Abstract
Fluorescence microscopy can be used to assess quantitatively the interaction between a ligand and its receptor, between two macromolecules, or between a macromolecule and a particular intracellular compartment by co-localization analysis. In general, this analysis involves tagging potential interacting partners with distinct fluorophores—by direct labeling of a small ligand, by expression of fluorescent cDNA constructs, by immunofluorescence labeling, or by some combination of these methods. Pairwise comparison of the fluorescence intensity of the two fluorophores at each pixel in a two channel digital image of the sample reveals regions where both are present. With appropriate protocols, the image data can be interpreted to indicate where the potential interacting partners are co-localized. Keeping in mind the limited resolution of the light microscope, co-localization is often used to support the claim that two molecules are interacting.
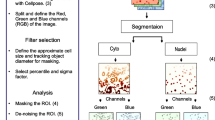
All quantitative methods for evaluating co-localization begin with identifying the pixels where the intensities of both color channels are above background. Typically this involves two sequential image segmentation steps: the first to exclude pixels where neither channel is above background, and the second to set overlap thresholds that exclude pixels where only one color channel is present. Following segmentation, various quantitative measures can be computed to describe the remaining subset of pixels where the two color channels overlap. These metrics range from simple calculation of the fraction of pixels where overlap occurs to more sophisticated image correlation metrics. Additional constraints may be employed to distinguish true co-localization from random overlap. Finally, an image map showing only the co-localized pixels may be displayed as an additional image channel in order to visualize the spatial distribution of co-localized pixels. Several commercial and open source software solutions provide this type of co-localization analysis, making image segmentation and calculation of metrics relatively straightforward. As an example, we provide a protocol for the time-dependent co-localization of fluorescently tagged lipoproteins with LDL receptor (LDLR) and with the early endosome marker EEA1.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Manders EM, Stap J, Brakenhoff GJ, van Driel R, Aten JA (1992) Dynamics of three-dimensional replication patterns during the S-phase, analysed by double labelling of DNA and confocal microscopy. J Cell Sci 103(Pt 3):857–862
Manders EMM, Verbeek FJ, Aten JA (1993) Measurement of co-localization of objects in dual-colour confocal images. J Microsc 169:375–382
Costes SV, Daelemans D, Cho EH, Dobbin Z, Pavlakis G, Lockett S (2004) Automatic and quantitative measurement of protein–protein colocalization in live cells. Biophys J 86:3993–4003
van Steensel B, van Binnendijk EP, Hornsby CD, van der Voort HT, Krozowski ZS, de Kloet ER, van Driel R (1996) Partial colocalization of glucocorticoid and mineralocorticoid receptors in discrete compartments in nuclei of rat hippocampus neurons. J Cell Sci 109(Pt 4):787–792
Li Q, Lau A, Morris TJ, Guo L, Fordyce CB, Stanley EF (2004) A syntaxin 1, Galpha(o), and N-type calcium channel complex at a presynaptic nerve terminal: analysis by quantitative immunocolocalization. J Neurosci 24:4070–4081
Bolte S, Cordelieres FP (2006) A guided tour into subcellular colocalization analysis in light microscopy. J Microsc 224:213–232
Comeau JW, Costantino S, Wiseman PW (2006) A guide to accurate fluorescence microscopy colocalization measurements. Biophys J 91:4611–4622
Adler J, Pagakis SN, Parmryd I (2008) Replicate-based noise corrected correlation for accurate measurements of colocalization. J Microsc 230:121–133
RamÍRez O, GarcÍA A, Rojas R, Couve A, HÄRtel S (2010) Confined displacement algorithm determines true and random colocalization in fluorescence microscopy. J Microsc 239:173–183.
Toomre D, Bewersdorf J (2010) A new wave of cellular imaging. Annu Rev Cell Dev Biol 26:285–314.
Zhao Z, Michaely P (2009) The role of calcium in lipoprotein release by the low-density lipoprotein receptor. Biochemistry 48:7313–7324
Image documentation for JaCoP can be found here: http://imagejdocu.tudor.lu/lib/exe/fetch.php?media=plugin:analysis:jacop_2.0:just_another_colocalization_plugin:jacop_ijconf2008.pdf Documentation and a link to the source code for Coloc 2, Colocalization Threshold and Colocalization Test can be found here: http://fiji.sc/wiki/Colocalization_Analysis
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this protocol
Cite this protocol
Pompey, S.N., Michaely, P., Luby-Phelps, K. (2013). Quantitative Fluorescence Co-localization to Study Protein–Receptor Complexes. In: Williams, M., Daviter, T. (eds) Protein-Ligand Interactions. Methods in Molecular Biology, vol 1008. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-398-5_16
Download citation
DOI: https://doi.org/10.1007/978-1-62703-398-5_16
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-397-8
Online ISBN: 978-1-62703-398-5
eBook Packages: Springer Protocols