Abstract
HMOs (Human milk oligosaccharide) has an impact on maternal and infant health. Colostrum samples of 70 breastfeeding women in China were collected and recorded clinical characteristics. The major oligosaccharides and microbiota were quantitated in colostrum. The concentration of fucosylated HMOs in primipara was higher than that of multipara (p = 0.030). The concentration of N-acetylated HMOs in vaginal delivery milk was less than that of cesarean (p = 0.038). Non-fucosylated HMOs of breastfeeding women were less than that of breast pump (p = 0.038). Meanwhile, the concentration of LNT was positively correlated with Lactobacillus (r = 0.250, p = 0.037). DS-LNT was negatively correlated with Staphylococcus (r = – 0.240, p = 0.045). There was a positive correlation of Streptococcus with LNFP II (r = 0.314, p = 0.011) and 3-SL (r = 0.322, p = 0.009). In addition, there was a negative correlation between 2'-FL and 3-FL (r = – 0.465, p = 0.001). There was a positive correlation between LNT and LNnT (r = 0.778, p = 0.001). Therefore, the concentration of HMOs is related to number of deliveries, delivery mode, lactation mode and perinatal antibiotic. The concentration of HMOs is related to Lactobacillus, Streptococcus and Streptococcus in colostrum. In addition, there are connections between different oligosaccharides in content. The study protocol was also registered in the ClinicalTrails.gov (ChiCTR2200064454) (Oct. 2022).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human milk, which is the biological norm of infant nutrition, has been reported to contain many oligosaccharides. The content of HMOs (Human milk oligosaccharide) in breast milk is second to lactose and lipids (Thum et al. 2021). Colostrum is usually a sticky yellowish liquid secreted by the mammary gland within 7 days after delivery. The content of HMOs in colostrum is more than that of mature milk (Donovan and Comstock 2016). The concentration of HMOs in colostrum is about 9–22 g/L, decreased with the duration of lactation (Samuel et al. 2019; Liu et al. 2021; Poulsen et al. 2022). As we all know, HMOs has important physiological functions. It can affect the health status of newborns by supplementing probiotics (Pamela et al. 2018; Triantis et al. 2018; Sakanaka et al. 2019), resisting harmful microbiota (Quinn et al. 2020), simulating intestinal epithelial cell binding ligands and regulating the immune response (Ayechu-Muruzabal et al. 2022). Infant formula containing 2'-FL has been approved by the European Union and the US Food and Drug Administration (Reverri et al. 2018).
So far, 200 kinds of HMOs have been found (Urashima et al. 2018). At least 157 different HMOs structures were isolated and identified (Oursel et al. 2017; Peterson and Nagy 2021). The diversity of HMOs is affected by glycosidic bonds of different sugar units. According to the substituents, oligosaccharides are mainly divided into fucosylated HMOs, sialylated HMOs and non-fucosylated HMOs (Samuel et al. 2019). The main fucosylated HMOs are 2'-fucosyllactose (2'-FL) and 3-fucosyllactose (3-FL). Sialylated HMOs mainly includes 3-sialyllactose (3'SL) and 6-sialyllactose (6'SL). Non-fucosylated HMOs mainly include lacto-N-tetraose (LNT) and lacto-N-neotetraose (LNnT).
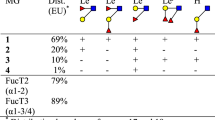
The concentration and composition of HMOs vary from individual to individual and from lactation to lactation (Thum et al. 2021). Fucosyltransferase 2 and 3 are encoded by secretory and Lewis genes, respectively. There are genetic variations that affect the activity (Soyyılmaz et al. 2021). In addition to genetic factors, other maternal factors may affect the composition of breast milk to some extent. The composition of HMOs was examined in 290 healthy breast milk samples. There were differences in the concentration of HMOs before pregnancy, delivery and parity (Samuel et al. 2019). Studies have provided data to show the correlation between HMOs and maternal factors, such as body weight, BMI, parity and age (McGuire et al. 2017; Azad et al. 2018). On the other hand, HMOs may also affect lactating mothers. HMOs can be used as a probiotic or antimicrobial agent to affect Staphylococci, Streptococci, Lactobacillus and Enterococci (Bode 2012). These oligosaccharides may affect the bacterial community in milk by promoting the growth of specific genera (Hunt et al. 2012), or directly regulate breast epithelial cell response and local immune response (Bode 2012).
Human milk microbiota affects maternal and infant health through breastfeeding. At the same time, HMOs affect the distribution and growth of infant intestinal microbiota. HMOs exist in human milk, which are associate with the milk microbiota and mother’s situation. Understanding the biological and environmental factors associated with the HMOs is an important part of the complex subject of maternal and child health. However, there are few studies on the relationship of HMOs and milk microbiota in China, especially in colostrum.
To our knowledge, the relationship between clinical characteristics and high levels of nutrition (including HMOs) in breast milk has not been confirmed. Though HMOs has been proved to have an impact on infant intestinal microbiota, it is not clear whether HMOs affected the microbiota of milk. Therefore, we collected human colostrum and pioneered the exploration of the potential relationship between HMOs and microbiota. At the same time, we observed the correlation of HMOs and maternal clinical characteristics. The findings may enrich the clinical data of breast milk research and guidelines for maternal and child health.
Materials and methods
Study participants
Healthy Chinese lactating women from the Shanghai Fourth People’s Hospital Affiliated to Tongji University School of Medicine were included in the study from October 2022 to December 2022. Informed consent was obtained from all subjects at enrollment. The study was approved by the Institutional Review Board (IRB) of the Shanghai Fourth People’s Hospital (20,211,124-001).
The inclusion criteria for the participants were as follows: (I) they were healthy and lactating; (II) postpartum colostrum was collected within 5 days after delivery. Exclusion criteria were as follows: (I) there was mammary abscess, (II) there was any other mammary pathology.
Collection and processing of milk samples
Colostrum samples were collected from healthy breastfeeding women within 5 days after delivery and 5 ml of milk was collected from each side of the breast. Women were asked to have not fed or expressed from the study breast for at least 2 h prior to sample collection. Before the milk collection, the areola and nipple area were sterilized by using 75% alcohol. The first drop of milk (about 150 μl) was discarded to avoid contamination. Samples were collected using a single-use, sterile containers and stored at − 20 °C. Microbiotal DNA was extracted within 24 h after collection.
Human milk oligosaccharide analysis
The HMO analysis was performed as previously described in the Laboratory of Immunopharmacology, Shanghai Institute of Materia Medical. After the samples were defatted and protein removed, the oligosaccharides were reduced to aldol with 1.0M sodium borohydride (NaBH4) and cleaned by solid phase extraction (SPE) with graphitized carbon cartridges. LC separation using binary gradient at 0.2 mL/min flow rate for 55 min. MS analysis runs in forward mode. Raffinose was added to each milk sample as an internal standard for absolute quantification. The concentration of HMOs was calculated as the specific oligosaccharides detected. The following HMOs were detected based on retention time comparison with commercial standard oligosaccharides (Zhenzhun Biotechnology, Shanghai, China) and mass spectrometry analysis: 2ʹ-fucosyllactose (2ʹ-FL Cat.ZMI-108106), 3-fucosyllactose (3-FL Cat.ZG10022), 3ʹ-sialyllactose (3ʹ-SL Cat.ZG-10015), lacto-N-tetraose (LNT Cat.ZEO-GLY-010), difucosyl-LNT (DS-LNT Cat. ZEO-GLY-066), lacto-N-neotetraose (LNnT Cat.ZB-038728), lacto-N-fucopentaose I (LNFP I Cat. ZG10048), LNFP II (Cat. ZG10049), and LNFP III (Cat. ZG10051).
Microbiotal DNA extraction
Total microbiotal DNA was extracted from the colostrum and mastitis milk samples using a microbiotal DNA Extraction Kit (TIANGEN, Beijing, China). Initially, milk samples (1 mL) were centrifuged at 12,000 rpm for 20 min at 4 °C. The supernatants with the fat and whey layer were removed. The protocol included an initial rupture of the microbiotal wall by 30 min of incubation with lysozyme (20 mg/mL) at 37 °C. Total DNA was then isolated from the pellets using the microbiotal DNA extraction kit (TIANGEN Cat: DP302) following the manufacturer’s instructions. A spectrophotometer (Thermo NanoDrop) was used to quantify the DNA. The purified DNA extracts were stored at – 20 °C.
Microbiotal quantitation of colostrum
Standard curves were created using serial tenfold dilutions of microbiotal DNA extracted from qPCR amplification products (Collado et al. 2009). A strain belonging to each of the microbiotal genera or groups targeted in this study was used to construct the standard curve. PCR standards were added, and the standard curve was obtained.
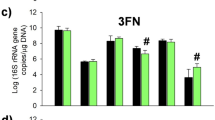
Microbial quantitation was based on the conserved marker gene 16s rRNA for microbiotal genus including Bifidobacterium, Lactobacillus, Staphylococcus and Streptococcus. Real-time PCR quantitation for target microbiotal gene was conducted with ABI 7900HT Fast Real-time PCR System (Applied Biosystems, Thermo Fisher, U.S.). Each reaction mixture (25 μl) was composed of 12.5 μl SYBR Green Master Mix (TIANGEN Cat: FP205), 0.1 ul of each of the specific primers at a con centration of 100 μM and 5.0 μL of template (1 ng/μl). Forty cycles of two-step polymerase chain reaction amplification were performed on the Applied Biosystems real-time polymerase chain reaction system (95 °C 5 s, 60 °C 32 s). The microbiotal concentration in each sample was measured as log10 genome equivalents by the interpolation of the Ct values obtained by the milk samples into the standard calibration curves. All samples were analyzed in two independent PCR assays, and the standard curve should be determined at all times.
The primer sequence was as follows: (Lactobacillus) LactoF TGGAAACAGRTGCTAATACCG; LactoR GTCCATTGTGGAAGATTCCC; (Bifidobacterium) T-Bifid426-F CTCGTAGGCGGTTCGTC; T-Bifid426-R GAACATGTCAAGCCCAGG; (Staphylococcus) TStaG422 GGCCGTGTTGAACGTGGTCAAATCA; TStag765 TACCATTTCAGTACCTTCTGGTAA;(Streptococcus) Tuf-Strep-1 GAAGAATTGCTTGAATTGGTTGAA; Tuf-Strep-R GGACGGTAGTTGTTGAAGAATGG.
Statistical analysis
All data were expressed as \(\overline{x }\pm s\) or [M (Q25, Q75)]. Continuous variables were compared using the t test or the Mann–Whitney U test. A single-factor analysis of variance (ANOVA) or Kruskal–Wallis test was used for multiple comparisons. A spearman correlation matrix was calculated for the HMO groups and the different microbiota, as well as individual HMO structures and the microbiota. p < 0.05 was considered the statistically significant level. All the PCR data represented three groups. The statistical analysis was done with IBM SPSS version 27.0 (Chicago, IL, U.S.) and Graphpad Prism version 9.0 (La Jolla, CA, U.S.).
Results
Clinical data of participants
A total of 70 subjects were included in this study, excluding those who lost follow-up and missing data (Fig. 1). Table 1 shows the participant’s description data. The average age of pregnant women was 29.7 ± 3.9, and the gestational weeks were normal. The maternal colostrum with an average of 2.9 ± 1.0 days was collected. The subjects were counted according to the number of deliveries, mode of delivery, lactation mode, situation before delivery, and whether to use antibiotics in perinatal period.
The concentration of HMOs
The concentrations of major oligosaccharides were as follow: 2'-FL 440.1 ± 215.6 ng/ml, 3-FL 665.7 ± 597.8 ng/ml, LNFP I 597.3 ± 470.1 ng/ml, LNFP II 373.4 ± 228.5 ng/ml, LNFP III 84.3 ± 33.9 ng/ml, 3-SL 560.7 ± 383.9 ng/ml, DS-LNT 575.6 ± 522.7 ng/ml, LNT 785.7 ± 460.5 ng/ml, LNnT 142.7 ± 99.4 ng/ml (Fig. 2). The substituents are divided into three categories according to their substituents, which are fucosylated HMOs (2ʹ-FL, 3ʹ-FL, LNFP I, LNFP II, LNFP III), sialylated HMOs (3-SL, DS-LNT) and non-fucosylated HMOs (LNT, LNnT). Furthermore, the correlation between different oligosaccharides was analyzed (Table 2). The results showed that there was a negative correlation between 2'-FL and 3-FL (r = – 0.465, p = 0.001), 3-FL and LNFP I (r = – 0.334, p = 0.005). There was a positive correlation between 2'-FL and LNFP I (r = 0.449, p = 0.001), LNT and LNnT (r = 0.778, p = 0.001) (Fig. 3).
The concentrations of HMOs in human colostrum (a the concentrations of several HMOs: 2'-FL 440.1 ± 215.6 ng/ml, 3-FL 665.7 ± 597.8 ng/ml, LNFP I 597.3 ± 470.1 ng/ml, LNFP II 373.4 ± 228.5 ng/ml, LNFP III 84.3 ± 33.9 ng/ml, 3-SL 560.7 ± 383.9 ng/ml, DS-LNT 575.6 ± 522.7 ng/ml, LNT 785.7 ± 460.5 ng/ml, LNnT 142.7 ± 99.4 ng/ml; b the concentrations of three kinds of HMOs: fucosylated HMOs 2314.3 ± 1994.4 ng/ml, sialylated HMOs 1136.3 ± 791.5 ng/ml, non-fucosylated HMOs928.3 ± 530.7 ng/ml)
The Relationship between HMOs and clinical characteristics
The HMO of different groups are compared according to the number of deliveries, mode of delivery, lactation mode, situation before delivery, and whether to use antibiotics in perinatal period (Fig. 4). The results showed that the concentration of non-fucosylated HMOs in vaginal delivery milk was less than that of cesarean (p = 0.038). The concentration of primipara Fucosylated HMOs was higher than that of multipara (p = 0.030). Non-fucosylated HMOs in breast milk of women who took antibiotics during the perinatal period were less than those who did not use (p = 0.020). Non-fucosylated HMOs in breast milk from breastfeeding women who were sucked by the baby were less than that of that breast pump (p = 0.038). Vacation or work before delivery and abortion history has no significant effect on HMOs concentration (p > 0.05).
The correlation between HMOs and microbiota
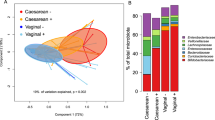
Bifidobacterium, Lactobacillus, Staphylococcus and Streptococcus was detected in milk (Fig. 5). We analyzed the correlation between microbiota and HMOs (Fig. 6, Table 3). The results showed that LNT was positively correlated with Lactobacillus (r = 0.250, p = 0.037). The concentration of DS-LNT was negatively correlated with Staphylococcus (r = – 0.240, p = 0.045). There was a positive correlation between Streptococcus and LNFP II (r = 0.314, p = 0.011), LNFP III (r = 0.251, p = 0.044), 3-SL (r = 0.322, p = 0.009), LNnT (r = 0.292, p = 0.018).
Discussion
In this study, we collected the precious colostrum. We quantitatively analyzed oligosaccharides in colostrum. Then, we found the relevance of oligosaccharides and the clinical characteristics of breastfeeding women. Furthermore, this study explored the relevance of the microbiota and HMOs in milk, which could guide lactation.
Human milk contains the plenty HMOs and various microbiota categories. The results from our study confirmed that colostrum samples of human milk consisted of high content of HMOs, with the evidences indicating that the Fucosylated HMOs as the highest compared with Sialylated HMOs or non-fucosylated HMOs. It was found that the ratio of the oligosaccharides in HMOs varied among the breastfeeding (Thurl et al. 2017; Xu et al. 2017). Our study exhibited the concentrations of 2'-FL, 3-FL, LNT and LNFP I were 0.58 g/L, 0.4 g/L, 0.55 g/L and 0.47 g/L correspondingly. It was reported that 2'-FL could be considered as the most abundant oligosaccharide in colostrum with the level almost 4.1 g/L, with the concentrations of 3-FL, LNT and LNFP I lower than it (Marriage et al. 2015; Thum et al. 2021). Meanwhile, the results from one meta study indicated that the mean content of 2'-FL, 3-FL and LNFP I varied from 0.14 to 2.74 g/L (Thurl et al. 2017), which was consistent with our founding. The concentrations of LNFP II and LNFP III were 0.19 g/L and 0.07 g/L, which were lower than the reported mean content of 0.21–0.58 g/L (Thurl et al. 2017). Previously research indicated that the level of LNnT was 0.36–1.12 g/L (Ma et al. 2018) which was higher than our results of 0.12 g/L. And the concentrations of 3-SL and DS-LNT were 0.50 g/L and 0.39 g/L in this study, which were different from the reported results of 0.19–0.29 g/L (3-SL) and 0.50–0.77 g/L(DS-LNT) (Gidrewicz and Fenton 2014; Van Niekerk et al. 2014). Previously studies mainly concentrated in the European or America, which rarely involved Asia, especially the China. Differences between our results and other reports would originate in the regional differences, which would be affected by the race, culture and living habits, etc. Besides, the concentrations of HMOs would be influenced by the personal status, including the α (1,2)-fucosyltransferase (FUT2) and enzyme FUT3 encoded by Lewis blood group gene (Morrow et al. 2011; Soyyılmaz et al. 2021). In addition, the differences would be attributed to the measurements conducted in the various studies (Austin and Bénet 2018; Huang et al. 2019). The method conducted in our results was LC–ESI–MS to determine the concentrations with the advancement of fast and convenient (Chaturvedi et al. 1997; Porfirio et al. 2020; Catenza and Donkor 2021). Other detection methods, such as refractive index detection (RID), evaporative light scattering detection (ELSD) and capillary electrophoresis (CE), have common disadvantages including complicated sample preparation steps and low sensitivity (Sarkozy et al. 2021).
The relationship between kinds of HMOs was observed in our study, including the negative correlation between 2'-FL and 3-FL which was consistent with the reported results (r = 0.78–0.99) (Austin et al. 2016; Thurl et al. 2017). The two HMOs contained the similar molecular structure which shared the same substrate (Guanosine 5′-diphosphate (GDP)-l-amylose), further resulting the negative correlation (Thum et al. 2021). The positive relationship of 2'-FL and LNFP I was observed in this study, which originated in α1-2 Rockweed glycosylation and the high dependence of FUT2 activity (Phipps et al. 2021). The results from our study also indicated that LNT positively related with LNnT, with supporting evidences that the two HMOs could be regulated by α1-2-fucosyltransferase (FUT2) (Sprenger et al. 2017). In addition, it was reported that LNT was negatively related with 2′-FL (Sprenger et al. 2017), which was different from our results that no relationship between LNT and 2′-FL founded. Meanwhile, our results exhibited that LNFP I was negatively related with 3-FL. However, Gabrielli et al. (2011) reported a positive correlation between the two HMOs, which would attributed to the different race and living habits, etc. Based on the above results, it was supposed that the HMOs with the similar molecular structure contained the homologous synthesis pathway, such as the regulation of Sialylated HMOs including 3'-SL and DS-LNT.
Oligosaccharides were formed through the conjunction of nucleotide sugar molecules based on the glycosidic bonds, which were catalyzed by the glycosyltransferases. The generation of HMOs was regulated by the cellular glycosyltransferases (including FUT2 and FUT3) in mammary gland cells, and simultaneous influenced by substrate. All the process were closely related with the physiological state of the mother, which caused the sightly influence on the glycosylation during the onset of the breastfeeding (Samuel et al. 2019). Concentrations of specific HMOs were related with maternal age, allergy history, pre-pregnancy body mass index, gestational age, mode of delivery, infant gestational age and sex (Wang et al. 2020). The results from this study indicated that higher concentrations of fucosylated HMOs in first-time mothers’ milk was founded compared with the milk from multiple births. Niekerk et al. (2014) also reported that the concentrations of HMOs were negatively correlated with births and the first contain the higher level, which was consistent with our results. But Azad et al. (2018) reported that there were relationships between the concentrations of HMOs and the times of birth. Meanwhile, this study exhibited that the non-fucosylated HMOs concentrations in breast milk from the mother underwent vaginal delivery were lower than it from the mother with cesarean delivery. However, it was reported that no relationship was founded between HMOs concentrations and delivery modes (Azad et al. 2018). In addition, the non-fucosylated HMOs concentrations in breast milk from breastfeeding mothers who are suckled by their babies was lower than it from the mothers who breastfeed the children through the breast-pumps. And the concentrations of non-fucosylated HMOs in the mother who unused antibiotics during the perinatal administration were higher. All the evidences suggested that there are differences in the susceptibility of various HMOs to maternal characteristics, further revealed that there may be different synthetic pathways for each type of HMOs.
It was noteworthy that there were complex relationships between HMOs and microorganisms. Previously studies reported that HMOs could directly mediate infant intestinal Staphylococci, Streptococci, Lactobacilli and Enterococci, or modulate maternal mammary epithelial cell responses and local immune responses (Bode 2012; Porro et al. 2022). Bifidobacterial proliferation in breastfed infants was correlated with the high levels of HMOs in breast milk from mothers (Le Doare et al. 2018; Moya-Gonzálvez et al. 2021). The breast milk was not considered as non-sterile and consisted of complex flora, which varies greatly among individuals (Tao et al. 2020). Based on the results from the measurements of colostrum samples, Lactobacillus and Streptococcus contents were higher compared with Bifidobacterium and Staphylococcus. The bacteria of breast milk could originate from the skin surface, and was associated with its own intestinal flora (Latuga et al. 2014). It was supposed that various bacteria utilized HMOs (such as Bifidobacterium and Lactobacillus), or benefited from their promotive/inhibitory effects without direct utilization (Craft and Townsend 2018; Zúñiga et al. 2018), with the possible relationships between HMOs and bacteria desired further exploration.
However, rare studies focused on the potential relationships between HMOs and bacteria of breast milk. The results from our study founded that DS-LNT was negatively related with Staphylococcus, which was consistent with the report that HMOs contained the bacteriostatic effect on pathogens from Yue et al. (2020). Rubio et al. (2019) also founded that the higher the level of staphylococcus in breast milk accompanied with the lower concentrations of total HMOs. Moossavi et al. (2019) analyzed the milk samples from 393 mothers and reported that oligosaccharides were associated with the diversity of lactic microbial communities with results including the negative relationship between HMOs and Staphylococcus (r = – 0.60 p = 0.038). It was believed that Staphylococci could bind to HMOs or HMOs contained the inhibitor effect on Staphylococci proliferation, which lead to the negative relationship between them. However, it was reported that the relative abundance of Staphylococcus was positively correlated with HMOs content based on the measurements from healthy breast milk samples (Williams et al. 2017). Therefore, the potential relationship still desired further confirmation through the clinical studies from with the expand sample size.
It was delighted that we found LNT was positively correlated with Lactobacillus. Rubio et al. (2019) also obtained the similar conclusion that higher levels of Lactobacillus were positively associated with concentrations of 2'-FL (r = 0.542, p = 0.038 in colostrum samples, and r = 0.700, p = 0.001 in mature samples). The results from our study proved that LNFP II, LNFP III, 3-SL, and LNnT possessed positive effects on Streptococcus proliferation. It was reported that Streptococcus levels were positively correlated with total HMOs in colostrum and transition samples of breast milk (Cabrera-Rubio et al. 2019). Besides, previously studies reported that oligosaccharides, such as 2'-FL et al., inhibited the growth of Group B Streptococci and Streptococcus pyogenes in vitro (Ackerman et al. 2017; Salli et al. 2020). However, the categories of Streptococcus genus also included the common probiotics such as Streptococcus thermophilus et al., in addition to Group B Streptococcus and Streptococcus pyogenes. It was important to conduct a series of studies to confirm the relationships between HMOs and Streptococcus. There were no relationships between HMOs and Bifidobacterium founded in our study. But evidences proved that Bifidobacterium was positively correlated with Sialylated HMOs and LNT (Aakko et al. 2017), and negatively correlated with DS-LNT (Moossavi et al. 2019). There were no associations between HMOs and bacteria of breast milk, which would be attributed to characteristic differences of HMO consumption by bacteria (Zúñiga et al. 2018). Therefore, the in-depth and comprehensive researches needed to be conducted to explore the potential relationships between HMOs and bacteria of breast milk.
In conclusion, there is a positive correlation between HMOs that depend on FUT2 in colostrum. We found that maternal clinical characteristics such as mode of delivery, number of delivery and lactation mode were associated with HMOs. In addition, there were correlations between HMOs and Lactobacillus, Staphylococcus and Streptococcus in milk. However, we were limited by multiple factors, such as the number of subjects, age, regional provinces and so on. In this study, secretor status was not determined on the basis of serological tests. In order to enrich the study of Chinese colostrum, it should be further studied in more groups in the future. Further experiments could be carried out to explore the mechanism of HMOs and Microbiota in vivo and in vitro.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Aakko J, Kumar H, Rautava S, Wise A, Autran C, Bode L, Isolauri E, Salminen S (2017) Human milk oligosaccharide categories define the microbiota composition in human colostrum. Benef Microbes 8(4):563–567
Ackerman DL, Doster RS, Weitkamp JH, Aronoff DM, Gaddy JA, Townsend SD (2017) Human milk oligosaccharides exhibit antimicrobial and antibiofilm properties against group B streptococcus. ACS Infect Dis 3(8):595–605
Austin S, Bénet T (2018) Quantitative determination of non-lactose milk oligosaccharides. Anal Chim Acta 1010:86–96
Austin S, De Castro CA, Bénet T, Hou Y, Sun H, Thakkar SK, Vinyes-Pares G, Zhang Y, Wang P (2016) Temporal change of the content of 10 oligosaccharides in the milk of Chinese urban mothers. Nutrients 8(6):346
Ayechu-Muruzabal V, Poelmann B, Berends AJ, Kettelarij N, Garssen J, Van’t Land B, Willemsen LEM (2022) Human milk oligosaccharide 2’-fucosyllactose modulates local viral immune defense by supporting the regulatory functions of intestinal epithelial and immune cells. Int J Mol Sci 23(18):10958
Azad M, Robertson B, Atakora F, Becker A, Subbarao P, Moraes T, Mandhane P, Turvey S, Lefebvre D, Sears M, Bode L (2018) Human milk oligosaccharide concentrations are associated with multiple fixed and modifiable maternal characteristics, environmental factors, and feeding practices. J Nutr 148(11):1733–1742
Bode L (2012) Human milk oligosaccharides: every baby needs a sugar mama. Glycobiology 22(9):1147–1162
Cabrera-Rubio R, Kunz C, Rudloff S, García-Mantrana I, Crehuá-Gaudiza E, Martínez-Costa C, Collado MC (2019) Association of maternal secretor status and human milk oligosaccharides with milk microbiota: an observational pilot study. J Pediatr Gastroenterol Nutr 68(2):256–263
Catenza KF, Donkor KK (2021) Recent approaches for the quantitative analysis of functional oligosaccharides used in the food industry: a review. Food Chem 355:129416
Chaturvedi P, Warren CD, Ruiz-Palacios GM, Pickering LK, Newburg DS (1997) Milk oligosaccharide profiles by reversed-phase HPLC of their perbenzoylated derivatives. Anal Biochem 251(1):89–97
Collado MC, Delgado S, Maldonado A, Rodriguez JM (2009) Assessment of the bacterial diversity of breast milk of healthy women by quantitative real-time PCR. Lett Appl Microbiol 48(5):523–528
Craft KM, Townsend SD (2018) The human milk glycome as a defense against infectious diseases: rationale, challenges, and opportunities. ACS Infect Dis 4(2):77–83
Donovan S, Comstock S (2016) Human milk oligosaccharides influence neonatal mucosal and systemic immunity. Ann Nutr Metab 69(Suppl 2):42–51
Gabrielli O, Zampini L, Galeazzi T, Padella L, Santoro L, Peila C, Giuliani F, Bertino E, Fabris C, Coppa GV (2011) Preterm milk oligosaccharides during the first month of lactation. Pediatrics 128(6):e1520-1531
Gidrewicz DA, Fenton TR (2014) A systematic review and meta-analysis of the nutrient content of preterm and term breast milk. BMC Pediatr 14:216
Huang X, Zhu B, Jiang T, Yang C, Qiao W, Hou J, Han Y, Xiao H, Chen L (2019) Improved simple sample pretreatment method for quantitation of major human milk oligosaccharides using ultrahigh pressure liquid chromatography with fluorescence detection. J Agric Food Chem 67(44):12237–12244
Hunt KM, Preuss J, Nissan C, Davlin CA, Williams JE, Shafii B, Richardson AD, McGuire MK, Bode L, McGuire MA (2012) Human milk oligosaccharides promote the growth of staphylococci. Appl Environ Microbiol 78(14):4763–4770
Latuga MS, Stuebe A, Seed PC (2014) A review of the source and function of microbiota in breast milk. Semin Reprod Med 32(1):68–73
Le Doare K, Holder B, Bassett A, Pannaraj PS (2018) Mother’s milk: a purposeful contribution to the development of the infant microbiota and immunity. Front Immunol 9:361
Liu S, Cai X, Wang J, Mao Y, Zou Y, Tian F, Peng B, Hu J, Zhao Y, Wang S (2021) Six oligosaccharides’ variation in breast milk: a study in South China from 0 to 400 days postpartum. Nutrients 13(11):4017
Ma L, McJarrow P, Jan Mohamed HJB, Liu X, Welman A, Fong BY (2018) Lactational changes in the human milk oligosaccharide concentration in Chinese and Malaysian mothers’ milk. Int Dairy J 87:1–10
Marriage B, Buck R, Goehring K, Oliver J, Williams J (2015) Infants fed a lower calorie formula with 2’FL show growth and 2’FL uptake like breast-fed infants. J Pediatr Gastroenterol Nutr 61(6):649–658
McGuire M, Meehan C, McGuire M, Williams J, Foster J, Sellen D, Kamau-Mbuthia E, Kamundia E, Mbugua S, Moore S, Prentice A, Kvist L, Otoo G, Brooker S, Price W, Shafii B, Placek C, Lackey K, Robertson B, Manzano S, Ruíz L, Rodríguez J, Pareja R, Bode L (2017) What’s normal? Oligosaccharide concentrations and profiles in milk produced by healthy women vary geographically. Am J Clin Nutr 105(5):1086–1100
Moossavi S, Atakora F, Miliku K, Sepehri S, Robertson B, Duan QL, Becker AB, Mandhane PJ, Turvey SE, Moraes TJ, Lefebvre DL, Sears MR, Subbarao P, Field CJ, Bode L, Khafipour E, Azad MB (2019) Integrated analysis of human milk microbiota with oligosaccharides and fatty acids in the CHILD cohort. Front Nutr 6:58
Morrow A, Meinzen-Derr J, Huang P, Schibler K, Cahill T, Keddache M, Kallapur S, Newburg D, Tabangin M, Warner B, Jiang X (2011) Fucosyltransferase 2 non-secretor and low secretor status predicts severe outcomes in premature infants. J Pediatr 158(5):745–751
Moya-Gonzálvez EM, Rubio-Del-Campo A, Rodríguez-Díaz J, Yebra MJ (2021) Infant-gut associated Bifidobacterium dentium strains utilize the galactose moiety and release lacto-N-triose from the human milk oligosaccharides lacto-N-tetraose and lacto-N-neotetraose. Sci Rep 11(1):23328
Oursel S, Cholet S, Junot C, Fenaille F (2017) Comparative analysis of native and permethylated human milk oligosaccharides by liquid chromatography coupled to high resolution mass spectrometry. J Chromatogr B, Anal Technol Biomed Life Sci 1071:49–57
Pamela T, Medina DA, Garrido D (2018) Human milk oligosaccharides and infant gut bifidobacteria: molecular strategies for their utilization. Food Microbiol 75:37–46
Peterson T, Nagy G (2021) Toward sequencing the human milk glycome: high-resolution cyclic ion mobility separations of core human milk oligosaccharide building blocks. Anal Chem 93(27):9397–9407
Phipps KR, Lynch B, Stannard DR, Gilby B, Baldwin N, Mikš MH, Lau A, Röhrig CH (2021) Genotoxicity and neonatal subchronic toxicity assessment of a novel mixture of the human-identical milk oligosaccharides lacto-N-fucopentaose I and 2’-fucosyllactose. J Appl Toxicol 41(4):632–649
Porfirio S, Archer-Hartmann S, Moreau GB, Ramakrishnan G, Haque R, Kirkpatrick BD, Petri WA, Azadi P (2020) New strategies for profiling and characterization of human milk oligosaccharides. Glycobiology 30(10):774–786
Porro M, Kundrotaite E, Mellor D, Munialo C (2022) A narrative review of the functional components of human breast milk and their potential to modulate the gut microbiome, the consideration of maternal and child characteristics, and confounders of breastfeeding, and their impact on risk of obesity later in life. Nutr Rev 81(5):597–609
Poulsen K, Meng F, Lanfranchi E, Young J, Stanton C, Ryan C, Kelly A, Sundekilde U (2022) Dynamic changes in the human milk metabolome over 25 weeks of lactation. Front Nutr 9:917659
Quinn E, Joshi L, Hickey R (2020) Symposium review: dairy-derived oligosaccharides-their influence on host-microbe interactions in the gastrointestinal tract of infants. J Dairy Sci 103(4):3816–3827
Reverri EJ, Devitt AA, Kajzer JA, Baggs GE, Borschel MW (2018) Review of the clinical experiences of feeding infants formula containing the human milk oligosaccharide 2’-fucosyllactose. Nutrients 10(10):1346
Sakanaka M, Hansen ME, Gotoh A, Katoh T, Yoshida K, Odamaki T, Yachi H, Sugiyama Y, Kurihara S, Hirose J, Urashima T (2019) Evolutionary adaptation in fucosylactose uptake systems supports bifidobacteria infant symbiosis. Sci Adv 5(8):eaw7696
Salli K, Söderling E, Hirvonen J, Gürsoy UK, Ouwehand AC (2020) Influence of 2’-fucosyllactose and galacto-oligosaccharides on the growth and adhesion of Streptococcus mutans. Br J Nutr 124(8):824–831
Samuel T, Binia A, de Castro C, Thakkar S, Billeaud C, Agosti M, Al-Jashi I, Costeira M, Marchini G, Martínez-Costa C, Picaud J, Stiris T, Stoicescu S, Vanpeé M, Domellöf M, Austin S, Sprenger N (2019) Impact of maternal characteristics on human milk oligosaccharide composition over the first 4 months of lactation in a cohort of healthy European mothers. Sci Rep 9(1):11767
Sarkozy D, Borza B, Domokos A, Varadi E, Szigeti M, Meszaros-Matwiejuk A, Molnar-Gabor D, Guttman A (2021) Ultrafast high-resolution analysis of human milk oligosaccharides by multicapillary gel electrophoresis. Food Chem 341(Pt 2):128200
Soyyılmaz B, Mikš MH, Röhrig CH, Matwiejuk M, Meszaros-Matwiejuk A, Vigsnæs LK (2021) The mean of milk: a review of human milk oligosaccharide concentrations throughout lactation. Nutrients 13(8):2737
Sprenger N, Lee LY, De Castro CA, Steenhout P, Thakkar SK (2017) Longitudinal change of selected human milk oligosaccharides and association to infants’ growth, an observatory, single center, longitudinal cohort study. PLoS ONE 12(2):e0171814
Tao Y, Tong X, Qian C, Wan H, Zuo J (2020) Microbial quantitation of colostrum from healthy breastfeeding women and milk from mastitis patients. Ann Palliat Med 9(4):1666–1680
Thum C, Wall CR, Weiss GA, Wang W, Szeto IM, Day L (2021) Changes in HMO concentrations throughout lactation: influencing factors, health effects and opportunities. Nutrients 13(7):2272
Thurl S, Munzert M, Boehm G, Matthews C, Stahl B (2017) Systematic review of the concentrations of oligosaccharides in human milk. Nutr Rev 75(11):920–933
Triantis V, Bode L, van Neerven RJJ (2018) Immunological effects of human milk oligosaccharides. Front Pediatr 6:190
Urashima T, Hirabayashi J, Sato S, Kobata A (2018) Human milk oligosaccharides as essential tools for basic and application studies on galectins. Trends Glycosci Glycotechnol 30(172):51–65
Van Niekerk E, Autran CA, Nel DG, Kirsten GF, Blaauw R, Bode L (2014) Human milk oligosaccharides differ between HIV-infected and HIV-uninfected mothers and are related to necrotizing enterocolitis incidence in their preterm very-low-birth-weight infants. J Nutr 144(8):1227–1233
Wang M, Zhao Z, Zhao A, Zhang J, Wu W, Ren Z, Wang P, Zhang Y (2020) Neutral human milk oligosaccharides are associated with multiple fixed and modifiable maternal and infant characteristics. Nutrients 12(3):826
Williams JE, Price WJ, Shafii B, Yahvah KM, Bode L, McGuire MA, McGuire MK (2017) Relationships among microbial communities, maternal cells, oligosaccharides, and macronutrients in human milk. J Hum Lact 33(3):540–551
Xu G, Davis JC, Goonatilleke E, Smilowitz JT, German JB, Lebrilla CB (2017) Absolute quantitation of human milk oligosaccharides reveals phenotypic variations during lactation. J Nutr 147(1):117–124
Yue H, Han Y, Yin B, Cheng C, Liu L (2020) Comparison of the antipathogenic effect toward Staphylococcus aureus of N-linked and free oligosaccharides derived from human, bovine, and goat milk. J Food Sci 85(8):2329–2339
Zúñiga M, Monedero V, Yebra MJ (2018) Utilization of host-derived glycans by intestinal lactobacillus and bifidobacterium species. Front Microbiol 9:1917
Funding
This work was supported by Shanghai Hospital Specialized Integrated Traditional Chinese and Western Medicine (ZHYY-ZXY-JHZX-202110); Shanghai Hongkou District Health Committee Meeting Project (HKQ-ZYY-2020-02, HKGYQYXM-2022-05, HKGYQYXM-2022-21); Subject Boosting Program of the Shanghai Fourth People’s Hospital Affiliated to Tongji University (SY-XKZT-2020-2002); Shanghai Hospital of Traditional Chinese Medicine High-level Talents Leading Program.
Author information
Authors and Affiliations
Contributions
WS and LT drafted the manuscript and revised it critically for important intellectual content. CQ and PX analyzed and interpreted the data. XT and LY directed the experimental methods. FL provided some samples. HW and YT contributed substantially to the conception and design of the study. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare that they have no competing interests.
Ethical approval
This study was approved by the Ethics Committee and Institutional Review Board, and all patients provided written informed consent. The study protocol was also registered in the ClinicalTrails.gov (ChiCTR2200064454).
Additional information
Communicated by Yusuf Akhter.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sun, W., Tao, L., Qian, C. et al. Human milk oligosaccharides and the association with microbiota in colostrum: a pilot study. Arch Microbiol 206, 58 (2024). https://doi.org/10.1007/s00203-023-03787-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03787-3