Abstract
Self-incompatibility (SI) in angiosperms prevents inbreeding and promotes outcrossing to generate genetic diversity. SI in the Brassicaceae is controlled by the S-haplotype-specific interaction between pollen ligand (S-locus protein 11, SP11 or SCR) and its stigmatic receptor (S-receptor kinase, SRK). SP11/SCR binding to cognate SRK induces autophosphorylation of SRK, which triggers a signaling cascade leading to the rejection of self-pollen. However, the mechanism of self-pollen rejection downstream of this ligand–receptor interaction is unknown. Here, we generated self-incompatible Arabidopsis thaliana accession C24 for the forward-genetic approach and live-cell imaging of SI in the Brassicaceae. Furthermore, for reverse-genetic analysis, we extended the Arabidopsis Targeting Induced Local Lesions IN Genomes (TILLING) resources by developing a new population of ethyl methanesulfonate (EMS)-induced mutant lines in A. thaliana accession C24. We believe that the reverse-genetic approach is a useful tool for identifying genes that function in the SI signaling pathway of the Brassicaceae.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Angiosperms have developed self-incompatibility (SI) as a genetic system to prevent inbreeding and thereby promote outcrossing to generate genetic diversity. SI is based on the self/non-self discrimination between male and female. In many angiosperms, SI is controlled by a single locus, designated S, with multiple haplotypes (de Nettancourt 2001). Each S-haplotype encodes both male-specificity and female-specificity determinants (S-determinants), and self/non-self discrimination is accomplished by the S-haplotype-specific interaction between these S-determinants. Both the male and female determinants are polymorphic and are inherited as one segregating unit. The variants of this gene complex are called S-haplotypes. Self/non-self recognition operates at the level of protein–protein interactions between the two determinants, and an incompatible response occurs when both determinants originate from the same S-haplotype.
2 Self/Non-Self Recognition System in the Brassicaceae
In the Brassicaceae, the self/non-self discrimination between male and female occurs on papilla cells covering the stigma surface of the pistil. When cross-pollen lands on the papilla cell, the pollen hydrates and germinates. The pollen tube penetrates the surface of the papilla cell and enters the style, ultimately resulting in cross-fertilization. By contrast, when self-pollen lands on the papilla cell, pollen hydration and germination are inhibited (Fig. 21.1).
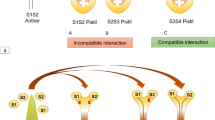
The female determinant is S-receptor kinase (SRK) (Takasaki et al. 2000). SRK consists of an SLG-like extracellular domain, a transmembrane domain, and an intracellular serine/threonine kinase domain. SRK spans the plasma membrane of the stigma papilla cell. The male determinant is S-locus protein 11 (SP11; also called S-locus cysteine-rich protein, SCR) (Schopfer et al. 1999; Takayama et al. 2000). SP11 is a small basic cysteine-rich protein that is predominantly expressed in the anther tapetum and accumulates in the pollen coat during pollen maturation (Iwano et al. 2003) (Fig. 21.2). Upon pollination, SP11/SCR penetrates the papilla cell wall and binds SRK in an S-haplotype-specific manner. This binding induces the autophosphorylation of SRK, triggering a signaling cascade that results in the rejection of self-pollen (Takayama et al. 2001; Takayama and Isogai 2005; Iwano and Takayama 2012) (Fig. 21.3).
Self-recognition self-incompatibility (SI) in Brassicaceae. The S-locus encodes female and male S-determinants, designated SRK and SP11 (or SCR), respectively. In self-pollination, the self (same S-haplotype)-specific SP11 binding to its cognate SRK stabilizes SRK in an active dimeric form on plasma membrane (PM), which triggers SI responses in the stigmatic papilla cell. MLPK and ARC1 were the only SP11/SRK downstream signaling molecular candidates
The self-recognition, that is, the S-haplotype-specific interaction between SP11 and its cognate SRK, has been shown by a series of biochemical studies in Brassica rapa. A binding experiment using 125I-labeled-S 8-SP11 suggested that it strongly binds to the stigmatic membrane of S 8-haplotype (K d = 0.7 nM) but not of the S-haplotype (Takayama et al. 2001). Cross-linking and immunological analyses suggested that 125I-labeled-S 8-SP11 directly binds to S 8-SRK and a 60-kDa protein in the stigmatic membrane of S 8-haplotype (Takayama et al. 2001). Affinity purification and LC-MS/MS analysis of SP11-binding stigmatic proteins have revealed that the 60-kDa protein is a truncated form of SRK (tSRK) containing the extracellular, transmembrane, and part of the intracellular juxtamembrane domains (Shimosato et al. 2007). Interestingly, an artificially expressed dimerized form of eSRK exhibited high-affinity binding to SP11. Another recent study suggested that two regions in the extracellular domain of SRK mediated the homo-dimerization of eSRK (Naithani et al. 2007). Taken together, these studies suggested that SRK on the stigmatic membrane is in an equilibrium between the inactive monomeric or dimeric low-affinity forms and the dimeric active high-affinity form, and that the SP11/SCR binding to its cognate SRK stabilizes its dimeric active form, which is expected to trigger the SI responses in the papilla cell (Shimosato et al. 2009) (Fig. 21.3).
3 SI Signaling Cascade Leading to Rejection of Self-Pollen
To date, the only candidates for signaling molecules acting downstream of SP11/SRK have been MLPK, the membrane-anchored M-locus protein kinase (Murase et al. 2004), and ARC1, an arm repeat-containing protein with E3 ubiquitin-ligase activity (Stone et al. 1999) (Fig. 21.3). MLPK was identified as a positive mediator of SI signaling in a genetic analysis of a self-compatible mutant of B. rapa var. Yellow Sarson (Murase et al. 2004). Upon self-pollination, MLPK is thought to interact directly with SRK to form an SRK-MLPK receptor complex on the plasma membrane and enhance SI signaling (Kakita et al. 2007). ARC1 is a potent positive mediator of this signal transduction pathway (Stone et al. 2003), and can be phosphorylated in vitro by both SRK and MLPK, suggesting that ARC1 is recruited to an SRK-MLPK complex at the plasma membrane (Samuel et al. 2008). ARC1 is predicted to promote self-pollen rejection in the self-incompatibility response by negatively regulating Exo70A1 and blocking the delivery of secretory vesicles to the pollen contact site (Samuel et al. 2009). However, the observation that suppression of ARC1 expression results in incomplete breakdown of SI in both Brassica napus and Arabidopsis lyrata (Stone et al. 1999; Indriolo et al. 2012) might suggest the existence of another unknown signaling pathway acting downstream of SP11/ SRK.
Pollen hydration is the earliest step of cross-pollination. The regulation of water transport from a papilla cell to a pollen grain is one of the most important steps in the rejection of self-pollen. The lipid and proteinaceous components of the pollen coat are essential to pollen hydration during pollen-foot formation in Brassica oleracea (Elleman and Dickinson 1986). In B. rapa, monitoring transiently expressed GFP-mTalin, and rhodamine-phalloidin staining showed the concentration of actin bundles at the cross-pollen attachment site and actin reorganization at the self-pollen attachment site (Iwano et al. 2007). Additionally, the application of cross-pollen coat induces actin polymerization in the apical region of the papilla, whereas the application of self-pollen coat is associated with a decrease in actin filaments in the apical region. The actin-depolymerizing drug cytochalasin D (CD) significantly inhibited pollen hydration and germination during cross-pollination, further emphasizing a role for actin in these processes. Furthermore, electron tomography using ultrahigh-voltage electron microscopy revealed the close association of the actin cytoskeleton with an apical vacuole network. Self-pollination disrupted the vacuole network, whereas cross-pollination led to vacuolar rearrangements toward the site of pollen attachment. Taken together, these data suggested that self- and cross-pollination differentially affect the dynamics of the actin cytoskeleton, leading to changes in vacuolar structure that might be associated with hydration and germination (Iwano et al. 2007). In B. napus and Arabidopsis thaliana, immunostaining using anti-tubulin antibodies found that moderate changes in the microtubule network were observed after self-incompatible pollinations, but a more distinct localized breakdown of the microtubule network was observed during compatible pollinations (Samuel et al. 2011). Visualization over time of the morphological and physiological changes in the stigmatic papilla cell during self- and cross-pollination is a useful method for understanding SI. However, the application of live-cell imaging to B. rapa and B. oleracea has been difficult because of the low efficiency of transformation and the variability of pollination timing. To investigate the SI downstream signaling cascade leading to rejection of self-pollen in the Brassicaceae, generation of SI Arabidopsis was thought to be useful.
4 Generations of SI Arabidopsis
Arabidopsis thaliana is a popular model plant in the Brassicaceae. A. thaliana is the first plant to have its genome sequenced and is a popular tool for understanding the molecular biology of many plant traits. Its small size, rapid life cycle, and easy genetic transformation are also advantageous for research. In addition, this plant is well suited for live-cell imaging during pollination. However, A. thaliana is a self-compatible species; by contrast, Arabidopsis lyrata is a self-incompatible species in genus Arabidopsis. Previously, a comparative analysis of the S-locus region of A. lyrata and its homologous region in A. thaliana (Col-0) identified orthologues of the SRK and SCR genes (Kusaba et al. 2001). However, none of the three candidate SCR orthologues was predicted to encode full-length SCR proteins; therefore, they were designated ΨSCR1, ΨSCR2, and ΨSCR3. The predicted SRK orthologue was also thought to be inactive because it contains a premature stop codon. Thus, self-compatibility in A. thaliana is associated with the inactivation of SI specificity genes. Introduction of functional SP11/SCR and SRK gene pairs isolated from A. lyrata into A. thaliana accession C24 conferred stable SI responses (Nasrallah et al. 2004). In the resulting SI Arabidopsis, pollen hydration and germination were arrested after self-pollination with SP11/SCR pollen, but normal pollen germination and pollen tube penetration were observed after pollination with wild-type (WT) pollen (SC). The establishment of the monitoring systems using this SI Arabidopsis is a useful tool to visualize the physiological events during SI- and SC-pollination.
5 A TILLING Resource for Functional Genomics in Arabidopsis thaliana Accession C24
Many reverse-genetic resources have been developed for functional genetic studies. Because site-directed mutagenesis is not effective in plants, random mutagenesis approaches, including insertional (Wisman et al. 1998; Alonso et al. 2003), chemical (McCallum et al. 2000), and fast neutron mutagenesis (Li et al. 2001), have been used to establish reverse-genetic platforms. In Arabidopsis, insertional mutagenic techniques using T-DNA or transposons have become popular tools for functional genomics. However, insertional mutagenesis often leads to complete gene knockouts, making it difficult to associate nuanced phenotypes with essential genes (Jander et al. 2002). Similarly, radiation mutagenesis, for example, fast neutrons, often induces large genomic deletions that affect multiple genes (Li et al. 2001). By contrast, classical chemical mutagenesis using a mutagen such as ethyl methanesulfonate (EMS) induces an array of interesting point mutations with different impacts on gene function. Such allelic series are desirable because they generate a wide repertoire of mutant phenotypes covering a range of severity, which provide more insight into a gene’s function. Moreover, individual plants carrying point mutations can be identified easily through a powerful method called TILLING (Targeting Induced Local Lesions IN Genomes).
TILLING is a reverse-genetic method that takes advantages of classical mutagenesis, sequence databases, and high-throughput PCR-based screening for point mutations in a targeted sequence (Henikoff et al. 2004). The key advantage of TILLING over competing methods is that it can be applied to any plant species, regardless of ploidy level, genome size, or genetic background (Kurowska et al. 2011). TILLING extends genomic resources, particularly in organisms lacking reverse-genetic tools, where mutants with a range of phenotypic severity are highly desirable. Since the inception of TILLING, this method has been applied to various organisms including Cucumis melo L. (González et al. 2011), Solanum lycopersium (Minioa et al. 2010), B. napus (Wang et al. 2008; Harloff et al. 2012), B. oleracea (Himelblau et al. 2009), B. rapa (Stephenson et al. 2010), Lotus japonicus (Perry et al. 2009), Zea mays (Till et al. 2004), Oryza sativa (Till et al. 2007), Drosophila (Winkler et al. 2005), and zebrafish (Wienholds et al. 2003).
To date, Arabidopsis TILLING resources are only available in accessions Columbia (Col-0) (Greene et al. 2003) and Landsberg erecta (Ler) (Martín et al. 2009). Reverse genetic tools for many commonly used Arabidopsis accessions are still limited, in particular accession C24, which is genetically distinct from accession Col-0 (Barth et al. 2002; Törjek et al. 2003). C24 is distinguished physiologically from other familial accessions in terms of tolerance to drought (Bechtold et al. 2010), ozone (Brosche et al. 2010), and frost (Rohde et al. 2004), and enhanced basal resistance to pathogens (Bechtold et al. 2010). The transgenic A. thaliana accession C24 also exhibited a robust and stable self-incompatible (SI) phenotype (Rea et al. 2010), which served as a good model for understanding SI signaling. In addition, a large portion of its genomic sequence was available (Schneeberger et al. 2011), making accession C24 an excellent alternative tool for plant research.
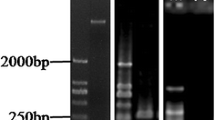
To take advantage of this tool, the Arabidopsis TILLING resources from the GAH molecules by developing a new population of EMS-induced mutant lines in A. thaliana accession C24 X (Lai et al. 2013). From approximately 8,000 A. thaliana C24 seeds treated with 25 mM EMS, 3,620 M1 seedlings were obtained, all of which were used to generate the M2 population. An M2 population with a total of 3,509 individual plants was successfully recovered for use in TILLING. This M2 population also contained 77 partial-seed set lines (semi-sterile) and 125 very low seed set lines (sterile). This population, including semi-sterile and sterile phenotypes, represents a valuable genetic resource for use in forward-genetic screens aimed at isolating novel genes affecting reproduction. Each M2 plant sampled for DNA used in TILLING was originally isolated from a distinct individual M1 plant to ensure independence of the mutations within the population. Ultimately, DNA from M2 plants and M3 seeds from 3,509 lines were stored for TILLING analysis (Lai et al. 2013). The TILLING collection represents the third TILLING resource reported for A. thaliana to date (Table 21.1). TILLING for selected genes from this new collection successfully identified allelic series of induced point mutations, including sense, missense, and nonsense mutations.
6 Conclusion
Self-recognition in the Brassicaceae is clearly mediated by a haplotype-specific interaction between pollen ligand (SP11/SCR) and its stigmatic receptor kinase (SRK). To clarify the downstream signaling pathway leading to self-pollen rejection, SI Arabidopsis was generated by the introduction of functional SP11/SCR and SRK gene pairs isolated from A. lyrata into A. thaliana accession C24 and conferred stable SI responses. The SI Arabidopsis is useful for the forward-genetic approach and live-cell imaging. In addition, TILLING resource was established in A. thaliana accession C24 for reverse genetic approach. The combination of forward- and reverse-genetic approaches with live-cell imaging will be a useful tool for identifying genes that function in the SI signaling pathway in the Brassicaceae.
References
Alonso JM, Stepanova AN, Leisse TJ et al (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Barth S, Melchinger AE, Lubberstedt T (2002) Genetic diversity in Arabidopsis thaliana L. Heynh. investigated by cleaved amplified polymorphic sequences (CAPS) and inter-simple sequence repeat (ISSR) markers. Mol Ecol 11:495–505
Bechtold U, Lawson T, Mejia-Carranza J et al (2010) Constitutive salicylic acid defenses do not compromise seed yield, drought tolerance and water productivity in Arabidopsis accession C24. Plant Cell Environ 33:1959–1973
Brosche M, Merilo E, Mayer F et al (2010) Natural variation in ozone sensitivity among Arabidopsis thaliana accessions and its relation to stomatal conductance. Plant Cell Environ 33:914–925
Elleman CJ, Dickinson HG (1986) Pollen–stigma interactions in Brassica. IV. Structural reorganization in the pollen grains during hydration. J Cell Sci 80:141–157
González M, Xu M, Esteras C et al (2011) Towards a TILLING platform for functional genomics in Piel de Sapo melons. BMC Res Notes 4:289
Greene EA, Codomo CA, Taylor NE et al (2003) Spectrum of chemically induced mutations from a large-scale reverse genetic screen in Arabidopsis. Genetics 164:731–740
Harloff HJ, Lemcke S, Mittasch J et al (2012) A mutation screening platform for rapeseed (Brassica napus L.) and the detection of sinapine biosynthesis mutants. Theor Appl Genet 124:957–969
Henikoff S, Till BJ, Comai L (2004) TILLING: traditional mutagenesis meets functional genomics. Plant Physiol 135:630–636
Himelblau E, Gilchrist EJ, Buono K et al (2009) Forward and reverse genetics of rapid-recycling Brassica oleracea. Theor Appl Genet 118:953–961
Indriolo E, Tharmapalan P, Wright SI, Goring DR (2012) The ARC1 E3 ligase gene is frequently deleted in self-compatible Brassicaceae species and has a conserved role in Arabidopsis lyrata self-pollen rejection. Plant Cell 24:4607–4620
Iwano M, Shiba H, Funato M et al (2003) Immunohistochemical studies on translocation of pollen S-haplotype determinant in self-incompatibility of Brassica rapa. Plant Cell Physiol 44:428–436
Iwano M, Shiba H, Matoba K et al (2007) Actin dynamics in papilla cells of Brassica rapa during self- and cross-pollination. Plant Physiol 144:72–81
Iwano M, Takayama S (2012) Self/non-self discrimination in angiosperm self-incompatibility. Curr Opin Plant Biol 15:78–83
Jander G, Norris SR, Rounsley SD et al (2002) Arabidopsis map-based cloning in the post-genome era. Plant Physiol 129:440–450
Kakita M, Murase K, Iwano M et al (2007) Two distinct forms of M-locus protein kinase localize to the plasma membrane and interact directly with S-locus receptor kinase to transduce self-incompatibility signaling in Brassica rapa. Plant Cell 19:3961–3973
Kurowska M, Daszkowska-Golec A, Gruszka D, Marzec M et al (2011) TILLING: a shortcut in functional genomics. J Appl Genet 52:371–390
Kusaba M, Dwyer K, Hendershot J et al (2001) Self-incompatibility in the genus Arabidopsis: characterization of the S locus in the outcrossing A. lyrata and its autogamous relative A. thaliana. Plant Cell 13:627–643
Lai KS, Kaothien-Nakayama P, Iwano M et al (2013) A TILLING resource for functional genomics in Arabidopsis thaliana accession C24. Genes Genet Syst 87:291–297
Li X, Song Y, Century K, Straight S et al (2001) A fast neutron deletion mutagenesis-based reverse genetics system for plants. Plant J 27:235–242
Martín B, Ramiro M, Martínez-Zapater JM, Alonso-Blanco C (2009) A high-density collection of EMS-induced mutations for TILLING in Landsberg erecta genetic background of Arabidopsis. BMC Plant Biol 9:147
Minioa S, Petrozza A, D’Onofrio O et al (2010) A new mutant genetic resource for tomato crop improvement by TILLING technology. BMC Res Notes 3:69
McCallum CM, Comai L, Greene EA, Henikoff S (2000) Targeted screening for induced mutations. Nat Biotechnol 18:455–457
Murase K, Shiba H, Iwano M et al (2004) A membrane-anchored protein kinase involved in Brassica self-incompatibility signaling. Science 303:1516–1519
Naithani S, Chookajorn T, Ripoll DR, Nasrallah JB (2007) Structural modules for receptor dimerization in the S-locus receptor kinase extracellular domain. Proc Natl Acad Sci USA 104: 12211–12216
Nasrallah ME, Liu P, Sherman-Broyles S et al (2004) Natural variation in expression of self-incompatibility in Arabidopsis thaliana: implications for the evolution of selfing. Proc Natl Acad Sci USA 101:16070–16074
de Nettancourt D (2001) Incompatibility and incongruity in wild and cultivated plants, 2nd edn. Springer, Berlin
Perry J, Brachmann A, Welham T et al (2009) TILLING in Lotus japonicus identified large allelic series for symbiosis genes and revealed a bias in functionally defective ethyl methanesulfonate alleles toward glycine replacements. Plant Physiol 51:1281–1291
Rea AC, Liu P, Nasrallah JB (2010) A transgenic self-incompatible Arabidopsis thaliana model for evolutionary and mechanistic studies of crucifer self-incompatibility. J Exp Bot 61:1897–1906
Rohde P, Hincha DK, Heyer AG (2004) Heterosis in the freezing tolerance of crosses between two Arabidopsis thaliana accessions (Columbia-O and C24) that show differences in non-acclimated and acclimated freezing tolerance. Plant J 38:790–799
Samuel MA, Mudgil Y, Salt JN et al (2008) Interactions between the S-domain receptor kinases and AtPUB-ARM E3 ubiquitin ligases suggest a conserved signaling pathway in Arabidopsis. Plant Physiol 147:2084–2095
Samuel MA, Chong YT, Haasen KE et al (2009) Cellular pathways regulating responses to compatible and self-incompatible pollen in Brassica and Arabidopsis stigmas intersect at Exo70A1, a putative component of the exocyst complex. Plant Cell 21:2655–2671
Samuel MA, Tang W, Jamshed M et al (2011) Proteomic analysis of Brassica stigmatic proteins following the self-incompatibility reaction reveals a role for microtubule dynamics during pollen responses. Mol Cell Proteomics 10:M111.011338
Schneeberger K, Ossowski S, Ott F et al (2011) Reference-guided assembly of four diverse Arabidopsis thaliana genomes. Proc Natl Acad Sci USA 108:10249–10254
Schopfer CR, Nasrallah ME, Nasrallah JB (1999) The male determinant of self-incompatibility in Brassica. Science 286:1697–1700
Shimosato H, Yokota N, Shiba H et al (2007) Characterization of the SP11/SCR high-affinity binding site involved in self/nonself recognition in Brassica self-incompatibility. Plant Cell 19:107–117
Stephenson P, Baker D, Girin T et al (2010) A rich TILLING resource for studying gene function in Brassica rapa. BMC Plant Biol 10:62
Stone SL, Arnoldo M, Goring DR (1999) A breakdown of Brassica self-incompatibility in ARC1 antisense transgenic plants. Science 286:1729–1731
Stone SL, Anderson EM, Mullen RT et al (2003) ARC1 is an E3 ubiquitin ligase and promotes the ubiquitination of proteins during the rejection of self-incompatible Brassica pollen. Plant Cell 15:885–898
Takasaki T, Hatakeyama K, Suzuki G et al (2000) The S receptor kinase determines self-incompatibility in Brassica stigma. Nature (Lond) 403:913–916
Takayama S, Isogai A (2005) Self-incompatibility in plants. Annu Rev Plant Biol 56:467–489
Takayama S, Shiba H, Iwano M et al (2000) The pollen determinant of self-incompatibility in Brassica campestris. Proc Natl Acad Sci USA 97:1920–1925
Takayama S, Shimosato H, Shiba H et al (2001) Direct ligand-receptor complex interaction controls Brassica self-incompatibility. Nature (Lond) 413:534–538
Till BJ, Reynolds SH, Weil C et al (2004) Discovery of induced point mutations in maize genes by TILLING. BMC Plant Biol 4:12
Till BJ, Cooper J, Tai TH et al (2007) Discovery of chemically induced mutations in rice by TILLING. BMC Plant Biol 7:19
Törjek O, Berger D, Meyer RC et al (2003) Establishment of a high-efficiency SNP-based framework marker set for Arabidopsis. Plant J 36:122–140
Wang N, Wang Y, Tian F et al (2008) A functional genomics resource for Brassica napus: development of an EMS mutagenized population and discovery of FAE1 point mutations by TILLING. New Phytol 180:751–765
Wienholds E, van Eeden F, Kosters M et al (2003) Efficient target-selected mutagenesis in zebrafish. Genome Res 13:2700–2707
Winkler S, Schwabedissen A, Backasch D et al (2005) Target-selected mutant screen by TILLING in Drosophila. Genome Res 15:718–723
Wisman E, Hartmann U, Sagasser M et al (1998) Knock-out mutants from an En-1 mutagenized Arabidopsis thaliana population generate phenylpropanoid biosynthesis phenotypes. Proc Natl Acad Sci USA 95:12432–12437
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
This chapter is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
Copyright information
© 2014 The Author(s)
About this paper
Cite this paper
Iwano, M., Ito, K., Shimosato-Asano, H., Lai, KS., Takayama, S. (2014). Self-Incompatibility in the Brassicaceae. In: Sawada, H., Inoue, N., Iwano, M. (eds) Sexual Reproduction in Animals and Plants. Springer, Tokyo. https://doi.org/10.1007/978-4-431-54589-7_21
Download citation
DOI: https://doi.org/10.1007/978-4-431-54589-7_21
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-54588-0
Online ISBN: 978-4-431-54589-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)