in fish community structure in the Bohai Sea, China. Estuar Coast Shelf Sci 59:163–171
Kyprianou TD, Pörtner HO, Anestis A, Kostoglou B, Feidantsis K, Michaelidis B (2010) Metabolic and molecular stress responses of gilthead seam bream Sparus aurata during exposure to low ambient temperature: an analysis of mechanisms underlying the winter syndrome. J Comp Physiol B 180:1005–1018
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359
Li Y, Han Z, Song N, Gao TX (2013) New evidence to genetic analysis of small yellow croaker (Larimichthys polyactis) with continuous distribution in China. Biochem Syst Ecol 50:331–338
Liang LQ, Chang YM, He XL, Tang R (2015) Transcriptome analysis to identify cold-responsive genes in Amur carp (Cyprinus carpio haematopterus). PLoS One 10:e0130526
Lin LS, Chen JH, Li HY (2008) The fishery biology of Trichiurus japonicus and Larimichthys polyactis in the East China Sea region. Marine Fish 30:126–134 (In Chinese with English abstract)
Liu LW, Sui YZ, Zhu WB, Guo A, Xu KD, Zhou YD (2017) In-depth transcriptome analysis of Larimichthys polyactis, de novo assembly, functional annotation. Marine Genomics 33:27–29
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2_DDCT method. Methods 25:402–408
Logan CA, Somero GN (2011) Effects of thermal acclimation on transcriptional responses to acute heat stress in the eurythermal fish Gillichthys mirabilis (Cooper). Am J Physiol 300:R1373–R1383
Long Y, Song G, Yan J, He X, Li Q, Cui Z (2013) Transcriptomic characterization of cold acclimation in larval zebrafish. BMC Genomics 14:612
Ma XY, Qiang J, He J, Gabriel NN, Xu P (2015) Changes in the physiological parameters, fatty acid metabolism, and SCD activity and expression in juvenile GIFT tilapia (Oreochromis niloticus) reared at three different temperatures. Fish Physiol Biochem 41:937–950
Mininni AN, Milan M, Ferraresso S, Petochi T, Di Marco P, Marino G, Livi S, Romualdi C, Bargelloni L, Patarnello T (2014) Liver transcriptome analysis in gilthead sea bream upon exposure to low temperature. BMC Genomics 15:765–777
Nagar R (2016) Autophagy: a brief overview in perspective of dermatology. Indian J Dermatol Venereol Leprol 83(3):290
Qi ZH, Liu YF, Luo SW, Chen CX, Liu Y, Wang WN (2013) Molecular cloning, characterization and expression analysis of tumor suppressor protein p53 from orangespotted grouper, Epinephelus coioides in response to temperature stress. Fish Shellfish Immunol 35:1466–1476
Qian B, Xue L (2016) Liver transcriptome sequencing and de novo annotation of the large yellow croaker (Larimichthy crocea) under heat and cold stress. Marine Genomics 25:95–102
Qiang J, Yang H, Wang H, Kpundeh MD, Xu P (2012) Growth and IGF-I response of juvenile Nile tilapia (Oreochromis niloticus) to changes in water temperature and dietary protein level. J Therm Biol 37:686–695
Qiang J, He J, Yang H, Wang H, Kpundeh MD, Xu P, Zhu ZX (2014) Temperature modulates hepatic carbohydrate metabolic enzyme activity and gene expression in juvenile GIFT tilapia (Oreochromis niloticus) fed a carbohydrate-enriched diet. J Therm Biol 40:25–31
Qin L, Peng D, Hu C, Xiang Y, Zhou YG, Tan YR, Qin XQ (2014) Differentiation of th subsets inhibited by nonstructural proteins of respiratory syncytial virus is mediated by ubiquitination. PLoS One 9:e101469
Salahudeen AK (2004) Cold ischemic injury of transplanted kidneys: new insights from experimental studies. Am J Physiol 287:F181–F187
Scott GR, Johnston IA (2012) Temperature during embryonic development has persistent effects on thermal acclimation capacity in zebrafish. Proc Natl Acad Sci 109:14247–14252
Seifert A, Werheid DF, Knapp SM, Tobiasch E (2015) Role of Hox genes in stem cell differentiation. World J Stem Cells 7:583–595
Snyder RJ, Hennessey TM (2003) Cold tolerance and homeoviscous adaptation in freshwater alewives (Alosa pseudoharengus). Fish Physiol Biochem 29:117–126
Somero GN (1995) Proteins and temperature. Annu Rev Physiol 57:43–68
Tiku PE, Gracey AY, Macartney AI, Beynon RJ, Cossins AR (1996) Cold-induced expression of delta 9-desaturase in carp by transcriptional and posttranslational mechanisms. Science 271(5250):815–818
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25(9):1105–1111
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28:511–515
Trueman RJ, Tiku PE, Caddick MX, Cossins AR (2000) Thermal thresholds of lipid restructuring and delta (9)-desaturase expression in the liver of carp (Cyprinus carpio L.). J Exp Biol 203:641–650
Vagner M, Santigosa E (2011) Characterization and modulation of gene expression and enzymatic activity of delta-6 desaturase in teleosts: a review. Aquaculture 315:131–143
Van Verk MC, Hickman R, Pieterse CM, Van Wees S (2013) RNA-Seq: revelation of the messengers. Trends Plant Sci 18:175–179
Yan WJ, Chen HM, Zhao XL, Yu H, Zhang M (2005) Cold stress causes early apoptosis in caudal fin cells of Colossoma brachypomum. Shi Yan Sheng Wu Xue Bao 38:105–110
Yang C, Jiang M, Wen H, Tian J, Liu W, Wu F, Gou G (2015) Analysis of differential gene expression under low temperature stress in Nile tilapia (Oreochromis niloticus) using digital gene expression. Gene 564:134–140
Zerai DB, Fitzsimmons KM, Collier RJ (2010) Transcriptional response of delta-9-desaturase gene to acute and chronic cold stress in Nile tilapia, Oreochromis niloticus. J World Aquac Soc 41:800–806
Acknowledgements
This research was supported by grants from the National Key Research and Development Program of China (No. 2018YFD0901204), the Special Fund for the Key Research and Development Project of Zhejiang Province (No. 2017C02013), and Zhejiang Provincial Natural Science Foundation of China (No. LQ19C190002).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Communicated by Bernd Pelster.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, F., Chu, T., Wang, M. et al. Transcriptome analyses provide the first insight into the molecular basis of cold tolerance in Larimichthys polyactis. J Comp Physiol B 190, 27–34 (2020). https://doi.org/10.1007/s00360-019-01247-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00360-019-01247-3