Abstract
The evaluation of genotoxicity plays an important role within hazard identification and risk assessment of chemicals and consumer products. For genotoxicity assessment, in vitro hepatic cells are often used as they have retained certain level of xenobiotic metabolic activity. However, current protocols are designed for the use on 2D monolayer models that are associated with several limitations due to the lack of numerous biological functions, which results in the loss of many hepatic properties. In this respect, an attractive alternative are three-dimensional (3D) models. The aim of our study was to develop physiologically more relevant 3D cell model (spheroids) from the human hepatocellular carcinoma (HepG2) cell line for genotoxicity testing. The spheroids were prepared by the forced floating method, which had been optimized for the production of a large number of uniform spheroids. The sensitivity of the spheroids to detect genotoxicity was determined by the comet assay after the exposure of spheroids to non-cytotoxic concentrations of model indirect acting genotoxic compounds, namely polycyclic aromatic hydrocarbon (B(a)P), mycotoxin (AFB1), two heterocyclic aromatic amines (PhIP and IQ) and a direct acting etoposide (ET). All five tested compounds concentration dependently induced DNA damage. Higher sensitivity of 3D cell model compared to 2D monolayer culture was noticed particularly for detection of the genotoxicity of the heterocyclic aromatic amines and BaP. Deregulation of mRNA expression (qPCR) by genotoxic compounds revealed that HepG2 cells in 3D express important genes encoding phase I and II metabolic enzymes, as well as DNA damage responsive genes in an inducible form. The newly developed HepG2 3D model shows improved sensitivity for detecting genotoxic compounds compared to 2D cultures and can provide a suitable experimental model for genotoxicity assessment.






Similar content being viewed by others
References
Altindag O, Karakoc M, Kocyigit A et al (2007) Increased DNA damage and oxidative stress in patients with rheumatoid arthritis. Clin Biochem 40:167–171. https://doi.org/10.1016/j.clinbiochem.2006.10.006
Baebler Š, Svalina M, Petek M et al (2017) quantGenius: implementation of a decision support system for qPCR-based gene quantification. BMC Bioinform 18:276. https://doi.org/10.1186/s12859-017-1688-7
Bromberg KD, Burgin AB, Osheroff N (2003) A two-drug model for etoposide action against human topoisomerase IIalpha. J Biol Chem 278:7406–7412. https://doi.org/10.1074/jbc.M212056200
Chang TT, Hughes-Fulford M (2009) Monolayer and spheroid culture of human liver hepatocellular carcinoma cell line cells demonstrate distinct global gene expression patterns and functional phenotypes. Tissue Eng Part A 15:559–567. https://doi.org/10.1089/ten.tea.2007.0434
Christmann M, Kaina B (2013) Transcriptional regulation of human DNA repair genes following genotoxic stress: trigger mechanisms, inducible responses and genotoxic adaptation. Nucleic Acids Res 41:8403–8420. https://doi.org/10.1093/nar/gkt635
Corvi R, Madia F (2017) In vitro genotoxicity testinge—can the performance be enhanced? NC-ND license. Food Chem Toxicol 106:600–608. https://doi.org/10.1016/j.fct.2016.08.024
den Braver-Sewradj SP, den Braver MW, Vermeulen NPE et al (2016) Inter-donor variability of phase I/phase II metabolism of three reference drugs in cryopreserved primary human hepatocytes in suspension and monolayer. Toxicol Vitr 33:71–79. https://doi.org/10.1016/J.TIV.2016.02.013
Denison MS, Whitlock JP (1995) Xenobiotic-inducible transcription of cytochrome P450 genes. J Biol Chem 270:18175–18178
Edmondson R, Broglie JJ, Adcock AF, Yang L (2014) Three-dimensional cell culture systems and their applications in drug discovery and cell-based biosensors. Assay Drug Dev Technol 12:207–218. https://doi.org/10.1089/adt.2014.573
Elje E, Hesler M, Rundén-Pran E et al (2019) The comet assay applied to HepG2 liver spheroids. Mutat Res Toxicol Environ Mutagen. https://doi.org/10.1016/J.MRGENTOX.2019.03.006
Ellinger-Ziegelbauer H, Stuart B, Wahle B et al (2005) Comparison of the expression profiles induced by genotoxic and nongenotoxic carcinogens in rat liver. Mutat Res Mol Mech Mutagen 575:61–84. https://doi.org/10.1016/j.mrfmmm.2005.02.004
Ewa B, Danuta M-Š (2017) Polycyclic aromatic hydrocarbons and PAH-related DNA adducts. J Appl Genet 58:321–330. https://doi.org/10.1007/s13353-016-0380-3
Fey SJ, Wrzesinski K (2012) Determination of drug toxicity using 3D spheroids constructed from an immortal human hepatocyte cell line. Toxicol Sci 127:403–411. https://doi.org/10.1093/toxsci/kfs122
Gajski G, Gerić M, Žegura B et al (2016) Genotoxic potential of selected cytostatic drugs in human and zebrafish cells. Environ Sci Pollut Res 23:14739–14750. https://doi.org/10.1007/s11356-015-4592-6
Gerets HHJ, Tilmant K, Gerin B et al (2012) Characterization of primary human hepatocytes, HepG2 cells, and HepaRG cells at the mRNA level and CYP activity in response to inducers and their predictivity for the detection of human hepatotoxins. Cell Biol Toxicol 28:69–87. https://doi.org/10.1007/s10565-011-9208-4
Godoy P, Hewitt NJ, Albrecht U, Andersen ME, Ansari N, Bhattacharya S, Bode JG, Bolleyn J, Borner C, Böttger J, Braeuning A, Budinsky RA, Burkhardt B, Cameron NR, Camussi G, Cho CS, Choi YJ, Craig Rowlands J, Dahmen U, Damm G, Dirsch O, Donato MT, Dong J, Dooley S, Drasdo D, Eakins R, Ferreira KS, Fonsato V, Fraczek J, Gebhardt R, Gibson A, Glanemann M, Goldring CEP, Gómez-Lechón MJ, Groothuis GMM, Gustavsson L, Guyot C, Hallifax D, Hammad S, Hayward A, Häussinger D, Hellerbrand C, Hewitt P, Hoehme S, Holzhütter HG, Houston JB, Hrach J, Ito K, Jaeschke H, Keitel V, Kelm JM, Kevin Park B, Kordes C, Kullak-Ublick GA, LeCluyse EL, Lu P, Luebke-Wheeler J, Lutz A, Maltman DJ, Matz-Soja M, McMullen P, Merfort I, Messner S, Meyer C, Mwinyi J, Naisbitt DJ, Nussler AK, Olinga P, Pampaloni F, Pi J, Pluta L, Przyborski SA, Ramachandran A, Rogiers V, Rowe C, Schelcher C, Schmich K, Schwarz M, Singh B, Stelzer EHK, Stieger B, Stöber R, Sugiyama Y, Tetta C, Thasler WE, Vanhaecke T, Vinken M, Weiss TS, Widera A, Woods CG, Xu JJ, Yarborough KM, Hengstler JG (2013) Recent advances in 2D and 3D in vitro systems using primary hepatocytes, alternative hepatocyte sources and non-parenchymal liver cells and their use in investigating mechanisms of hepatotoxicity, cell signaling and ADME. Arch Toxicol 87(8):1315–1530. https://doi.org/10.1007/s00204-013-1078-5
Gomez-Lechon M, Donato M, Castell J, Jover R (2003) Human hepatocytes as a tool for studying toxicity and drug metabolism. Curr. Drug Metab. 4:292–312. https://doi.org/10.2174/1389200033489424
Gomez-Lechon M, Donato M, Castell J, Jover R (2004) Human hepatocytes in primary culture: the choice to investigate drug metabolism in man. Curr Drug Metab 5:443–462. https://doi.org/10.2174/1389200043335414
Guerre P, Pineau T, Costet P et al (2000) Effects of AFB1 on CYP 1A1, 1A2 and 3A6 mRNA, and P450 expression in primary culture of rabbit hepatocytes. Toxicol Lett 111:243–251. https://doi.org/10.1016/S0378-4274(99)00181-2
Gunness P, Mueller D, Shevchenko V et al (2013) 3D organotypic cultures of human heparg cells: a tool for in vitro toxicity studies. Toxicol Sci 133:67–78. https://doi.org/10.1093/toxsci/kft021
Guo L, Dial S, Shi L et al (2011) Similarities and differences in the expression of drug-metabolizing enzymes between human hepatic cell lines and primary human hepatocytes. Drug Metab Dispos 39:528–538. https://doi.org/10.1124/dmd.110.035873
Hercog K, Maisanaba S, Filipič M et al (2017) Genotoxic potential of the binary mixture of cyanotoxins microcystin-LR and cylindrospermopsin. Chemosphere 189:319–329. https://doi.org/10.1016/j.chemosphere.2017.09.075
Hoeijmakers JHJ (2009) DNA damage, aging, and cancer. N Engl J Med 361:1475–1485. https://doi.org/10.1056/NEJMra0804615
Hollander MC, Zhan Q, Bae I, Fornace AJ (1997) Mammalian GADD34, an apoptosis- and DNA damage-inducible gene. J Biol Chem 272:13731–13737
Hreljac I, Zajc I, Lah T, Filipič M (2008) Effects of model organophosphorous pesticides on DNA damage and proliferation of HepG2 cells. Environ Mol Mutagen 49:360–367. https://doi.org/10.1002/em.20392
Hurrell T, Lilley KS, Cromarty AD (2019) Proteomic responses of HepG2 cell monolayers and 3D spheroids to selected hepatotoxins. Toxicol Lett 300:40–50. https://doi.org/10.1016/J.TOXLET.2018.10.030
Kamdem LK, Meineke I, Gödtel-Armbrust U et al (2006) Dominant contribution of P450 3A4 to the hepatic carcinogenic activation of aflatoxin B 1. Chem Res Toxicol 19:577–586. https://doi.org/10.1021/tx050358e
Le Hégarat L, Mourot A, Huet S et al (2014) Performance of comet and micronucleus assays in metabolic competent HepaRG cells to predict in vivo genotoxicity. Toxicol Sci 138:300–309. https://doi.org/10.1093/toxsci/kfu004
LeCluyse EL (2001) Human hepatocyte culture systems for the in vitro evaluation of cytochrome P450 expression and regulation. Eur J Pharm Sci 13:343–368
Lee C-M, Pohl J, Morgan ET (2009) Dual mechanisms of CYP3A protein regulation by proinflammatory cytokine stimulation in primary hepatocyte cultures. Drug Metab Dispos 37:865–872. https://doi.org/10.1124/dmd.108.026187
Li C-L, Tian T, Nan K-J et al (2008) Survival advantages of multicellular spheroids vs monolayers of HepG2 cells in vitro. Oncol Rep 20:1465–1471
Luckert C, Schulz C, Lehmann N et al (2017) Comparative analysis of 3D culture methods on human HepG2 cells. Arch Toxicol 91:393–406. https://doi.org/10.1007/s00204-016-1677-z
Majer BJ, Mersch-Sundermann V, Darroudi F et al (2004) Genotoxic effects of dietary and lifestyle related carcinogens in human derived hepatoma (HepG2, Hep3B) cells. Mutat Res Mol Mech Mutagen 551:153–166. https://doi.org/10.1016/j.mrfmmm.2004.02.022
Mazzoleni G, Di Lorenzo AD, Steimberg AN (2009) Modelling tissues in 3D: the next future of pharmaco-toxicology and food research? Genes Nutr 4:13–22. https://doi.org/10.1007/s12263-008-0107-0
Novak M, Žegura B, Baebler Š et al (2016) Influence of selected anti-cancer drugs on the induction of DNA double-strand breaks and changes in gene expression in human hepatoma HepG2 cells. Environ Sci Pollut Res 23:14751–14761. https://doi.org/10.1007/s11356-015-5420-8
Petković J, Žegura B, Stevanović M et al (2011) DNA damage and alterations in expression of DNA damage responsive genes induced by TiO 2 nanoparticles in human hepatoma HepG2 cells. Nanotoxicology 5:341–353. https://doi.org/10.3109/17435390.2010.507316
Pezdirc M, Žegura B, Filipič M (2013) Genotoxicity and induction of DNA damage responsive genes by food-borne heterocyclic aromatic amines in human hepatoma HepG2 cells. Food Chem Toxicol 59:386–394. https://doi.org/10.1016/j.fct.2013.06.030
Ramaiahgari SC, Den Braver MW, Herpers B et al (2014) A 3D in vitro model of differentiated HepG2 cell spheroids with improved liver-like properties for repeated dose high-throughput toxicity studies. Arch Toxicol 88:1083–1095. https://doi.org/10.1007/s00204-014-1215-9
Reisinger K, Blatz V, Brinkmann J et al (2018) Validation of the 3D Skin Comet assay using full thickness skin models: transferability and reproducibility. Mutat Res Toxicol Environ Mutagen 827:27–41. https://doi.org/10.1016/J.MRGENTOX.2018.01.003
Richert L, Liguori MJ, Abadie C et al (2006) Gene expression in human hepatocytes in suspension after isolation is similar to the liver of origin, is not affected by hepatocyte cold storage and cryopreservation, but is strongly changed after hepatocyte plating. Drug Metab Dispos 34:870–879. https://doi.org/10.1124/dmd.105.007708
Shah UK, de Mallia JO, Singh N et al (2018) A three-dimensional in vitro HepG2 cells liver spheroid model for genotoxicity studies. Mutat Res Genet Toxicol Environ Mutagen 825:51–58. https://doi.org/10.1016/j.mrgentox.2017.12.005
Shimada T (2006) Xenobiotic-metabolizing enzymes involved in activation and detoxification of carcinogenic polycyclic aromatic hydrocarbons. Drug Metab Pharmacokinet 21:257–276. https://doi.org/10.2133/dmpk.21.257
Singh NP, McCoy MT, Tice RR, Schneider EL (1988) A simple technique for quantitation of low levels of DNA damage in individual cells. Exp Cell Res 175:184–191. https://doi.org/10.1016/0014-4827(88)90265-0
Snykers S, De Kock J, Rogiers V, Vanhaecke T (2009) In Vitro differentiation of embryonic and adult stem cells into hepatocytes: state of the art. Stem Cells 27:577–605. https://doi.org/10.1634/stemcells.2008-0963
Soldatow VVY, Lecluyse EEL, Griffith LLG, Rusyn I (2013) In vitro models for liver toxicity testing. Toxicol Res (Camb) 2:23–39. https://doi.org/10.1039/C2TX20051A.In
Stiborová M, Moserová M, Černá V et al (2014) Cytochrome b5 and epoxide hydrolase contribute to benzo[a]pyrene-DNA adduct formation catalyzed by cytochrome P450 1A1 under low NADPH:p450 oxidoreductase conditions. Toxicology 318:1–12. https://doi.org/10.1016/j.tox.2014.02.002
Štraser A, Filipič M, Žegura B (2011) Genotoxic effects of the cyanobacterial hepatotoxin cylindrospermopsin in the HepG2 cell line. Arch Toxicol 85:1617–1626. https://doi.org/10.1007/s00204-011-0716-z
Takahashi Y, Hori Y, Yamamoto T et al (2015) Three-dimensional (3D) spheroid cultures improve the metabolic gene expression profiles of HepaRG cells. Biosci Rep. https://doi.org/10.1042/bsr20150034
Terashima J, Goto S, Hattori H et al (2015) CYP1A1 and CYP1A2 expression levels are differentially regulated in three-dimensional spheroids of liver cancer cells compared to two-dimensional monolayer cultures. Drug Metab Pharmacokinet 30:434–440. https://doi.org/10.1016/J.DMPK.2015.10.001
Tice RR, Agurell E, Anderson D et al (2000) Single cell gel/comet assay: guidelines for in vitro and in vivo genetic toxicology testing. Environ Mol Mutagen 35:206–221
Tomc J, Kološa K, Žegura B et al (2018) Adipose tissue stem cell-derived hepatic progenies as an in vitro model for genotoxicity testing. Arch Toxicol. https://doi.org/10.1007/s00204-018-2190-3
Turesky RJ (2010) Heterocyclic aromatic amines: potential human carcinogens. In: Fishbein J (ed) Advances in molecular toxicology. Elseiver, Amsterdam, pp 37–83
Waldherr M, Mišík M, Ferk F et al (2018) Use of HuH6 and other human-derived hepatoma lines for the detection of genotoxins: a new hope for laboratory animals? Arch Toxicol 92:921–934. https://doi.org/10.1007/s00204-017-2109-4
Westerink WMA, Schoonen WGEJ (2007) Cytochrome P450 enzyme levels in HepG2 cells and cryopreserved primary human hepatocytes and their induction in HepG2 cells. Toxicol Vitr 21:1581–1591. https://doi.org/10.1016/J.TIV.2007.05.014
Wilkening S, Stahl F, Bader A (2003) Comparison of primary human hepatocytes and hepatoma cell line HepG2 with regard to their biotransformation properties. Drug Metab Dispos 31:1035–1042. https://doi.org/10.1124/dmd.31.8.1035
Wong SF, No DY, Choi YY et al (2011) Concave microwell based size-controllable hepatosphere as a three-dimensional liver tissue model. Biomaterials 32:8087–8096. https://doi.org/10.1016/j.biomaterials.2011.07.028
Wrzesinski K, Fey SJ (2015) From 2D to 3D—a new dimension for modelling the effect of natural products on human tissue. Curr Pharm Des 21:5605–5616. https://doi.org/10.2174/1381612821666151002114227
Xu C, Li CY-T, Kong A-NT (2005) Induction of phase I, II and III drug metabolism/transport by xenobiotics. Arch Pharm Res 28:249–268. https://doi.org/10.1007/BF02977789
Žegura B, Volčič M, Lah TT, Filipič M (2008) Different sensitivities of human colon adenocarcinoma (CaCo-2), astrocytoma (IPDDC-A2) and lymphoblastoid (NCNC) cell lines to microcystin-LR induced reactive oxygen species and DNA damage. Toxicon 52:518–525. https://doi.org/10.1016/j.toxicon.2008.06.026
Zhang X, Yang S-T (2011) High-throughput 3-D cell-based proliferation and cytotoxicity assays for drug screening and bioprocess development. J Biotechnol 151:186–193. https://doi.org/10.1016/j.jbiotec.2010.11.012
Zhou B-BS, Elledge SJ (2000) The DNA damage response: putting checkpoints in perspective. Nature 408:433–439. https://doi.org/10.1038/35044005
Zhou S, Koh H-L, Gao Y et al (2004) Herbal bioactivation: the good, the bad and the ugly. Life Sci 74:935–968. https://doi.org/10.1016/J.LFS.2003.09.035
Acknowledgements
The authors acknowledge the financial support from the Slovenian Research Agency [research core funding P1-0245, J1-6730 and MR-MStampar], and COST Action CA16119 (In vitro 3-D total cell guidance and fitness) and CA15132 (The comet assay as a human biomonitoring tool).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supporting Information
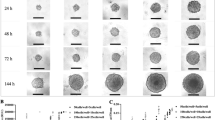
The Figure showing the results of protein expression. The Table showing the results of planimetry of formed spheroids after 24, 48, 72 and 96 h. (DOCX 160 kb)
Rights and permissions
About this article
Cite this article
Štampar, M., Tomc, J., Filipič, M. et al. Development of in vitro 3D cell model from hepatocellular carcinoma (HepG2) cell line and its application for genotoxicity testing. Arch Toxicol 93, 3321–3333 (2019). https://doi.org/10.1007/s00204-019-02576-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-019-02576-6