Abstract
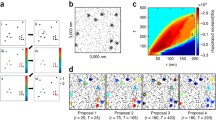
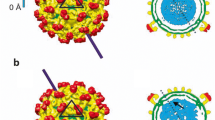
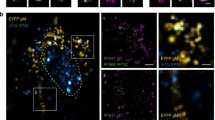
We apply single-molecule super-resolution microscopy and coordinate-based cluster analysis to extract information on the distribution and on the morphology and size of clusters of the human immunodeficiency virus (HIV-1) Gag polyprotein in fixed cells. Three different patterns of Gag distribution could be distinguished. A major type of assembly observed was in accordance with previous electron microscopy analyses revealing ~140 nm-sized HIV-1 buds at the plasma membrane of virus-producing cells. The distribution of Gag molecules in the 2D projection at these sites was consistent with a semi-spherical 3D assembly. We compared different methods of cluster analysis and demonstrated that we can reliably distinguish different distribution patterns of the Gag polyprotein. These methods were applied to extract information on the properties of the different Gag clusters.



Similar content being viewed by others
Abbreviations
- CA:
-
Capsid domain
- dSTORM:
-
Direct stochastic optical reconstruction microscopy
- EMCCD:
-
Electron-multiplying charge device
- HIV-1:
-
Human immunodeficiency virus type 1
- MA:
-
Matrix domain
- MEA:
-
Mercaptoethylamine
- NC:
-
Nucleocapsid domains
- ROI:
-
Region of interest
- Ryr:
-
Ryanodine receptors
- TIRF:
-
Total internal reflection fluorescence
References
Abramoff MDM, Paulo J, Ram, Sunanda J (2004) Image processing with ImageJ. Biophotonics Int 11:36–42
Baddeley D, Jayasinghe ID, Lam L, Rossberger S, Cannell MB, Soeller C (2009) Optical single-channel resolution imaging of the ryanodine receptor distribution in rat cardiac myocytes. Proc Natl Acad Sci USA 106:22275–22280
Betzig E, Patterson GH, Sougrat R, Lindwasser OW, Olenych S, Bonifacino JS, Davidson MW, Lippincott-Schwartz J, Hess HF (2006) Imaging intracellular fluorescent proteins at nanometer resolution. Science 313:1642–1645
Briggs JA, Krausslich HG (2011) The molecular architecture of HIV. J Mol Biol 410:491–500
Briggs JA, Riches JD, Glass B, Bartonova V, Zanetti G, Krausslich HG (2009) Structure and assembly of immature HIV. Proc Natl Acad Sci USA 106:11090–11095
Carlson LA, Briggs JA, Glass B, Riches JD, Simon MN, Johnson MC, Muller B, Grunewald K, Krausslich HG (2008) Three-dimensional analysis of budding sites and released virus suggests a revised model for HIV-1 morphogenesis. Cell Host Microbe 4:592–599
Eckhardt M, Anders M, Muranyi W, Heilemann M, Krijnse-Locker J, Muller B (2011) A SNAP-tagged derivative of HIV-1—a versatile tool to study virus-cell interactions. PLoS ONE 6:e22007
Endesfelder U, Malkusch S, Flottmann B, Mondry J, Liguzinski P, Verveer PJ, Heilemann M (2011) Chemically induced photoswitching of fluorescent probes—a general concept for super-resolution microscopy. Molecules 16:3106–3118
Haase P (1995) Spatial pattern analysis in ecology based on Ripley K-function: introduction and methods of edge correction. J Veg Sci 6:575–582
Heilemann M (2010) Fluorescence microscopy beyond the diffraction limit. J Biotechnol 149:243–251
Heilemann M, van de Linde S, Schuttpelz M, Kasper R, Seefeldt B, Mukherjee A, Tinnefeld P, Sauer M (2008) Subdiffraction-resolution fluorescence imaging with conventional fluorescent probes. Angew Chem Int Ed Engl 47:6172–6176
Heilemann M, van de Linde S, Mukherjee A, Sauer M (2009) Super-resolution imaging with small organic fluorophores. Angew Chem Int Ed Engl 48:6903–6908
Hermida-Matsumoto L, Resh MD (2000) Localization of human immunodeficiency virus type 1 Gag and Env at the plasma membrane by confocal imaging. J Virol 74:8670–8679
Ivanchenko S, Godinez WJ, Lampe M, Krausslich HG, Eils R, Rohr K, Brauchle C, Muller B, Lamb DC (2009) Dynamics of HIV-1 assembly and release. PLoS Pathog 5:e1000652
Jouvenet N, Neil SJ, Bess C, Johnson MC, Virgen CA, Simon SM, Bieniasz PD (2006) Plasma membrane is the site of productive HIV-1 particle assembly. PLoS Biol 4:e435
Jouvenet N, Bieniasz PD, Simon SM (2008) Imaging the biogenesis of individual HIV-1 virions in live cells. Nature 454:236–240
Jouvenet N, Simon SM, Bieniasz PD (2011) Visualizing HIV-1 assembly. J Mol Biol 410:501–511
Kiskowski MA, Hancock JF, Kenworthy AK (2009) On the use of Ripley’s K-function and its derivatives to analyze domain size. Biophys J 97:1095–1103
Lampe M, Briggs JA, Endress T, Glass B, Riegelsberger S, Krausslich HG, Lamb DC, Brauchle C, Muller B (2007) Double-labelled HIV-1 particles for study of virus-cell interaction. Virology 360:92–104
Larson DR, Johnson MC, Webb WW, Vogt VM (2005) Visualization of retrovirus budding with correlated light and electron microscopy. Proc Natl Acad Sci USA 102:15453–15458
Lehmann M, Rocha S, Mangeat B, Blanchet F, Uji IH, Hofkens J, Piguet V (2011) Quantitative multicolor super-resolution microscopy reveals tetherin HIV-1 interaction. PLoS Pathog 7:e1002456
Manley S, Gillette JM, Patterson GH, Shroff H, Hess HF, Betzig E, Lippincott-Schwartz J (2008) High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat Methods 5:155–157
Mayhew TM, Griffiths G, Lucocq JM (2004) Applications of an efficient method for comparing immunogold labelling patterns in the same sets of compartments in different groups of cells. Histochem Cell Biol 122:171–177
Mortensen KI, Churchman LS, Spudich JA, Flyvbjerg H (2010) Optimized localization analysis for single-molecule tracking and super-resolution microscopy. Nat Methods 7:377–381
Muller B, Daecke J, Fackler OT, Dittmar MT, Zentgraf H, Krausslich HG (2004) Construction and characterization of a fluorescently labeled infectious human immunodeficiency virus type 1 derivative. J Virol 78:10803–10813
Ono A (2009) HIV-1 assembly at the plasma membrane: Gag trafficking and localization. Future Virol 4:241–257
Owen DM, Rentero C, Rossy J, Magenau A, Williamson D, Rodriguez M, Gaus K (2010) PALM imaging and cluster analysis of protein heterogeneity at the cell surface. J Biophotonics 3:446–454
Ripley BD (1977) Modelling spatial patterns. J R Stat Soc 39:172–212
Ripley BD (1979) Tests of ‘randomness’ for spatial point patterns. J R Stat Soc 41:368–374
Rust MJ, Bates M, Zhuang X (2006) Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat Methods 3:793–795
Schermelleh L, Heintzmann R, Leonhardt H (2010) A guide to super-resolution fluorescence microscopy. J Cell Biol 190:165–175
van de Linde S, Sauer M, Heilemann M (2008) Subdiffraction-resolution fluorescence imaging of proteins in the mitochondrial inner membrane with photoswitchable fluorophores. J Struct Biol 164:250–254
van de Linde S, Loschberger A, Klein T, Heidbreder M, Wolter S, Heilemann M, Sauer M (2011) Direct stochastic optical reconstruction microscopy with standard fluorescent probes. Nat Protoc 6:991–1009
Williamson DJ, Owen DM, Rossy J, Magenau A, Wehrmann M, Gooding JJ, Gaus K (2011) Pre-existing clusters of the adaptor Lat do not participate in early T cell signaling events. Nat Immunol 12:655–662
Wolter S, Schuttpelz M, Tscherepanow M, Van De Linde S, Heilemann M, Sauer M (2010) Real-time computation of subdiffraction-resolution fluorescence images. J Microsc 237:12–22
Wombacher R, Heidbreder M, van de Linde S, Sheetz MP, Heilemann M, Cornish VW, Sauer M (2010) Live-cell super-resolution imaging with trimethoprim conjugates. Nat Methods 7:717–719
Wright ER, Schooler JB, Ding HJ, Kieffer C, Fillmore C, Sundquist WI, Jensen GJ (2007) Electron cryotomography of immature HIV-1 virions reveals the structure of the CA and SP1 Gag shells. EMBO J 26:2218–2226
Xu K, Babcock HP, Zhuang X (2012) Dual-objective STORM reveals three-dimensional filament organization in the actin cytoskeleton. Nat Methods 9:185–188
Acknowledgments
This work was supported by the Systems Biology Initiative (FORSYS) of the German Ministry of Research and Education (BMBF), Project VIROQUANT and Grant No. 0315262.
Author information
Authors and Affiliations
Corresponding author
Additional information
S. Malkusch and W. Muranyi contributed equally.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Malkusch, S., Muranyi, W., Müller, B. et al. Single-molecule coordinate-based analysis of the morphology of HIV-1 assembly sites with near-molecular spatial resolution. Histochem Cell Biol 139, 173–179 (2013). https://doi.org/10.1007/s00418-012-1014-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-012-1014-4