Abstract
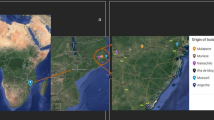
The Prespa lakes plain is an isolated area where about 1000 ha are seeded to Phaseolus vulgaris L. and Phaseolus coccineus L. Nodulation, arbuscular mycorrhizal fungal (AMF) presence and the genetic diversity of rhizobia were evaluated by 16S-ITS-23S-RFLP patterns and by sequencing. The bean rhizobial population in the region was diverse, despite its geographic isolation. No biogeographic relationships were detected, apart from a Rhizobium tropici-related strain that originated from an acidic soil. No clear pattern was detected in clustering with bean species and all isolates formed nodules with both bean species. Most strains were related to Rhizobium leguminosarum and a number of isolates were falling outside the already characterized species of genus Rhizobium. Application of heavy fertilization has resulted in high soil N and P levels, which most likely reduced nodulation and AMF spore presence. However, considerable AMF root length colonization was found in most of the fields.


Similar content being viewed by others
References
Abbaszadeh-dahaji P, Savaghebi GhR, Asadi-rahmani H, Rejali F, Farahbakhsh M, Moteshareh-zadeh D, Omidvari M, Lindstrom K (2012) Symbiotic effectiveness and plant growth promoting traits in some Rhizobium strains isolated from Phaseolus vulgaris L. Plant Growth Regul 68:361–370
Adhikari D, Itoh K, Suyama L (2013) Genetic diversity of common bean (Phaseolus vulgaris L.) nodulating rhizobia in Nepal. Plant Soil 368:341–353
Akter Z, Pageni BB, Lupwayi NZ, Balasubramanian PM (2014) Biological nitrogen fixation and nifH gene expression in dry beans (Phaseolus vulgaris L.). Can J Plant Sci 94:203–212
Álvarez-Martínez ER, Valverde Á, Ramírez-Bahena MH, García-Fraile P, Tejedor C, Mateos PF, Santillana N, Zúñiga D, Peix A, Velázquez E (2009) The analysis of core and symbiotic genes of rhizobia nodulating Vicia from different continents reveals their common phylogenetic origin and suggests the distribution of Rhizobium leguminosarum strains together with Vicia seeds. Arch Microbiol 191:659–668
Amarger N, Machere V, Laguerre G (1997) Rhizobium gallicum sp. nov. and Rhizobium giardinii sp. nov., from Phaseolus vulgaris nodules. Int J Syst Bacteriol 47:996–1006
Andrade DS, Murphy PJ, Giller KE (2002) The diversity of Phaseolus-nodulating rhizobial populations is altered by liming of acid soils planted with Phaseolus vulgaris L. in Brazil. Appl Environ Micorbiol 68:4025–4034
Anguilar OM, López MV, Donato M, Morón B, Soria-Diaz ME, Mateos C, Gil-Serrano A, Sousa C, Megías M (2006) Phylogeny and nodulation signal molecule of rhizobial populations able to nodulate common beans—other than the predominant species Rhizobium etli—present in soils from the northwest of Argentina. Soil Biol Biochem 38:573–586
Aoki S, Kondo T, Prévost D, Nakata S, Kajita T, Ito M (2010) Genotypic and phenotypic diversity of rhizobia isolated from Lathyrus japonicus indigenous to Japan. Syst Appl Microbiol 33:383–397
Aouani ME, Mhamdi R, Mars M, Elayeb M, Ghtir R (1997) Potential for inoculation of common bean by effective rhizobia in Tunisian soils. Agronomie 17:445–454
Aserse AA, Räsänen LA, Assefa F, Hailemariam A, Lindström K (2012) Phylogeny and genetic diversity of native rhizobia nodulating common bean (Phaseolus vulgaris L.) in Ethiopia. Syst Appl Microbiol 35:120–131
Baginsky C, Brito B, Scherson R, Pertuzé R, Seguelm O, Cañete A, Araneda C, Johnson WE (2015) Genetic diversity of Rhizobium from nodulating beans grown in a variety of Mediterranean climate soils of Chile. Arch Microbiol 197:419–429
Bernal G, Graham PH (2001) Diversity in the rhizobia associated with Phaseolus vulgaris L. in Ecuador, and comparisons with Mexican bean rhizobia. Can J Microbiol 47:526–534
Bustos P, Santamaria RI, Pérez-Carrascal OM, Acosta JL, Lozano L, Juárez S, Martínez-Flores I, Martínez-Romero E, Cevallos MA, Romero D, Dávila G, Vinuesa P, Miranda F, Ormeρo E, González V (2017) Complete genome sequences of three Rhizobium gallicum symbionts associated with common bean (Phaseolus vulgaris). Genome Announc 11:e00030–17
Caballero-Mellado J, Martínez-Romero E (1999) Soil fertilization limits the genetic diversity of Rhizobium in bean nodules. Symbiosis 26:111–121
Cao Y, Wang ET, Zhao L, Chen WM, Wei GH (2014) Diversity and distribution of rhizobia nodulated with Phaseolus vulgaris in two ecoregions of China. Soil Biol Biochem 78:128–137
Dall’Agnol RF, Ribeiro RA, Ormeño-Orrillo E, Rogel MA, Delamuta JRM, Andrade DS, Martínez-Romero E, Hungria M (2013) Rhizobium freirei sp. nov., a symbiont of Phaseolus vulgaris that is very effective at fixing nitrogen. Int J Syst Evol Microbiol 63:4167–4173
Dar GH, Zargar MY, Beigh GM (1997) Biocontrol of Fusarium root rot in the common bean (Phaseolus vulgaris L.) by using symbiotic Glomus mosseae and Rhizobium leguminosarum. Microb Ecol 34:74–80
Darriba D, Taboad GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772
Elbanna K, Elbadry M, Gamal-Eldin H (2009) Genotypic and phenotypic characterization of rhizobia that nodulate snap bean (Phaseolus vulgaris L.) in Egyptian soils. Syst Appl Microbiol 32:522–530
Embalomatis A, Papakosta DK, Katinakis P (1994) Evaluation of Rhizobium meliloti strains isolated from indigenous populations in northern Greece. J Agron Crop Sci 172:73–80
García-Fraile P, Mulas-García D, Peix A, Rivas R, González- Andrés F, Velázquez E (2010) Phaseolus vulgaris is nodulated in northern Spain by Rhizobium leguminosarum strains harboring two nodC alleles present in American Rhizobium etli strains: biogeographical and evolutionary implications. Can J Microbiol 56:657–666
Giller KE, Cadisch G (1995) Future benefits from biological nitrogen fixation—an ecological approach to agriculture. Plant Soil 174:255–277
Graham PH, Ranalli P (1997) Common bean (Phaseolus vulgaris L.). Field Crops Res 53:131–146
Graham PH, Rosas JC, Estevez de Jensen C, Peralta E, Tlusty B, Acosta-Gallegos J, Arraes Perreira PA (2003) Addressing edaphic constrains to bean production: the Bean/Cowpea CRSP project in perspective. Field Crops Res 82:179–192
Gryndler H, Leština J, Moravec V, Přikryl Z, Lipavsky J (1989) Colonization of maize roots by VAM-fungi under conditions of long-term fertilization of varying intensity. Agric Ecosyst Environ 29:183–186
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Hardarson G, Bliss FA, Cigales-Rivero MR, Henson RA, Kipe-Nolt JA, Longeri L, Manrique A, Pena-Cabriales JJ, Pereira PAA, Sanabria CA, Tsai SM (1993) Genotypic variation in biological nitrogen fixation by common bean. Plant Soil 152:59–70
Haukka K, Lindström K, Young JPW (1996) Diversity of partial 16S rRNA sequences among and within strains of African rhizobia isolated from Acacia and Prosopis. Syst Appl Microbiol 19:352–359
Havlin JJ, Tisdale SL, Nelson WL, Beaton JD (2014) Chapter 4: Nitrogen. In: Havlin JJ, Tisdale SL, Nelson WL, Beaton JD (eds) Soil fertility and fertilizers, 8th edn. Pearson, Prentice Hall, Upper Saddle River, pp 117–184
Hayman DS (1986) Mycorrhizae of nitrogen-fixing legumes. MIRCEN J 2:121–145
Hayman DS, Barea JM, Azcon R (1976) Vesicular-arbuscular mycorrhiza in southern Spain: its distribution in crops growing in soil of different fertility. Phytopathol Mediterr 15:1–6
Herrera-Cervera JA, Caballero-Mellado J, Laguerre G, Tichy HV, Requena N, Amarger N, Martínez-Romero E, Olivares J, Sanjuan J (1999) At least five rhizobial species nodulate Phaseolus vulgaris in a Spanish soil. FEMS Microbiol Ecol 30:87–97
Hou BC, Wang ET, Li Y, Jia RZ, Chen WF, Man CX, Sui XH, Chen WX (2009) Rhizobial resource associated with epidemic legumes in Tibet. Microb Ecol 57:69–81
Huang YY, Cho ST, Lo WS, Wang YC, Lai EML, Kuo CH (2015) Complete genome sequence of Agrobacterium tumefaciens Ach5. Genome Announc 3:e00570–15
Junier P, Alfaro M, Guevara R, Witzel KP, Carú M (2014) Genetic diversity of Rhizobium present in nodules of Phaseolus vulgaris L. cultivated in two soils of the central region in Chile. Appl Soil Ecol 80:60–66
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Kosmas C, Moustakas N, Tsatiris B, Danalatos N (1990) Evaluation of soil resources of the Prespa region, Greece. EEC Project B6617-25-89. Agricultural University of Athens, Athens
Kwon SW, Park JY, Kim JS, Kang JW, Cho YH, Lim CK, Parker MA, Lee GB (2005) Phylogenetic analysis of the genera Bradyrhizobium, Mesorhizobium, Rhizobium and Sinorhizobium on the basis of 16S rRNA gene and internally transcribed spacer region sequences. Int J Syst Evol Microbiol 55:263–270
Laguerre G, Mavingui P, Allard MR, Charnay MP, Louvrier P, Mazurier SI, Rigottier-Gois L, Amarger N (1996) Typing of rhizobia by PCR DNA fingerprinting and PCR-restriction fragment length polymorphism analysis of chromosomal and symbiotic gene regions: application to Rhizobium leguminosarum and its different biovars. Appl Environ Microbiol 62:2029–2036
López-López A, Rogel-Hernández MA, Barois I, Ceballos AIO, Martínez J, Ormeño-Orrillo E, Martínez-Romero E (2012) Rhizobium grahamii sp. nov., from nodules of Dalea leporina, Leucaena leucocephala and Clitoria ternatea, and Rhizobium mesoamericanum sp. nov., from nodules of Phaseolus vulgaris, siratro, cowpea and Mimosa pudica. Int J Syst Evol Microbiol 62:2264–2271
Mavromatis AG, Arvanitoyannis S, Korkovelos AE, Giakountis A, Chatzitheodorou VA, Goulas CK (2010) Genetic diversity among common bean (Phaseolus vulgaris L.) Greek landraces and commercial cultivars: nutritional components, RAPD and morphological markers. Span J Agric Res 8:986–994
Mhamdi R, Jebara M, Aouani ME, Ghrir R, Mars M (1999) Genotypic diversity and symbiotic effectiveness of rhizobia isolated from roots nodules of Phaseolus vulgaris L., grown in Tunisian soils. Biol Fertil Soil 28:313–320
Miller RM, Reinhardt DR, Jastrow JD (1995) External hyphal production of vesicular-arbuscular mycorrhizal fungi in pasture and tallgrass prairie communities. Oecologia 103:17–23
Mnasri B, Mrabet M, Laguerre G, Aouani ME, Mhamdi R (2007) Salt-tolerant rhizobia isolated from a Tunisian oasis that are highly effective for symbiotic N2-fixation with Phaseolus vulgaris constitute a novel biovar (bv. mediterranense) of Sinorhizobium meliloti. Arch Microbiol 187:79–85
Mnasri B, Saïdi S, Chihaou SA, Mhamdi R (2012) Sinorhizobium americanum symbiovar mediterranense is a predominant symbiont that nodulates and fixes nitrogen with common bean (Phaseolus vulgaris L.) in a northern Tunisian field. Syst Appl Microbiol 35:263–269
Mnasri B, Liu TY, Saidi S, Chen WF, Chen WX, Zhang XX, Mhamdi R (2014) Rhizobium azibense sp. nov., a nitrogen fixing bacterium isolated from root nodules of Phaseolus vulgaris. Int J Syst Evol Microbiol 64:1501–1506
Nelson M, Guhlin J, Epstein B, Tiffin P, Sadowsky MJ (2018) The complete replicons of 16 Ensifer meliloti strains offer insights into intra-and inter-replicon gene transfer, transposon-associated loci, and repeat elements. Microb Genom 4:e000174
Oliveira JP, Galli-Terasawa LV, Enke CG, Cordeiro VK, Armstrong LCT, Hungria M (2011) Genetic diversity of rhizobia in a Brazilian oxisol nodulating Mesoamerican and Andean genotypes of common bean (Phaseolus vulgaris L.). World J Microbiol Biotechnol 27:643–650
Pavel BA, Vasile CI (2012) PyElph—a software tool for gel images analysis and phylogenetices. BMC Bioinform 13:9
Pérez-Ramírez NO, Rogel MA, Wang E, Castellanos JZ, Martínez-Romero E (1998) Seeds of Phaseolus vulgaris bean carry Rhizobium etli. FEMS Microbiol Ecol 26:289–296
Prévost D, Antoun H (2007) Root nodule bacteria and symbiotic nitrogen fixation, Chapter 31. In: Carter MR, Gregorich EG (eds) Soil sampling and methods of analysis. CRC Press, Boca Raton, pp 379–397
Rahmani ΗΑ, Räsänen LA, Afshari M, Lindström K (2011) Genetic diversity and symbiotic effectiveness of rhizobia isolated from root nodules of Phaseolus vulgaris L. grown in soils of Iran. Appl Soil Ecol 48:287–293
Redecker D, von Berswordt-Wallrabe P, Beck DP, Werner D (1997) Influence of inoculation with arbuscular mycorrhizal fungi on stable isotopes of nitrogen in Phaseolus vulgaris. Biol Fert Soils 24:344–346
Rodriguez-Navarro DN, Buendia AM, Camacho M, Lucas MM, Santamaria C (2000) Characterization of Rhizobium spp. bean isolates from south west Spain. Soil Biol Biochem 32:1601–1613
Santamaría RI, Bustos P, Pérez-Carrascal OM, Miranda-Sánchez F, Vinuesa P, Martínez-Flores I, Juárez S, Lozano L, Martínez-Romero E, Cevallos MA, Romero D, Dávila G, Ormeño-Orrillo E, González V (2017) Complete genome sequences of eight Rhizobium symbionts associated with common bean (Phaseolus vulgaris). Genome Announc 5:e00645–17
Sessitch A, Ramirez-Saad H, Hardarson G, Akkermans ADL, DeVos WM (1997) Classification of Austrian rhizobia and the Mexican isolate FL27 obtained from Phaseolus vulgaris L. as Rhizobium gallicum. Int J Syst Bacteriol 47:1097–1101
Smith SE, Read DJ (1997) Mycorrhizal symbiosis. Academic Press, San Diego
Somasegaran P, Hoben HJ (1985) Methods in legume-rhizobium technology. University of Hawaii NifTAL Project and MIRCEN
Sylvia DM (1994) Vesicular-arbuscular mycorrhizal (VAM) fungi. In: Weaver RW, Angle JS, Bottomley PJ, Bezdicek D, Smith S, Tabatabai A, Wollum AG (eds) Methods of soil analysis, Part 2. Microbiological and biochemical properties, vol 5. Soil Science Society of America, Madison, pp 351–378
Tamimi SM (2002) Genetic diversity and symbiotic effectiveness of rhizobia isolated from root nodules of common bean (Phaseolus vulgaris L.) grown in the soils of the Jordan valley. Appl Soil Ecol 19:183–190
Tertivanidis K, Koutita O, Papadopoulos II, Tokatlidis IS, Tamoutsidis EG, Pappa-Michailidou V, Koutsika-Sotiriou M (2008) Genetic diversity in bean populations based on random amplified polymorphic DNA markers. Biotechnology 7:1–9
Thies JE, Bohlool BB, Singleton PW (1992) Environmental effects on competition for nodule occupancy between introduced and indigenous rhizobia and among introduced strains. Can J Microbiol 38:493–500
Valverde A, Ingua JM, Peix A, Cervantes E, Velásquez E (2006) Rhizobium lusitanum sp. nov. a bacterium that nodulates Phaseolus vulgaris. Int J Syst Evol Microbiol 56:2631–2637
Van Cauwenberghe J, Verstraete B, Lemaire B, Lievens B, Michiels J, Honnay O (2014) Population structure of root nodulating Rhizobium leguminosarum in Vicia cracca populations at local to regional geographic scales. Syst Appl Microbiol 37:613–621
Wang L, Cao Y, Wang ET, Qiao YJ, Jiao S, Liu ZS, Zhao L, Wei GH (2016) Biodiversity and biogeography of rhizobia associated with common bean (Phaseolus vulgaris L.) in Shaanxi province. Syst Appl Microbiol 39:211–219
Wei GH, Zhang ZX, Chen C, Chen WM, Ju WT (2008) Phenotypic and genetic diversity of rhizobia isolated from nodules of the legume genera Astragalus, Lespedeza and Hedysarum in northwestern China. Microbiol Res 163:651–662
Yan H, Ji ZJ, Jiao YS, Wang ET, Chen WF, Guo BL, Chen WX (2016) Genetic diversity and distribution of rhizobia associated with the medicinal legumes Astragalus spp. and Hedysarum polybotrys in agricultural soils. Syst Appl Microbiol 39:141–149
Zurdo-Piñeiro JL, García-Fraile P, Rivas R, Peix A, León-Barrios M, Willems A, Mateos PF, Martínez-Molina E, Velázquez E, van Berkum P (2009) Rhizobia from Lanzarote, the Canary Islands, that nodulate Phaseolus vulgaris have characteristics in common with Sinorhizobium meliloti isolates from mainland Spain. Appl Environ Microbiol 75:2354–2359
Acknowledgements
This work was supported by the Research Committee of Aristotle University of Thessaloniki (Grant number 89313). The authors would also like to thank Kyriaki Kosmanidou and the “Pelecanos” cooperative for their assistance in field sampling.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Yusuf Akhter.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
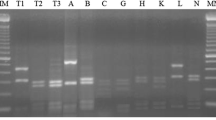
S 1
. Agarose gel of the PCR products where multiple or single band may be observed before (left) and after HaeII digestion (right). Highlighted are the bands that were successfully sequenced. M: 100 base DNA ladder marker, 1: PYL2, 2: ORM3, 3: ORM1, 4: PYL6, 5: PYL7, 6: SLL3, 7: SLL7, 8: PYL6, 9: KAR6, 10: ORM4, 11: SLL8, 12: PYL5, 13: ERG2, 14: LAI3 (DOCX 142 kb)
S 2
. Agarose gel of the PCR products where multiple or single band may be observed before (up) and after HaeII digestion (down). Highlighted are the bands that were successfully sequenced. M: 100 base DNA ladder marker, 1: ORM2, 2: PYL3, 3: KAR1, 4: KAR7, 5: KAR9, 6: SLL1, 7: PYL2, 8: MIK3, 9: KAR2, 10: KAR8, 11: SLL6, 12: SLA2, 13: LAI2, 14: PYL4, 16: SLL12, 17: SLA5, 18: OPA1, 19: GRA4, 20: GRA5, 21: PYL1, 22: PYL9, 23: SLA1, 24: LAI5, 25: GRA2, 26: SLL4, 27: GRA3, 28: MIK 7, 29: MIK5, 30: JUM1, 31: MIK2, 32: LAI3, 33: GRA1, 34: LAI6, 35: LAI4, 36: OPA2, 37: MIK1 38: SLA4, 39: LAI8, 40: LAI7, 41: SLL5, 42: KAR4, 43: MIK4, 44: KAR5, 45: SLL13, 46: MIK6, 47: ERG1, 48: SLL2, 49: PYL10, 50: PYL11, 51: ORM5, 52: KAR10, 53: KAR11, 54: ORM6, 55: KAR12. Note that JUM1 is an isolate from Florina, an area close, but outside the Prespa lakes area (DOCX 328 kb)
S 3.
Agarose gel of the PCR products after MspI digestion. M: 100 base DNA ladder marker. 1: ERG1, 2: SLL11, 3: MIK6, 4: KAR3, 6: KAR6, 7: PYL5, 8: KAR1, 9: KAR4, 10: KAR7, 11: KAR9, 12: LAI6, 13: GRA1, 14: LAI7, 15: LAI1, 16: GRA1, 17: KAR5, 18: MIK4, 19: OPA1, 21: GRA2, 22: MIK5, 23: MIK2, 24: MIK7, 25: MIK3, 26: PYL2, 27: LAI3, 28: ERG2, 30: SLL3, 31: SLL7, 32: ORM1, 33: SLL4, 34: PYL4, 36: GRA2, 37: KAR2, 38: SLA1, 39: SLA1, 40: SLL1, 43: KAR3, 44: LAI2, 45: PYL6, 46: PYL7, 47: SLA2, 48: KAR8, 50: KAR9, 51: SLL7, 52: SLL12, 53: GRA3, 54: ERG1 (DOCX 200 kb)
Rights and permissions
About this article
Cite this article
Ipsilantis, I., Lotos, L. & Tsialtas, I.T. Diversity and nodulation effectiveness of rhizobia and mycorrhizal presence in climbing dry beans grown in Prespa lakes plain, Greece. Arch Microbiol 201, 1151–1161 (2019). https://doi.org/10.1007/s00203-019-01679-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-019-01679-z