Abstract
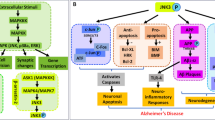
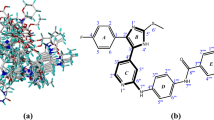
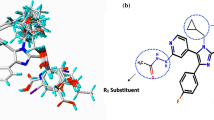
JNK3, a protein kinase of the MAPK family that is potently activated by a variety of environmental stress and pro-inflammatory cytokines, has been recognized as an important therapeutic target for several neurodegenerative diseases. However, due to the long development cycle and the cost of R&D, no related drugs have been released. By applying the CoMFA and CoMSIA methods, QSAR models that explore the structure-activity relationship of the JNK3 inhibitors are established and validated, the parameters are satisfactory (q2 = 0.774, r2 = 0.991 for CoMFA model and q2 = 0.666, r2 = 0.990 for CoMSIA model) and base on which a series of novel molecules are designed. Molecular docking is conducted to verify the potential of the new compounds, the results shows promising activity. We speculated that the series of compounds S1–S11 might have some therapeutic activity in the c-jnk3 pathway, particularly the “S8” may prove to be the best one, which provided useful guidance for the design of new and efficient subtype selective JNK3 inhibitors.







Similar content being viewed by others
References
Ansideri F, Macedo JT, Eitel M, El-Gokha A, Zinad DS, Scarpellini C, Laufer SA (2018) Structural Optimization of a Pyridinylimidazole scaffold: shifting the selectivity from p38α mitogen-activated protein kinase to c-Jun N-terminal kinase 3. ACS Omega 3(7):7809–7831
Buckingham J, Glen RC, Hill AP, Hyde RM, Martin GR, Robertson AD, Woollard PM (1995) Computer-aided design and synthesis of 5-substituted tryptamines and their pharmacology at the 5-HT1D receptor: discovery of compounds with potential anti-migraine properties. J Med Chem 38(18):3566–3580
Cervigni RI, Bonavita R, Barretta ML, Spano D, Ayala I, Nakamura N, Colanzi A (2015) JNK2 controls fragmentation of the Golgi complex and the G2/M transition through phosphorylation of GRASP65. J Cell Sci 128(12):2249–2260
Chambers JW, Howard S, LoGrasso PV (2013) Blocking c-Jun N-terminal kinase (JNK) translocation to the mitochondria prevents 6-hydroxydopamine-induced toxicity in vitro and in vivo. J Biol Chem 288(2):1079–1087
Chambers JW, Pachori A, Howard S, Ganno M, Hansen Jr D, Kamenecka T, Cherry L (2011) Small molecule c-jun-N-terminal kinase inhibitors protect dopaminergic neurons in a model of Parkinson’s disease. Acs Chem Neurosci 2(4):198–206
Dérijard B, Hibi M, Wu IH, Barrett T, Su B, Deng T, Davis RJ (1994) JNK1: a protein kinase stimulated by UV light and Ha-Ras that binds and phosphorylates the c-Jun activation domain. Cell 76(6):1025–1037
Dorsey BD, Levin RB, McDaniel SL, Vacca JP, Guare JP, Darke PL, Schleif WA (1994) L-735,524: the design of a potent and orally bioavailable HIV protease inhibitor. J Med Chem 37(21):3443–3451
Gupta S, Barrett T, Whitmarsh AJ, Cavanagh J, Sluss HK, Derijard B, Davis RJ (1996) Selective interaction of JNK protein kinase isoforms with transcription factors. EMBO J 15(11):2760–2770
Gutierrez GJ, Tsuji T, Chen M, Jiang W, Ze’ev AR (2010) Interplay between Cdh1 and JNK activity during the cell cycle. Nat Cell Biol 12(7):686
Hibi M, Lin ANNING, Smeal T, Minden A, Karin M (1993) Identification of an oncoprotein-and UV-responsive protein kinase that binds and potentiates the c-Jun activation domain. Gene dev 7(11):2135–2148
Huang X, Liu T, Gu J, Luo X, Ji R, Cao Y, Jiang H (2001) 3D-QSAR model of flavonoids binding at benzodiazepine site in GABAA receptors. J Med Chem 44(12):1883–1891
Jain AN (2003) Surflex: fully automatic flexible molecular docking using a molecular similarity-based search engine. J Med Chem 46(4):499–511
Kamenecka T, Jiang R, Song X, Duckett D, Chen W, Ling YY, Cameron MD (2009) Synthesis, biological evaluation, X-ray structure, and pharmacokinetics of aminopyrimidine c-jun-N-terminal kinase (JNK) inhibitors. J Med Chem 53(1):419–431
Klebe G, Abraham U (1999) Comparative molecular similarity index analysis (CoMSIA) to study hydrogen-bonding properties and to score combinatorial libraries. J Comput Aid Mol Des 13(1):1–10
Koch P, Gehringer M, Laufer SA (2014) Inhibitors of c-Jun N-terminal kinases: an update. J Med Chem 58(1):72–95
Krenitsky VP, Nadolny L, Delgado M, Ayala L, Clareen SS, Hilgraf R, Wright J (2012) Discovery of CC-930, an orally active anti-fibrotic JNK inhibitor. Bioorg Med Chem Lett 22(3):1433–1438
Kyriakis JM, Banerjee P, Nikolakaki E, Dai T, Rubie EA, Ahmad MF, Woodgett JR (1994) The stress-activated protein kinase subfamily of c-Jun kinases. Nature 369(6476):156
Lange A, Günther M, Büttner FM, Zimmermann MO, Heidrich J, Hennig S, Koch P (2015) Targeting the gatekeeper MET146 of C-Jun N-terminal kinase 3 induces a bivalent halogen/chalcogen bond. J Am Chem Soc 137(46):14640–14652
Li S, Fan J, Peng C, Chang Y, Guo L, Hou J, Xiao G (2017) New molecular insights into the tyrosyl-tRNA synthase inhibitors: CoMFA, CoMSIA analyses and molecular docking studies. Sci Rep 7(1):11525
Mishra P, Günther S (2018) New insights into the structural dynamics of the kinase JNK3. Sci Rep 8(1):9435
Mohit AA, Martin JH, Miller CA (1995) p493F12 kinase: a novel MAP kinase expressed in a subset of neurons in the human nervous system. Neuron 14(1):67–78
Muth F, El-Gokha A, Ansideri F, Eitel M, Döring E, Sievers-Engler A, Laufer SA (2017) Tri-and tetrasubstituted pyridinylimidazoles as covalent inhibitors of c-Jun N-terminal kinase 3. J Med Chem 60(2):594–607
Qin LT, Liu SS, Xiao QF, Wu QS (2013) Internal and external validtions of QSAR model: Review. Environ Chem 32(7):1205–1211
Rech JC, Yato M, Duckett D, Ember B, LoGrasso PV, Bergman RG, Ellman JA (2007) Synthesis of potent bicyclic bisarylimidazole c-jun N-terminal kinase inhibitors by catalytic C−H bond activation. J Am Chem Soc 129(3):490–491
Schattel V, Hinselmann G, Jahn A, Zell A, Laufer S (2011) Modeling and benchmark data set for the inhibition of c-Jun N-terminal kinase-3. J Chem Inf Model 51(3):670–679
Sharma P, Ghoshal N (2006) Exploration of a binding mode of benzothiazol-2-yl acetonitrile pyrimidine core based derivatives as potent c-Jun N-terminal kinase-3 inhibitors and 3D-QSAR analyses. J Chem Inf Model 46(4):1763–1774
Siddiqui MA, Reddy PA (2010) Small molecule JNK (c-Jun N-terminal kinase) inhibitors. J Med Chem 53(8):3005–3012
Srivastava V, Kumar A, Mishra BN, Siddiqi MI (2008) Molecular docking studies on DMDP derivatives as human DHFR inhibitors. Bioinformation 3(4):180
Wang Z, Chang Y, Han Y, Liu K, Hou J, Dai C, Chen W (2016) 3D-QSAR and docking studies on 1-hydroxypyridin-2-one compounds as mutant isocitrate dehydrogenase 1 inhibitors. J Mol Struct 1123:335–343
Xiao A, Zhang Z, An L, Xiang Y (2008) 3D-QSAR and docking studies of 3-arylquinazolinethione derivatives as selective estrogen receptor modulators. J Mol Model 14(2):149–159
Zhao H, Serby MD, Xin Z, Szczepankiewicz BG, Liu M, Kosogof C, Pederson T (2006) Discovery of potent, highly selective, and orally bioavailable pyridine carboxamide c-Jun NH2-terminal kinase inhibitors. J Med Chem 49(15):4455–4458
Zheng K, Iqbal S, Hernandez P, Park H, LoGrasso PV, Feng Y (2014) Design and synthesis of highly potent and isoform selective JNK3 inhibitors: SAR studies on aminopyrazole derivatives. J Med Chem 57(23):10013–10030
Acknowledgements
This work was supported by the Natural Science Foundation of China (No. 81573294, 81773593, 81872759), the Excellent Young Teachers Program of Guangdong Provincial Colleges and Universities (YQ 2015061) and the Pearl River S&T Nova Program of Guangzhou (201610010100). We thank the software support from Dr. Junxia Zheng of the Guangdong University of Technology.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Yanda Liu, Yewei Xie
Rights and permissions
About this article
Cite this article
Liu, Y., Xie, Y., Liu, Y. et al. Insights into the c-Jun N-terminal kinase 3 (JNK3) inhibitors: CoMFA, CoMSIA analyses and molecular docking studies. Med Chem Res 28, 1796–1805 (2019). https://doi.org/10.1007/s00044-019-02416-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-019-02416-3