Abstract
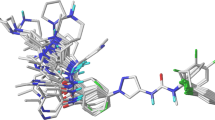
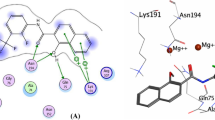
c-Jun N-terminal kinase is an important regulator, activating several transcription factors in response to proinflammatory cytokines, ultraviolet radiations, environmental stress, hypoxia and osmotic shock and is known to be reported a cause for many diseases, such as diabetes, cancer, inflammation, stroke, etc. In the present study, we aim to predict novel therapeutic leads against c-Jun N-terminal kinase-3 by employing structure based virtual screening in combination with various in silico toxicity filters. We screened ZINC database virtually using a known potent c-Jun N-terminal kinase inhibitor, SP600125, as reference molecule. We obtained 128 molecules sharing ≥70% structure identity with SP600125. These 128 compounds were subjected to virtual screening and various toxicity filters. Finally, three molecules were identified as novel c-Jun N-terminal kinase inhibitors. Further binding mode analysis suggested that these molecules inhibit c-Jun N-terminal kinase activity through binding the adenosine triphosphate binding pocket.


Similar content being viewed by others
References
Asano Y, Kitamura S, Ohra T, Itoh F, Kajino M, Tamura T, Kaneko M, Ikeda S, Igata H, Kawamoto T, Sogabe S (2008) Discovery, synthesis and biological evaluation of isoquinolones as novel and highly selective JNK inhibitors (2). Bioorg Med Chem 16:4699–4714
Bennett BL, Sasaki DT, Murray BW, O’Leary EC, Sakata ST, Xu W, Leisten JC, Motiwala A, Pierce S, Satoh Y, Bhagwat SS (2001) SP600125, an anthrapyrazolone inhibitor of Jun N-terminal kinase. Proc Natl Acad Sci 98:13681–13686
Bode AM, Dong Z (2007) The functional contrariety of JNK. Mol Carcinog 46:591–598
Dallakyan S, Olson AJ (2015) Small-molecule library screening by docking with PyRx. Chem Biol 1263:243–250
Gupta S, Barrett T, Whitmarsh AJ, Cavanagh J, Sluss HK, Derijard B, Davis RJ (1996) Selective interaction of JNK protein kinase isoforms with transcription factors. EMBO J 15:2760
Hibi M, Lin AN, Smeal T, Minden A, Karin M (1993) Identification of an oncoprotein-and UV-responsive protein kinase that binds and potentiates the c-Jun activation domain. Genes Dev 7:2135–2148
Irwin JJ, Shoichet BK (2005) ZINC-a free database of commercially available compounds for virtual screening. J Chem Inf model 45:177–182
Kyriakis JM, Avruch J (1990) pp54 microtubule-associated protein 2 kinase. A novel serine/threonine protein kinase regulated by phosphorylation and stimulated by poly-l-lysine. J Biol Chem 265:17355–17363
Lajiness MS, Vieth M, Erickson J (2004) Molecular properties that influence oral drug-like behavior. Curr Opin Drug Discov Dev 7:470–477
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2012) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 64:4–17
Meng XY, Zhang HX, Mezei M, Cui M (2011) Molecular docking: a powerful approach for structure-based drug discovery. Curr Comput Aided Drug Des 7:146–157
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Probst GD, Bowers S, Sealy JM, Truong AP, Hom RK, Galemmo RA, Konradi AW, Sham HL, Quincy DA, Pan H, Yao N (2011) Highly selective c-Jun N-terminal kinase (JNK) 2 and 3 inhibitors with in vitro CNS-like pharmacokinetic properties prevent neurodegeneration. Bioorg Med Chem Lett 21:315–319
Sander T, Freyss J, von Korff M, Rufener C (2015) Data warrior: an open-source program for chemistry aware data visualization and analysis. J Chem Inf Model 55:460–473
Stierand K, Rarey M (2010) Poseview–molecular interaction patterns at a glance. J Chem Inform 2:1
Vyas VK, Ghate M, Goel A (2013) Pharmacophore modeling, virtual screening, docking and in silico ADMET analysis of protein kinase B (PKB β) inhibitors. J Mol Graph Model 42:17–25
Xie X, Gu Y, Fox T, Coll JT, Fleming MA, Markland W, Caron PR, Wilson KP, Su MS (1998) Crystal structure of JNK3: a kinase implicated in neuronal apoptosis. Structure 6:983–991
Zhou YY, Li Y, Jiang WQ, Zhou LF (2015) MAPK/JNK signalling: a potential autophagy regulation pathway. Biosci Rep 35:e00199
Acknowledgements
DT is thankful to University Grants Commission, New Delhi for financial support in the form of UGC—BSR (RFSMS) Senior Research Fellowship. Y.S. is thankful to Human Resource Development for Health Research, New Delhi (F.No.V.25011/542-HRD/2016-HR). We are very grateful to the Journal Editor, anonymous reviewers, and Prof. P. Sreedhara Reddy, Department of Physics, Sri Venkateswara University, Tirupati for their constructive and useful comments which improve the scientific content of the original paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Daggupati, T., Chitrala, K., Pamanji, R. et al. Molecular screening and analysis of novel therapeutic inhibitors against c-Jun N-terminal kinase. Med Chem Res 26, 2112–2118 (2017). https://doi.org/10.1007/s00044-017-1919-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-017-1919-5