Abstract
The human pregnane X receptor (PXR) is a ligand dependent transcription factor that can be activated by structurally diverse agonists including steroid hormones, bile acids, herbal drugs, and prescription medications. PXR regulates the transcription of several genes involved in xenobiotic detoxification and apoptosis. Activation of PXR has the potential to initiate adverse effects by altering drug pharmacokinetics or perturbing physiological processes. Hence, more reliable prediction of PXR activators would be valuable for pharmaceutical drug discovery to avoid potential toxic effects. Ligand- and protein structure-based computational models for PXR activation have been developed in several studies. There has been limited success with structure-based modeling approaches to predict human PXR activators, which can be attributed to the large and promiscuous site of this protein. Slightly better success has been achieved with ligand-based modeling methods including quantitative structure–activity relationship (QSAR) analysis, pharmacophore modeling and machine learning that use appropriate descriptors to account for the diversity of the ligand classes that bind to PXR. These combined computational approaches using molecular shape information may assist scientists to more confidently identify PXR activators. This chapter reviews the various ligand and structure based methods undertaken to date and their results.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Ruegg J, Penttinen-Damdimopoulou P, Makela S, Pongratz I, Gustafsson JA (2009) Receptors mediating toxicity and their involvement in endocrine disruption. EXS 99:289–323
Tabb MM, Blumberg B (2006) New modes of action for endocrine-disrupting chemicals. Mol Endocrinol 20:475–482
Safe S, Bandiera S, Sawyer T, Robertson L, Safe L, Parkinson A, Thomas PE, Ryan DE, Reik LM, Levin W et al (1985) PCBs: structure-function relationships and mechanism of action. Environ Health Perspect 60:47–56
Gustafsson JA (1995) Receptor-mediated toxicity. Toxicol Lett 82–83:465–470
Sewall CH, Lucier GW (1995) Receptor-mediated events and the evaluation of the Environmental Protection Agency (EPA) of dioxin risks. Mutat Res 333:111–122
Zollner G, Wagner M, Trauner M (2010) Nuclear receptors as drug targets in cholestasis and drug-induced hepatotoxicity. Pharmacol Ther 126:228–243
Lee EJ, Lean CB, Limenta LM (2009) Role of membrane transporters in the safety profile of drugs. Expert Opin Drug Metab Toxicol 5:1369–1383
Giguere V (1999) Orphan nuclear receptors: from gene to function. Endocr Rev 20:689–725
Ashby J, Houthoff E, Kennedy SJ, Stevens J, Bars R, Jekat FW, Campbell P, Van Miller J, Carpanini FM, Randall GL (1997) The challenge posed by endocrine-disrupting chemicals. Environ Health Perspect 105:164–169
Kelce WR, Gray LE, Wilson EM (1998) Antiandrogens as environmental endocrine disruptors. Reprod Fertil Dev 10:105–111
Melnick R, Lucier G, Wolfe M, Hall R, Stancel G, Prins G, Gallo M, Reuhl K, Ho SM, Brown T, Moore J, Leakey J, Haseman J, Kohn M (2002) Summary of the National Toxicology Program’s report of the endocrine disruptors low-dose peer review. Environ Health Perspect 110:427–431
Goldman JM, Laws SC, Balchak SK, Cooper RL, Kavlock RJ (2000) Endocrine-disrupting chemicals: prepubertal exposures and effects on sexual maturation and thyroid activity in the female rat. A focus on the EDSTAC recommendations. Crit Rev Toxicol 30:135–196
Ekins S, Chang C, Mani S, Krasowski MD, Reschly EJ, Iyer M, Kholodovych V, Ai N, Welsh WJ, Sinz M, Swaan PW, Patel R, Bachmann K (2007) Human pregnane X receptor antagonists and agonists define molecular requirements for different binding sites. Mol Pharmacol 72:592–603
Ekins S, Kholodovych V, Ai N, Sinz M, Gal J, Gera L, Welsh WJ, Bachmann K, Mani S (2008) Computational discovery of novel low micromolar human pregnane X receptor antagonists. Mol Pharmacol 74:662–672
Biswas A, Mani S, Redinbo MR, Krasowski MD, Li H, Ekins S (2009) Elucidating the ‘Jekyll and Hyde’ nature of PXR: the case for discovering antagonists or allosteric antagonists. Pharm Res 26:1807–1815
Mnif W, Pascussi JM, Pillion A, Escande A, Bartegi A, Nicolas JC, Cavailles V, Duchesne MJ, Balaguer P (2007) Estrogens and antiestrogens activate PXR. Toxicol Lett 170:19–29
Teotico DG, Bischof JJ, Peng L, Kliewer SA, Redinbo MR (2008) Structural basis of human pregnane X receptor activation by the hops constituent colupulone. Mol Pharmacol 74:1512–1520
Wada T, Gao J, Xie W (2009) PXR and CAR in energy metabolism. Trends Endocrinol Metab 20:273–279
Gong H, Guo P, Zhai Y, Zhou J, Uppal H, Jarzynka MJ, Song WC, Cheng SY, Xie W (2007) Estrogen deprivation and inhibition of breast cancer growth in vivo through activation of the orphan nuclear receptor liver X receptor. Mol Endocrinol 21:1781–1790
Rotroff DM, Beam AL, Dix DJ, Farmer A, Freeman KM, Houck KA, Judson RS, LeCluyse EL, Martin MT, Reif DM, Ferguson SS (2010) Xenobiotic-metabolizing enzyme and transporter gene expression in primary cultures of human hepatocytes modulated by ToxCast chemicals. J Toxicol Environ Health B Crit Rev 13:329–346
Tirona RG, Leake BF, Podust LM, Kim RB (2004) Identification of amino acids in rat pregnane X receptor that determine species-specific activation. Mol Pharmacol 65:36–44
Kortagere S, Krasowski MD, Reschly EJ, Venkatesh M, Mani S, Ekins S (2010) Evaluation of computational docking to identify pregnane X receptor agonists in the ToxCast database. Environ Health Perspect 118:1412–1417
Ekins S, Kortagere S, Iyer M, Reschly EJ, Lill MA, Redinbo M, Krasowski MD (2009) Challenges Predicting Ligand-Receptor Interactions of Promiscuous Proteins: The Nuclear Receptor PXR. PLoS Comput Biol 5:e1000594
Yasuda K, Ranade A, Venkataramanan R, Strom S, Chupka J, Ekins S, Schuetz E, Bachmann K (2008) A comprehensive in vitro and in silico analysis of antibiotics that activate pregnane X receptor and induce CYP3A4 in liver and intestine. Drug Metab Dispos 36:1689–1697
Khandelwal A, Krasowski MD, Reschly EJ, Sinz MW, Swaan PW, Ekins S (2008) Machine learning methods and docking for predicting human pregnane X receptor activation. Chem Res Toxicol 21:1457–1467
Ekins S, Andreyev S, Ryabov A, Kirillov E, Rakhmatulin EA, Sorokina S, Bugrim A, Nikolskaya T (2006) A Combined approach to drug metabolism and toxicity assessment. Drug Metab Dispos 34:495–503
Bachmann K, Patel H, Batayneh Z, Slama J, White D, Posey J, Ekins S, Gold D, Sambucetti L (2004) PXR and the regulation of apoA1 and HDL-cholesterol in rodents. Pharmacol Res 50:237–246
Ekins S, Erickson JA (2002) A pharmacophore for human pregnane-X-receptor ligands. Drug Metab Dispos 30:96–99
Pan Y, Li L, Kim G, Ekins S, Wang H, Swaan PW (2011) Identification and validation of Novel hPXR activators amongst prescribed drugs via ligand-based virtual screening. Drug Metab Dispos 39(2):337–44
Gao YD, Olson SH, Balkovec JM, Zhu Y, Royo I, Yabut J, Evers R, Tan EY, Tang W, Hartley DP, Mosley RT (2007) Attenuating pregnane X receptor (PXR) activation: a molecular modelling approach. Xenobiotica 37:124–138
Lemaire G, Benod C, Nahoum V, Pillon A, Boussioux AM, Guichou JF, Subra G, Pascussi JM, Bourguet W, Chavanieu A, Balaguer P (2007) Discovery of a highly active ligand of human pregnane x receptor: a case study from pharmacophore modeling and virtual screening to “in vivo” biological activity. Mol Pharmacol 72:572–581
Schuster D, Langer T (2005) The identification of ligand features essential for PXR activation by pharmacophore modeling. J Chem Inf Model 45:431–439
Huang H, Wang H, Sinz M, Zoeckler M, Staudinger J, Redinbo MR, Teotico DG, Locker J, Kalpana GV, Mani S (2007) Inhibition of drug metabolism by blocking the activation of nuclear receptors by ketoconazole. Oncogene 26:258–268
Lill MA, Dobler M, Vedani A (2005) In silico prediction of receptor-mediated environmental toxic phenomena-application to endocrine disruption. SAR QSAR Environ Res 16:149–169
Lin YS, Yasuda K, Assem M, Cline C, Barber J, Li CW, Kholodovych V, Ai N, Chen JD, Welsh WJ, Ekins S, Schuetz EG (2009) The major human pregnane X receptor (PXR) splice variant, PXR.2, exhibits significantly diminished ligand-activated transcriptional regulation. Drug Metab Dispos 37:1295–1304
Watkins RE, Davis-Searles PR, Lambert MH, Redinbo MR (2003) Coactivator binding promotes the specific interaction between ligand and the pregnane X receptor. J Mol Biol 331:815–828
Xue Y, Chao E, Zuercher WJ, Willson TM, Collins JL, Redinbo MR (2007) Crystal structure of the PXR-T1317 complex provides a scaffold to examine the potential for receptor antagonism. Bioorg Med Chem 15:2156–2166
Chrencik JE, Orans J, Moore LB, Xue Y, Peng L, Collins JL, Wisely GB, Lambert MH, Kliewer SA, Redinbo MR (2005) Structural disorder in the complex of human pregnane X receptor and the macrolide antibiotic rifampicin. Mol Endocrinol 19:1125–1134
Schuster D, Laggner C, Steindl TM, Palusczak A, Hartmann RW, Langer T (2006) Pharmacophore modeling and in silico screening for new P450 19 (aromatase) inhibitors. J Chem Inf Model 46:1301–1311
Khandelwal, A., Krasowski, M. D., Reschly, E. J., Sinz, M., Swaan, P. W., and Ekins, S. (2007) A Comparative Analysis of Quantitative Structure Activity Relationship Methods and Docking For Human Pregnane X Receptor Activation, submitted.
Jacobs MN (2004) In silico tools to aid risk assessment of endocrine disrupting chemicals. Toxicology 205:43–53
Hassan M, Brown RD, Varma-O'brien S, Rogers D (2006) Cheminformatics analysis and learning in a data pipelining environment. Mol Divers 10:283–299
Ghose AK, Viswanadhan VN, Wendoloski JJ (1998) Prediction of hydrophobic (lipophilic) properties of small organic molecules using fragmental methods: an analysis of ALOGP and CLOGP methods. J Phys Chem 102:3762–3772
Watkins RE, Wisely GB, Moore LB, Collins JL, Lambert MH, Williams SP, Willson TM, Kliewer SA, Redinbo MR (2001) The human nuclear xenobiotic receptor PXR: structural determinants of directed promiscuity. Science 292:2329–2333
Kortagere S, Chekmarev D, Welsh WJ, Ekins S (2009) Hybrid scoring and classification approaches to predict human pregnane X receptor activators. Pharm Res 26:1001–1011
Ekins S, Kortagere S, Iyer M, Reschly EJ, Lill MA, Redinbo MR, Krasowski MD (2009) Challenges predicting ligand-receptor interactions of promiscuous proteins: the nuclear receptor PXR. PLoS Comput Biol 5:e1000594
Judson RS, Houck KA, Kavlock RJ, Knudsen TB, Martin MT, Mortensen HM, Reif DM, Rotroff DM, Shah I, Richard AM, Dix DJ (2010) In vitro screening of environmental chemicals for targeted testing prioritization: the ToxCast project. Environ Health Perspect 118:485–492
Martin MT, Dix DJ, Judson RS, Kavlock RJ, Reif DM, Richard AM, Rotroff DM, Romanov S, Medvedev A, Poltoratskaya N, Gambarian M, Moeser M, Makarov SS, Houck KA (2010) Impact of environmental chemicals on key transcription regulators and correlation to toxicity end points within EPA’s ToxCast program. Chem Res Toxicol 23:578–590
Knudsen TB, Martin MT, Kavlock RJ, Judson RS, Dix DJ, Singh AV (2009) Profiling the activity of environmental chemicals in prenatal developmental toxicity studies using the U.S. EPA’s ToxRefDB. Reprod Toxicol 28:209–219
Mani S, Huang H, Sundarababu S, Liu W, Kalpana G, Smith AB, Horwitz SB (2005) Activation of the steroid and xenobiotic receptor (human pregnane X receptor) by nontaxane microtubule-stabilizing agents. Clin Cancer Res 11:6359–6369
Chen Y, Tang Y, Wang MT, Zeng S, Nie D (2007) Human pregnane X receptor and resistance to chemotherapy in prostate cancer. Cancer Res 67:10361–10367
Reynolds CH, Tounge BA, Bembenek SD (2008) Ligand binding efficiency: trends, physical basis, and implications. J Med Chem 51:2432–2438
Ekins S, Reschly EJ, Hagey LR, Krasowski MD (2008) Evolution of pharmacologic specificity in the pregnane X receptor. BMC Evol Biol 8:103
Krasowski MD, Yasuda K, Hagey LR, Schuetz EG (2005) Evolution of the pregnane x receptor: adaptation to cross-species differences in biliary bile salts. Mol Endocrinol 19:1720–1739
Gasteiger JAMM (1980) Iterative partial equalization of orbital electronegativity—a rapid access to atomic charges. Tetrahedron 36:10
Kortagere S, Welsh WJ (2006) Development and application of hybrid structure based method for efficient screening of ligands binding to G-protein coupled receptors. J Comput Aided Mol Des 20:789–802
Xue Y, Moore LB, Orans J, Peng L, Bencharit S, Kliewer SA, Redinbo MR (2007) Crystal structure of the pregnane X receptor-estradiol complex provides insights into endobiotic recognition. Mol Endocrinol 21:1028–1038
Jones G, Willett P, Glen RC, Leach AR, Taylor R (1997) Development and validation of a genetic algorithm for flexible docking. J Mol Biol 267:727–748
Kogej T, Engkvist O, Blomberg N, Muresan S (2006) Multifingerprint based similarity searches for targeted class compound selection. J Chem Inf Model 46:1201–1213
Acknowledgments
We would like to thank all our collaborators who have contributed to our studies on PXR. SK is supported by Scientist Development Grant awarded by American Heart Association.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Appendix
Appendix
Structure-based Methods. Data collection: A comprehensive hPXR dataset consisting of a variety of chemical classes, namely, androstanes, pregnanes, estratrienes, bile acids, and regular xenobiotics is available from previously published studies. (53, 54). The Environmental Protection Agency has developed a dataset comprising of everyday chemicals, pesticides and other related chemicals aptly called Toxcast dataset (ToxcastTM) which is available for use by researchers ((47, 49). For all these datasets, either the direct binding data to hPXR or relative fold induction changes have been published. However, the EC50 cutoff value used for classifying the molecules into hPXR activators and nonactivators is not always available or does not follow a strict pattern (25, 33), SMILES strings for compounds listed in datasets or 2D structures in mol or sdf formats can be obtained from the respective publications (25). Using these as input, single low energy three-dimensional structure of molecules can be generated using Molecular Operating Environment (MOE, Chemical Computing Group, Montreal, Canada) or CORINA (Molecular Networks GmbH, Nägelsbachstr. 25, 91052 Erlangen, Germany. http://www.mol-net.de) with partial charges assigned according to the Gasteiger-Marsili scheme (55).
Molecular Descriptors. hPXR binds a variety of ligands that differ in shape, size, and chemical composition. However, analysis of well known hPXR agonists show that the interactions are dominated by hydrophobic and hydrogen bonding features of the ligands. Hence to capture these properties, molecular descriptors that represent shape, size, flexibility and hydrogen bonding, and hydrophobic properties must be chosen. These include FCFP_6 fingerprints, volume, weight, KierA1-A3, Kier1-3, number of rotatable bonds, number of rings and KierFlex, electrostatic features like logP, TPSA, logs of lip_don, lip_acc, number of N atoms, and number of O atoms. The values for these specific molecular descriptors can be derived from MOE. In addition to analyze the specific role of shape and electrostatics, specific shape based descriptors such as shape signatures can be derived (56).
Molecular Docking. Five crystal structures of hPXR cocrystalized with a variety of ligands are available in the protein databank (PDB) under the codes 1 M13, resolution 2.00 Å (36), 1SKX, resolution 2.80 Å (38), 2O9I, resolution 2.80 Å (37), 1NRL, resolution 2.00 Å (36), and 2QNV, resolution 2.80 Å (37). In addition, another structure in complex with 17-β estradiol that is yet to be deposited in PDB was also used for docking studies (57). In all cases, the protein structure is first prepared by removing the crystal structure ligand and adding hydrogen atoms to the amino acids and the resulting structures are energy minimized to remove any steric contacts. All amino acids within 6 Å of the cocrystallized ligand are generally chosen as being part of the binding site. The docking program GOLD (ver 4) (58) or FlexX could be used for docking. GOLD uses a genetic algorithm to explore the various conformations of ligands and flexible receptor side chains in the binding pocket. In our studies, we have chosen to perform 20 independent docking runs for each ligand to sample the ligand and protein conformational space. The resulting docked complexes can be scored using GoldScore and ChemScore.
Scoring functions. One of the bottlenecks in docking studies is the scoring functions. Although most docking programs are capable of sampling the ligand in the binding pocket and generating solutions, the scoring functions are not sensitive enough to identify the truly best docked solutions. This is because, most scoring functions are empirical energy based schemes as shown below:
Where ΔGbind is the binding free energy, ΔGsolvent is the penalty for desolvation, ΔGint is the internal energy, ΔGrot represents the energy contribution for bond rotation, and ΔGvib represents the energy contribution for vibration component. However, the terms for solvation is approximated to a best fit model and the terms for entropy are most likely dropped from the energy equation due to the complexity involved in computing the entropy factors. Thus, in practice no single scoring function can work for every target and has to be customized to suit the needs of the target. In case of hPXR, given that the binding site is very promiscuous and binds a variety of ligands, developing a scoring scheme is challenging. We have used a number of methods to derive consensus scoring schemes including contact score, shape based scores, and molecular descriptor based schemes.
-
1.
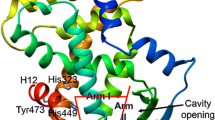
Contact scoring scheme. The docked receptor–ligand complexes were scored using a contact based scoring function. Accordingly, an in-house program was used to scan the docked complexes for contacts between the ligand and protein atoms (56). These contacts were scored based on a weighting scheme that was derived from the nature of interaction between the ligands cocrystallized with hhPXR. For example, hyperforin has hydrogen bond interactions with residues Gln285, His407, and Ser247 of the hhPXR protein in the crystal structure (PDB ID:1 M13) (Fig. 3). Thus the contact scoring function weighted all those docked protein–ligand complexes that featured the hydrogen bonding between the ligands and these three residues, higher than the rest of the interactions. Similarly, other nonbonded interactions were weighted based on the interactions of the ligands in the hPXR crystal structures. All interaction scores were then summed and normalized against all crystal structures. A consensus scoring scheme was developed for final classification based on the following rule: Only those compounds that had at least half the value of the highest GoldScore and a nonzero contact score were assigned as activators and the rest of the molecules were classified as nonactivators.
-
2.
Shape based scoring scheme. In this scheme, the ligands were compared with the hPXR ligands from the five crystal structures for their shape based similarities using two different approaches. The first was based on the 2D similarity encoded in MDL public fingerprint keys calculation using Discovery Studio 2.0 (Accelrys, San Diego, CA). The Tanimoto coefficient was used as the metric to compare the molecular fingerprints. The coefficients varied between 0 and 1, where 0 meant maximally dissimilar and 1 coded for maximally similar. The Tanimoto coefficient between fingerprints X and Y has been defined to be: [number of features in intersect (A,B)]/[number of features in union(A,B)], where A and B are two compounds.
In an approach, the 3D shapes of the molecules from the combined dataset were compared with the shapes of each of the four crystal structure ligands. This was achieved by comparing their corresponding 1D Shape Signatures and a dissimilarity score was computed for each ligand pair. The dissimilarity score was then converted to a similarity score, which was in turn used as weighting factor for the GoldScore. In all these scoring schemes the consensus score was calculated as shown below in Eq. (1).
-
3.
Molecular descriptor based scoring. In this scheme, the molecular descriptors computed using MOE were used to calculate Euclidean distances from the crystal structure ligands. These Euclidean distances were used as weighting factors to GoldScore. Similarly, the values of the molecular descriptors were also used to calculate Tanimoto similarity indices (59) with reference to the cocrystal structure ligands. The values for the Tanimoto indices for each ligand in the combined set were calculated against each of the crystal structure ligands and then used as weighting factors to the GoldScores.
The weighted docking score of an active compound j with i conformations was described as
$$ {S_{{i,j}}} = {w_i}{s_{{ij}}} $$(1)where s ij was the original GoldScore for the compound i in its jth conformation and w i is the weighting factor for compound i from either of the schemes described above.
Rights and permissions
Copyright information
© 2012 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Kortagere, S., Krasowski, M.D., Ekins, S. (2012). Ligand- and Structure-Based Pregnane X Receptor Models. In: Reisfeld, B., Mayeno, A. (eds) Computational Toxicology. Methods in Molecular Biology, vol 929. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-050-2_15
Download citation
DOI: https://doi.org/10.1007/978-1-62703-050-2_15
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-049-6
Online ISBN: 978-1-62703-050-2
eBook Packages: Springer Protocols