Abstract
Background
Studies show that abnormalities in non-coding genes can contribute to carcinogenesis; microRNA levels may modulate cancer growth and metastatic diffusion.
Method
MicroRNA libraries were built and sequenced from two osteosarcoma cell lines (MG-63 and 143B), which differ in proliferation and transmigration. By cloning and transfection, miR-93, expressed in both cell lines, was then investigated for its involvement in osteosarcoma progression.
Results
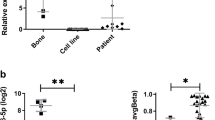
Six of the 19 miRNA identified were expressed in both cell lines with higher expression levels of miR-93 in 143B and in primary osteosarcoma cultures compared to normal osteoblasts. Interestingly, levels of miR-93 were significantly higher in metastases from osteosarcoma than in paired primary tumours. When 143B and MG-63 were transfected with miR-93, clones appeared to respond differently to microRNA overexpression. Ectopic expression of miR-93 more significantly increased cell proliferation and invasivity in 143B than in MG-63 clones. Furthermore, increased mRNA and protein levels of E2F1, one of the potential miR-93 targets, were seen in osteosarcoma cellular clones and its involvement in 143B cell proliferation was confirmed by E2F1 silencing.
Conclusion
Although further studies are needed to evaluate miRNA involvement in osteosarcoma progression, miR-93 overexpression seems to play an important role in osteosarcoma cell growth and invasion.










Similar content being viewed by others
References
M. Campanacci, Bone and soft tissue tumors, 2nd edn. (Springer, Wien, 1999)
S.S. Bielack, B. Kempf-Bielack, G. Delling, G.U. Exner, S. Flege, K. Helmke, R. Kotz, M. Salzer-Kuntschik, M. Werner, W. Winkelmann, A. Zoubek, H. Jurgens, K. Winkler, Prognostic factors in high-grade osteosarcoma of the extremities or trunk: an analysis of 1,702 patients treated on neoadjuvant cooperative osteosarcoma study group protocols. J. Clin. Oncol. 20, 776–790 (2002)
M. Wachtel, B.W. Schäfer, Targets for cancer therapy in childhood sarcomas. Cancer Treat. Rev. 36, 318–327 (2010)
J. Toguchida, T. Nakayama, Molecular genetics of sarcomas: applications to diagnoses and therapy. Cancer Sci. 100, 1573–1580 (2009)
R. Schieckel, B. Boyerinas, S.M. Park, M.E. Peter, MicroRNAs: key players in the immune system, differentiation, tumorigenesis and cell death. Oncogene 27, 5959–5974 (2008)
S.M. Hammond, Dicing and ilincing: the core machinery of the RNA interference pathway. FEBS Lett. 579, 5822–5829 (2005)
A. Fire, S. Xu, M.K. Montgomery, S.A. Kostas, S.E. Driver, C.C. Mello, Potent and specific genetic interference by double stranded RNA in Caenorhabditis elegans. Nature 391, 744–745 (1998)
I. Alvarez-Garcia, E.A. Miska, MicroRNAs functions in animal development and human disease. Development 132, 4653–4662 (2005)
A. Esquela-Kerscher, F.J. Slack, Oncomirs-microRNAs with a role in cancer. Nat. Rev. Cancer 53, 627–635 (2006)
L. Ma, R.A. Weinberg, Micromanagers of malignancy: role of microRNAs in regulating metastasis. Cell 24, 448–456 (2008)
P.M. Voorhoeve, C. le Sage, M. Schrier, A.J. Gillis, H. Stoop, R. Nagel, Y.P. Liu, J. van Duijse, J. Drost, A. Griekspoor, E. Zlotorynski, N. Yabuta, G. De Vita, H. Nojima, L.H. Looijenga, R. Agami, A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumours. Cell 124, 1169–1181 (2006)
Q. Huang, K. Gumireddy, M. Schrier, C. le Sage, R. Nagel, S. Nair, D.A. Egan, A. Li, G. Huang, A.J. Klein-Szanto, P.A. Gimotty, D. Katsaros, G. Coukos, L. Zhang, E. Puré, R. Agami, The microRNAs miR-373 and mir-520c promote tumour invasion and metastasis. Nat. Cell. Biol. 10, 202–210 (2008)
J.A. Chan, A.M. Krichevsky, K.S. Kosik, MicroRNA-21 is an anti-apoptotic factor in human glioblastoma cell. Cancer Res. 65, 6029–6033 (2005)
S.A. Ciafrè, S. Galardi, A. Mangiola, M. Ferracin, C.G. Liu, G. Sabatino, M. Negrini, G. Maira, C.M. Croce, M.G. Farace, Extensive modulation of a set of microRNAs in primary glioblastoma. Biochem. Biophys. Res. Commun. 334, 1351–1358 (2005)
C. Roldo, E. Missiaglia, J.P. Hagan, M. Falconi, P. Capelli, S. Bersani, G.A. Calin, S. Volinia, C.G. Liu, A. Scarpa, C.M. Croce, MicroRNA expression abnormalities in pancreatic and acinar tumors are associated with distinctive pathologic features and clinical behavior. J. Clin. Oncol. 24, 4677–4684 (2006)
F. Yu, H. Yao, P. Zhu, X. Zhang, Q. Pan, C. Gong, Y. Huang, X. Hu, F. Su, J. Lieberman, E. Song, Let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 131, 1109–1123 (2007)
S.F. Tavazoie, C. Alarcon, T. Oskarsson, D. Padua, Q. Wang, P.D. Bos, W.L. Gerald, J. Massague, Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451, 147–152 (2008)
D. Yan, X. Zhou, X. Chen, D.N. Hu, X.D. Dong, J. Wang, F. Lu, L. Tu, J. Qu, MicroRNA-34a inhibits uveal melanoma cell proliferation and migration thrugh down-regulation of c-Met. Invest. Ophthalmol. Vis. Sci. 50, 1559–1565 (2009)
P.A. Gregory, A.G. Bert, E.L. Paterson, S.C. Barry, A. Tsykin, G. Farshid, M.A.Y. Vadas, Y. Khew-Goodall, G.J. Goodall, The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 10, 593–601 (2008)
M. Korpal, E.S. Lee, G. Hu, Y. Kang, The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of e-cadherin transcriptional repressor ZEB1 and ZEB2. J. Biol. Chem. 283, 14910–14914 (2008)
M. Crawford, E. Brawner, K. Batte, L. Yu, M.G. Hunter, G.A. Otterson, G. Nuovo, C.B. Marsh, S.P. Nana-Sinkam, MicroRNA-126 inhibits invasion in non-small cell lung carcinoma cell lines. Biochem. Biophys. Res. Commun. 373, 607–612 (2008)
C. Evangelisti, M.C. Florian, I. Massimi, C. Dominici, G. Giannini, S. Galardi, M.C. Bluè, S. Massalini, H.P. McDowell, E. Messi, A. Gulino, M.G. Farace, S.A. Ciafrè, MiR-128 up-regulation inhibits Reelin and DCX expression and reduces neuroblastoma cell motility and invasiveness. FASEB J. 23, 4276–4287 (2009)
S. Subramanian, W.O. Lui, C.H. Lee, I. Espinosa, T.O. Nielsen, M.C. Heinrich, C.L. Corless, A.Z. Fire, M. van de Rijn, MicroRNA expression signature of human sarcomas. Oncogene 27, 2015–2026 (2008)
C. He, J. Xiong, X. Xu, W. Lu, L. Liu, D. Xiao, D. Wang, Functional elucidation of miR-34 in osteosarcoma cells and primary tumor samples. Biochem. Biophys. Res. Commun. 388, 35–40 (2009)
P. Spessotto, K. Lacrima, P.A. Nicolosi, E. Pivetta, M. Scapolan, R. Perris, Fluorescence-based assays for in vitro analysis of cell adhesion and migration. Methods Mol. Biol. 522, 221–250 (2009)
S.M. Elbashir, W. Lendeckel, T. Tuschl, RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev. 15, 188–200 (2001)
O.A. Kent, J.T. Mendell, A small piece in the cancer puzzle: microRNAs as tumor suppressors and oncogenes. Oncogene 25, 6188–6196 (2006)
W. Zhang, J.E. Dahlberg, W. Tam, MicroRNAs in tumorigenesis: a primer. Am. J. Pathol. 171, 728–738 (2007)
M. Osaki, F. Takeshita, Y. Sugimoto, N. Kosaka, Y. Yamamoto, Y. Yoshioka, E. Kobayashi, T. Yamada, A. Kawai, T. Inoue, H. Ito, M. Oshimura, T. Ochiya, MicroRNA-143 regulates human osteosarcoma metastasis by regulating matrix metalloproteinase-13 expression, Mol. Ther. Mar 22 (2011), [Epub ahead of print], PMID:21427707.
B. Song, Y. Wang, Y. Xi, K. Kudo, S. Bruheim, G.I. Botchkina, E. Gavin, Y. Wan, A. Formentini, M. Kornmann, O. Fodstad, J. Ju, Mechanism of chemoresistance mediated by miR-140 in human osteosarcoma and colon cancer cells. Oncogene 28, 4065–4074 (2009)
A. Gougelet, J. Perez, D. Pissaloux, A. Besse, A. Duc, A.V. Decouvelaere, D. Ranchere-Vince, J.Y. Blay, L. Alberti, miRNA Profiling: how to bypass the current difficulties in the diagnosis and treatment of sarcomas, Sarcoma Feb 22(2011), PMID:21437224.
G. Maire, J.W. Martin, M. Yoshimoto, S. Chilton-MacNeill, M. Zielenska, A. Jeremy, Analysis of miRNA-gene expression-genomic profiles reveals complex mechanisms of microRNA deregulation in osteosarcoma. Cancer Genet. 204, 138–146 (2011)
J. Gao, T.T. Yang, X.C. Qiu, B. Yu, J.W. Han, Q.Y. Fan, B.A. Ma, Cloning and identification of microRNA from human osteosarcoma cell line SOSP-9607. Ai Zheng 26, 561–565 (2007)
S. Baranwal, S.K. Alahari, MiRNA contro and tumour cell invasion and metastasis. Int. J. Cancer 126, 1283–1290 (2010)
W. Ziyan, W. Shuhua, W. Xiufang, L. Xiaoyun, MicroRNA-p21 is involved in osteosarcoma cell invasion and migration, Med. Oncol. May 18 (2010), [Epub ahead of print] PMID: 20480266.
H. Zhang, X. Cai, Y. Wang, H. Tang, D. Tong, F. Ji, MicroRNA 143, down-regulated in osteosarcoma, promotes apoptosis and suppresses tumorigenicity by targeting Bcl-2. Oncol. Rep. 24, 1363–1369 (2010)
Y. Li, W. Tan, T.W. Neo, M.O. Aung, S. Wasser, S.G. Lim, T.M. Tan, Role of miR-106b-25 microRNA cluster in hepatocellular carcinoma. Cancer Sci. 100, 1234–1242 (2009)
C. Volinia, G. Calin, C.G. Liu, S. Ambs, A. Cimmino, F. Petrocca, R. Visone, M. Iorio, C. Roldo, M. Ferracin, R.L. Prueitt, N. Yanaihara, G. Lanza, A. Scarpa, A. Vecchione, M. Negrini, C.C. Harris, C.M. Croce, A microRNA expression segnature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. USA 103, 2257–2261 (2006)
Y. Chen, D.H. Gorski, Regulation of angiogenesis through a microRNA (miR-130a) that down-regulates anti-angiogenic homeobox genes GAX and HOXA5. Blood 111, 1217–1226 (2008)
M.T. Van Joorsveld, J. Helleman, E.M. Berns, E.A. Wiemer, MicroRNAs in ovarian cancer biology and therapy resistance, Int. J. Biochem. Cell. Biol. (2010) 18 [Epub ahead of print] PMID: 20083225.
G. Calin, A. Cimmino, M. Fabbri, M. Ferracin, S.E. Wojcik, M. Shimizu, C. Taccioli, N. Zanesi, R. Garzon, R.I. Aqeilan, H. Alder, S. Volinia, L. Rassenti, X. Liu, C.G. Liu, T.J. Kipps, M. Negrini, C.M. Croce, Mir-15a e Mir-16-1 clusters function in human leukaemia. Proc. Natl. Acad. Sci. USA 105, 5166–5171 (2008)
A. Ventura, A.G. Young, M.M. Winslow, L. Lintault, A. Meissner, S.J. Erkeland, J. Newman, R.T. Bronson, D. Crowley, J.R. Stone, R. Jaenisch, P.A. Sharp, T. Jacks, Targeted dekletion reveals essential and overlapping functions of the miR-17 trough 92 family of miRNA clusters. Cell 132, 875–886 (2008)
C. Xiao, L. Srinivasan, D.P. Calado, H.C. Patterson, B. Zhang, J. Wang, J.M. Henderson, J.L. Kutok, K. Rajewsky, Lymphoprolferative disease and autoimmunity in mice with increased miR-17-92 expression in lymphocytes. Nat. Immunol. 9, 405–414 (2008)
K.M. Foshay, G.I. Gallicano, Small RNAs, big potential: the role of MicroRNAs in stem cell function. Curr. Stem Cell Res. Ther. 2, 264–271 (2007)
F. Petrocca, R. Visone, M.R. Onelli, M.H. Shah, M.S. Nicoloso, I. de Martino, D. Iliopoulos, E. Pilozzi, C.G. Liu, M. Negrini, L. Cavazzini, S. Volinia, H. Alder, L.P. Ruco, G. Baldassarre, C.M. Croce, A. Vecchione, E2F1-regulated microRNAs impair TGFbeta-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell 13, 272–286 (2008)
M.L. Yeung, J. Yasunaga, Y. Bennasser, N. Dusetti, D. Harris, N. Ahmad, M. Matsuoka, K.T. Jeang, Roles of MicroRNAs, miR-93 and miR-103b, and tumor protein 53-induced nuclear protein 1 tumor suppressor in cell growth dysregulation by human T-cell lymphotrophic virus 1. Cancer Res. 68, 8976–8985 (2008)
K.E. Resnick, H. Alder, J.P. Hagan, D.L. Richardson, C.M. Croce, D.E. Cohn, The detection of differentially expressed microRNAs from the serum of ovarian cancer patients using a novel real-time PCR platform. Gynecol. Oncol. 112, 55–59 (2009)
J. Brugarolas, C. Chandrasekaran, J.I. Gordon, D. Beach, T. Jacks, G.J. Hannon, Radiation-induced cell cycle arrest compromised by p21 deficiency. Nature 377, 552–557 (2002)
Acknowledgements
The authors thank Paola Spessotto for transwell and invasion assays, Arcispedale Santa Maria Nuova for providing umbilical cord, Prof. Paolo Malatesta for technical support, Cristina Ghinelli for the graphic work and Alba Balladelli, B.A. for text revision. This work was supported by grants from the Italian Ministry of Health (MSB, RP), Italian Istituto Superiore della Sanità (MSB, RP), EuroBoNeT Consortium (MSB) and Institutional funds from the University of Parma (RP).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Montanini, L., Lasagna, L., Barili, V. et al. MicroRNA cloning and sequencing in osteosarcoma cell lines: differential role of miR-93. Cell Oncol. 35, 29–41 (2012). https://doi.org/10.1007/s13402-011-0059-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-011-0059-z