Abstract
Background
Disruptions in lipid metabolism and amino acids have been increasingly linked to resistance to immunotherapy. However, the underlying mechanisms by which dysregulated serum lipid metabolism and amino acids affect the efficacy of immunotherapies through PD-1/PD-L1 expression and function remain poorly understood.
Methods
To elucidate the potential associations between lipid metabolism, amino acids, and PD-1/PD-L1, we employed the powerful Mendelian randomization (MR) method, leveraging large-scale genome-wide association studies.
Results
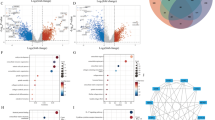
In the present MR study, we identified a noteworthy negative association between alanine and PD-1 expression, implicating a regulatory role for alanine metabolism in modulating the immune response to cancer treatment. Additionally, we elucidated fourteen specific lipid metabolism biomarkers that were significantly linked to PD-L1 expression, including cholesterol and triglycerides. Glutamine and phenylalanine were also found to showcase an intriguing causal association with the expression of PD-L1. Eventually, we confirmed the potential roles of key genes involved in lipid and amino acids metabolism in influencing the response to immunotherapy.
Conclusions
These findings provided new insights into the role of lipid metabolism as well as amino acids in regulating PD-1/PD-L1, suggesting that strategies targeting lipid and amino acid metabolisms may have therapeutic potential for improving the efficacy of immunotherapy.

Similar content being viewed by others
Data availability
The original data are available in the open GWAS project (https://gwas.mrcieu.ac.uk/).
Abbreviations
- CIs:
-
Confidence intervals
- HDL:
-
High-density lipoprotein
- ICB:
-
Immune checkpoint blockade
- LDL:
-
Low-density lipoprotein particles
- MR:
-
Mendelian randomization
- ORs:
-
Odd ratios
- PD-1:
-
Programmed cell death protein-1
- PD-L1:
-
Programmed cell death ligand-1
- SNPs:
-
Single nucleotide polymorphisms
- VLDL:
-
Very low-density lipoprotein
References
Bagchi S, Yuan R, Engleman E (2021) Immune checkpoint inhibitors for the treatment of cancer: clinical impact and mechanisms of response and resistance. Annu Rev Pathol 16:223–249. https://doi.org/10.1146/annurev-pathol-042020-042741
Morad G, Helmink BA, Sharma P, Wargo JA (2021) Hallmarks of response, resistance, and toxicity to immune checkpoint blockade. Cell 184:5309–5337. https://doi.org/10.1016/j.cell.2021.09.020
Bian X, Liu R, Meng Y, Xing D, Xu D, Lu Z (2021) Lipid metabolism and cancer. J Exp Med 218:e20201606. https://doi.org/10.1084/jem.20201606
Zhang H-L, Hu B-X, Li Z-L et al (2022) PKCβII phosphorylates ACSL4 to amplify lipid peroxidation to induce ferroptosis. Nat Cell Biol 24:88–98. https://doi.org/10.1038/s41556-021-00818-3
Yuan H, Wu X, Wu Q et al (2023) Lysine catabolism reprograms tumour immunity through histone crotonylation. Nature 617:818–826. https://doi.org/10.1038/s41586-023-06061-0
Davey Smith G, Hemani G (2014) Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum Mol Genet 23:R89-98. https://doi.org/10.1093/hmg/ddu328
Li W, Wang R, Wang W (2023) Exploring the causality and pathogenesis of systemic lupus erythematosus in breast cancer based on Mendelian randomization and transcriptome data analyses. Front Immunol 13:1029884. https://doi.org/10.3389/fimmu.2022.1029884
Wang Y, Guo Z, Isah AD, Chen S, Ren Y, Cai H (2023) Lipid metabolism and tumor immunotherapy. Front Cell Dev Biol 11:1187989. https://doi.org/10.3389/fcell.2023.1187989
Luo Y, Wang H, Liu B, Wei J (2022) Fatty acid metabolism and cancer immunotherapy. Curr Oncol Rep 24:659–670. https://doi.org/10.1007/s11912-022-01223-1
Tanaka S, Nemoto Y, Takei Y et al (2020) High-fat diet-derived free fatty acids impair the intestinal immune system and increase sensitivity to intestinal epithelial damage. Biochem Biophys Res Commun 522:971–977. https://doi.org/10.1016/j.bbrc.2019.11.158
Chowdhury PS, Chamoto K, Kumar A, Honjo T (2018) PPAR-induced fatty acid oxidation in T cells increases the number of tumor-reactive CD8+ T cells and facilitates anti-PD-1 therapy. Cancer Immunol Res 6:1375–1387. https://doi.org/10.1158/2326-6066.CIR-18-0095
Yang Z, Zhang D, Sima X et al (2023) Levels of pretreatment serum lipids predict responses to PD-1 inhibito r treatment in advanced intrahepatic cholangiocarcinoma. Int Immunopharmacol 115:109687. https://doi.org/10.1016/j.intimp.2023.109687
Leone RD, Zhao L, Englert JM et al (2019) Glutamine blockade induces divergent metabolic programs to overcome tumor immune evasion. Science 366:1013–1021. https://doi.org/10.1126/science.aav2588
Sikalidis AK (2015) Amino acids and immune response: a role for cysteine, glutamine, phenylalanine, tryptophan and arginine in T-cell function and cancer? Pathol Oncol Res 21:9–17. https://doi.org/10.1007/s12253-014-9860-0
Acknowledgements
We appreciate the work of the open GWAS project (https://gwas.mrcieu.ac.uk/).
Funding
This work was supported by the National Natural Science Foundation of China Grant (82172642 to W. Wang) and Natural Science Foundation of Guangdong Province (2021A1515011683 to W. Wang).
Author information
Authors and Affiliations
Contributions
WL & WW: Conceptualization, methodology, data curation, software, and writing—review & editing. The work reported in the paper has been performed by the authors, unless clearly specified in the text.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the study was performed in the absence of the conflict of interest.
Ethical approval
No ethics approval and written consent were needed for the secondary analysis of public data.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, W., Wang, W. Decoding the genetic links between serum lipidomic profile, amino acid biomarkers, and programmed cell death protein-1/programmed cell death-ligand-1. Cancer Immunol Immunother 72, 3395–3399 (2023). https://doi.org/10.1007/s00262-023-03501-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-023-03501-8