Abstract
Codon substitution models (CSMs) are commonly used to infer the history of natural section for a set of protein-coding sequences, often with the explicit goal of detecting the signature of positive Darwinian selection. However, the validity and success of CSMs used in conjunction with the maximum likelihood (ML) framework is sometimes challenged with claims that the approach might too often support false conclusions. In this chapter, we use a case study approach to identify four legitimate statistical difficulties associated with inference of evolutionary events using CSMs. These include: (1) model misspecification, (2) low information content, (3) the confounding of processes, and (4) phenomenological load, or PL. While past criticisms of CSMs can be connected to these issues, the historical critiques were often misdirected, or overstated, because they failed to recognize that the success of any model-based approach depends on the relationship between model and data. Here, we explore this relationship and provide a candid assessment of the limitations of CSMs to extract historical information from extant sequences. To aid in this assessment, we provide a brief overview of: (1) a more realistic way of thinking about the process of codon evolution framed in terms of population genetic parameters, and (2) a novel presentation of the ML statistical framework. We then divide the development of CSMs into two broad phases of scientific activity and show that the latter phase is characterized by increases in model complexity that can sometimes negatively impact inference of evolutionary mechanisms. Such problems are not yet widely appreciated by the users of CSMs. These problems can be avoided by using a model that is appropriate for the data; but, understanding the relationship between the data and a fitted model is a difficult task. We argue that the only way to properly understand that relationship is to perform in silico experiments using a generating process that can mimic the data as closely as possible. The mutation-selection modeling framework (MutSel) is presented as the basis of such a generating process. We contend that if complex CSMs continue to be developed for testing explicit mechanistic hypotheses, then additional analyses such as those described in here (e.g., penalized LRTs and estimation of PL) will need to be applied alongside the more traditional inferential methods.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
- Codon substitution model
- dN/dS
- False positives
- Maximum likelihood
- Mechanistic model
- Model misspecification
- Mutation-selection model
- Parameter confounding
- Phenomenological load
- Phenomenological model
- Positive selection
- Reliability
- Statistical inference
- Site-specific fitness landscape
1 Introduction
Codon substitution models (CSMs) fitted to an alignment of homologous protein-coding genes are commonly used to make inferences about evolutionary processes at the molecular level (see Chapter 10 for examples of different applications of CSMs). Such processes (e.g., mutation and selection) are represented by a vector of parameters θ that can be estimated using maximum likelihood (ML) or Bayesian statistical methods. Here, we focus on ML and for convenience use CSM to indicate a model that is used in conjunction with the ML approach (see [21], for an example of the Bayesian approach). Considerable apprehension was expressed about the statistical validity of CSMs during their initial phase of development. In particular were concerns over the risk of falsely inferring that a sequence or codon site evolved by adaptive evolution [11, 22, 23, 46, 60,61,62,63, 85]. Many of the studies employed in the critique of CSMs were later shown to be flawed due to statistical errors or incorrect interpretation of results [70, 72, 77, 84]. In their reanalysis of the iconic MHC dataset [24], for example, Suzuki and Nei [61] based their criticism of the ML approach on results that were incorrect due to computational issues [70]. And in simulation studies by Suzuki [60] and Nozawa et al. [46], the branch-site model of Yang and Nielsen [79] was criticized as being too liberal because it falsely inferred positive selection at 32 out of 14,000 simulated sites, despite that this rate (0.0023) was well below the level of significance of the test (α = 0.05) [77]. Concerns about the ML approach were eventually mollified by numerous simulation studies showing that the false positive rate is no greater than the specified level of significance of the LRT under a wide range of evolutionary scenarios [2, 3, 29, 31, 37, 70, 77, 82, 85, 86]. The validity and success of the approach is now well established [84], and this has led to the formulation of CSMs of ever-increasing sophistication [31, 41, 48,49,50, 55, 64, 65].
The most common use of a CSM is to infer whether a given process, such as adaptive evolution somewhere in the gene, the fixation of double and triple mutations, or variations in the synonymous substitution rate, actually occurred when the alignment was generated. Several factors can potentially undermine the reliability of such inferences. These include:
-
1.
Model misspecification, which can result in biased parameter estimates;
-
2.
Low information content, which can cause parameter estimates to have large sampling errors and can lead to excessive false positive rates;
-
3.
Confounding, which can cause patterns in the data generated by one evolutionary process to be attributed to a different process;
-
4.
Phenomenological load, which can cause a model parameter to be statistically significant even if the process it represents did not actually occur when the data was generated.
These same factors can impact any model-based effort to make inferences from data generated by complex biological processes, not only to the CSMs described here. The possibility of false inference due to any combination of these factors does not imply that the CSM approach is unreliable in principle. As has been demonstrated by numerous successful applications, CSMs generally extract accurate and useful information provided that the model is well suited for the data at hand [1, 71, 76]. We maintain that the validity of inferences is not a function of the model in and of itself, but is a consequence of the relationship between the model and the data.
Here, we explore this relationship via case studies taken from the historical development of CSMs. Our objective is to be candid about the limitations of CSMs to reliably extract information from an alignment. But, we emphasize that the impact of these limitations (i.e., false positives and confounding) is a consequence of a mismatch between the parameters included in the model and the often limited information contained in the alignment. The case studies are divided into two parts, each corresponding to a distinct phase in the development of CSMs. Phase I is characterized by pioneering efforts to formulate CSMs to account for the most prominent components of variation in an alignment [16, 42]. These include the M-series models that were among the first CSMs to account for variations in selection effects across sites [81], and the branch-site model of Yang and Nielsen [79] (hereafter, YN-BSM) formulated to account for variations in selection effects across both sites and branches. The first pair of case studies exemplifies concerns about the impact of low information content (Case Study A) and model misspecification (Case Study B) on the probability of falsely detecting positive selection in a gene or at a particular codon site. We also include a description of methods recently developed to mitigate the problem of false inference.
Phase II in the historical development is characterized by the general increase in the complexity of CSMs aimed to account for more subtle components of variation in an alignment.Footnote 1 Models used to detect temporal changes in site-specific selection effects (e.g., [18, 31, 55]) or “heterotachy” [36] are representative. The movement toward complex parameter-rich models has resulted in a new set of concerns that are not yet widely appreciated. Principal among these is an increase in the possibility of confounding. Two components of the alignment-generating process are confounded if they can produce the same or similar patterns in the data. Such components can be impossible to disentangle without the input of further biological information, and their existence can lead to a statistical pathology that we call phenomenological load (PL). The second pair of case studies illustrates the possibility of false inference due to confounding (Case Study C) and PL (Case Study D). An essential feature of these studies is the use of a much more realistic generating model to produce alignments for the purpose of model evaluation.
Recent discoveries made using the mutation-selection (MutSel; [80]) framework of Halpern and Bruno [19], which is based on a realistic approximation of population dynamics at individual codon sites, have challenged the way we think about the relationship between parameters of traditional CSMs and components of the process of molecular evolution they are meant to summarize (e.g., [25, 26, 56, 57]). Previously, there has been a tendency to think about alignment-generating processes as if they occur in the same way they are modeled by a CSM. This way of thinking can be misleading because mechanisms of protein evolution can differ in important and substantial ways from traditional CSMs. To redress this issue, we begin this chapter with a brief overview of the conceptual foundations of MutSel as a more realistic way of thinking about the actual process of molecular evolution. This material is followed by a novel presentation of the ML statistical framework intended to illustrate potential limitations in what can reasonably be inferred when a CSM is fitted to data.
2 Conceptual Foundations
2.1 How Should We Think About the Alignment-Generating Process?
A codon substitution model represents an attempt to explain the way a target protein-coding gene changed over time by a combination of mutation, selection (purifying as well as adaptive), and drift. Adaptive evolution occurs at each site within a protein in response to a hierarchy of effects, including, but not limited to, changes in the network of the protein’s interactions, changes in the functional properties of that network, and changes in both the cellular and organismal environment over time. The result of the complex interplay between these effects is typically viewed through the narrow lens of an alignment of homologous sequences X obtained from extant species, possibly accompanied by a tree topology τ (for our purposes, it is always assumed that τ is known). The information contained in X is evidently insufficient to resolve all of the effects of the true generating process, which would in any case be difficult or even impossible to parameterize with any accuracy. It is therefore necessary to base the formulation of a CSM on a number of simplifying assumptions. The usual assumptions include that:
-
1.
Sites evolved independently;
-
2.
Each site evolved via a homogenous substitution process over the tree (formally, by a Markov process governed by a substitution rate matrix Q);
-
3.
The selection regime at a site is determined by Q j drawn from a small set of possible substitution rate matrices {Q 1, …, Q k};
-
4.
All sites share a common vector of stationary frequencies and evolved via a common mutation process.
The elements q ij of a substitution rate matrix Q are typically defined for codons i ≠ j as follows [44]:
where κ is the transition bias and π i is the stationary frequency of the ith codon, both assumed to be the same for all codon sites. The ratio ω = dN∕dS of the nonsynonymous substitution rate dN to the synonymous substitution rate dS (both adjusted for “opportunity”Footnote 2) quantifies the stringency of selection at the site, with values closer to zero corresponding to sites that are more strongly conserved. We follow standard notation and use \( \widehat{\omega} \) to represent the maximum likelihood estimate (MLE) of ω obtained by fitting Eq. 1 to an alignment.
Equation 1 provides the building block for most CSMs, yet it is unsuitable as a means to think about the substitution process at a site. For instance, the rate ratio in Eq. 1 is assumed to be the same for all nonsynonymous pairs of codons. If interpreted mechanistically, this is tantamount to the assumption that the amino acid occupying a site has fitness f and all other amino acids have fitness f + df, and that, with each substitution, the newly fixed amino acid changes its fitness to f and the previous occupant changes it fitness to f + df. Such a narrow view of the substitution process, akin to frequency-dependent selection [6, 25], is conceptually misleading for the majority of proteins. To be clear, CSMs are undoubtedly a valuable tool to make inferences about the evolution of a protein (e.g., [8, 52, 71, 76]); our point is that they do not necessarily provide the best way to think about the process.
The way we think about the substitution process should not be limited to unrealistic assumptions used to formulate a tractable CSM. It is more informative to conceptualize evolution at a codon site using the traditional metaphor of a fitness landscape upon which greater height represents greater fitness as depicted in Fig. 1. If sites are assumed to evolve independently, a site-specific fitness landscape can be defined for the hth site by a vector of fitness coefficients f h and its implied vector of equilibrium codon frequencies π h. Combined with a model for the mutation process, π h determines the evolutionary dynamics at the site, or the way it “moves” over its landscape (more formally, the way mutation and fixation events occur at a codon site in a population over time). This provides a way to think about evolution at a codon site in terms of three possible dynamic regimes: shifting balance, under which the site moves episodically away from the peak of its fitness landscape (i.e., the fittest amino acid) via drift and back again by positive selection (Fig. 1a); adaptive evolution, under which a change in the landscape is followed by movement of the site toward its new fitness peak (Fig. 1b); and neutral or nearly neutral evolution, under which drift dominates and the site is free to move over a relatively flat landscape limited primarily by biases in the mutation process. This way of thinking about the alignment-generating process is encapsulated by the MutSel framework [6, 7, 25]. The precise relationship between the MutSel framework and the three dynamic regimes will be presented in Case Study C.
It can be useful to think of the substitution process at a site as movement on a site-specific fitness landscape. The horizontal axis in each figure shows the amino acids at a hypothetical site in order of their stationary frequencies indicated by the height of the bars. Frequency is a function of mutation and selection, but can be construed as a proxy for fitness. The site-specific dN∕dS ratio [25] is a function of the amino acid that occupies the site, and can be <1 (left of the red dashed line) or >1 (right of the dashed red line). (a) Suppose phenylalanine (F, TTT) is the fittest amino acid. The site-specific dN∕dS ratio is much less than one when occupied by F because any nonsynonymous mutation will always be to an amino acid that is less fit. Nevertheless, it is possible for an amino acid such as valine (V, GTT) to be fixed on occasion, provided that selection is not too stringent. When this happens, dN∕dS at the site is temporarily elevated to a value greater than one as positive selection moves the site back to F by a series of replacement substitutions, e.g., V (GTT) → G (GGT) → C (TGT) → F (TTT). We call the episodic recurrence of this process shifting balance on a static fitness landscape. Shifting balance on a landscape for which all frequencies are approximately equal corresponds to nearly neutral evolution (not depicted), when dN∕dS is always ≈1. (b) Now, consider what happens following a change in one or more external factors that impact the functional significance of the site. The relative fitnesses of the amino acids might change from that depicted in a to that in b for instance, where glutamine (Q) is fittest. If at the time of the change the site is occupied by F (as is most likely), then dN∕dS would be temporarily elevated as positive selection moves the site toward its new peak at Q, e.g., F (TTT) → Y (TAT) → H (CAT) → Q (CAA). This process of adaptive evolution is followed by a return to shifting balance once the site is occupied by Q
2.2 What Is the Objective of Model Building?
CSMs have become increasingly complex with the addition of more free parameters since the introduction of the M-series models in Yang et al. [81]. The prima facie objective of this trend is to produce models that provide better mechanistic explanations of the data. The assumption is that this will lead to more accurate inferences about evolutionary processes, particularly as the volume of genetic data increases [35]. However, the significance of a new model parameter is assessed by a comparison of site-pattern distributions without reference to mechanism. Combined with the possibility of confounding, this feature of the ML framework means that the objective of improving model fit does not necessarily coincide with the objective of providing a better representation of the mechanisms of the true generating process.
Given any CSM with parameters θ M, it is possible to compute a vector P that assigns a probability to each of the 61N possible site patterns for an N-taxon alignment (i.e., a multinomial distribution for 61N categories). We refer to P = P M(θ M) as the site-pattern distribution for that model. Figure 2 depicts the space of all possible site-pattern distributions for an N-taxon alignment. Each ellipse represents the family of distributions {P M(θ M)∣θ M ∈ ΩM}, where ΩM is the vector space of all possible values of θ M. For example, {P M0(θ M0)∣θ M0 ∈ ΩM0} is the family of distributions that can be specified using M0, the simplest CSM that assumes a common substitution rate matrix Q for all sites and branches. This is nested inside {P M1(θ M1)∣θ M1 ∈ ΩM1}, where M1 is a hypothetical model that is the same as M0 but for a few extra parameters. Likewise, M1 is nested in M2. The location of the site-pattern distribution for the true generating process is represented by P PG. Its location is fixed but unknown. It is therefore not possible to assess the distance between it and any other distribution. Instead, comparisons are made using the site-pattern distribution inferred under the saturated model.
The (61N − 1)-dimensional simplex containing all possible site-pattern distributions for an N-taxon alignment is depicted. The innermost ellipse represents the subspace {P M0(θ M0)∣θ M0 ∈ ΩM0} that is the family of distributions that can be specified using M0, the simplest of CSMs. This is nested in the family of distributions that can be specified using M1 (blue ellipse), a hypothetical model that has the same parameters as M0 plus some extra parameters. Similarly, M1 is nested in M2 (red ellipse). Whereas models are represented by subspaces of distributions, the true generating process is represented by a single point P GP, the location of which is unknown. The empirical site-pattern distribution \( {P}_{\mathrm{S}}\left({\widehat{\theta}}_{\mathrm{S}}\right) \) corresponds to the saturated model fitted to the alignment; with large samples, \( {P}_{\mathrm{S}}\left({\widehat{\theta}}_{\mathrm{S}}\right)\approx {P}_{\mathrm{GP}} \). For any other model M, the member \( {P}_{\mathrm{M}}\left({\widehat{\theta}}_{\mathrm{M}}\right)\in \left\{{P}_{\mathrm{M}}\left({\theta}_{\mathrm{M}}\right)\mid {\theta}_{\mathrm{M}}\in {\Omega}_{\mathrm{M}}\right\} \) most consistent with X is the one that minimizes deviance, which is twice the difference between the maximum log-likelihood of the data under the saturated model and the maximum log-likelihood of the data under M
Whereas a CSM {P M(θ M)∣θ M ∈ ΩM} can be thought of as a family of multinomial distributions for the 61N possible site patterns, the fitted saturated model \( {P}_{\mathrm{S}}\left({\widehat{\theta}}_{\mathrm{S}}\right) \) is the unique distribution defined by the MLE \( {\widehat{\theta}}_{\mathrm{S}}={\left({y}_1/ n,\dots, {y}_m/ n\right)}^T \), where y i > 0 is the observed frequency of the ith site pattern, m is the number of unique site patterns, and n is the number of codon sites. In other words, the fitted saturated model is the empirical site-pattern distribution for a given alignment. Because it takes none of the mechanisms of mutation or selection into account, ignores the phylogenetic relationships between sequences, and excludes the possibility of site patterns that were not actually observed (i.e., y i∕n = 0 for site patterns i not observed in X), \( {P}_{\mathrm{S}}\left({\widehat{\theta}}_{\mathrm{S}}\right) \) can be construed as the maximally phenomenological explanation of the observed alignment. An alignment is always more likely under the saturated model than it is under any other CSM. \( {P}_{\mathrm{S}}\left({\widehat{\theta}}_{\mathrm{S}}\right) \) therefore provides a natural benchmark for model improvement.
For any alignment, the MLE over the family of distributions {P M(θ M)∣θ M ∈ ΩM} is represented by a fixed point \( {P}_{\mathrm{M}}\left({\widehat{\theta}}_{\mathrm{M}}\right) \) in Fig. 2. \( {P}_{\mathrm{M}}\left({\widehat{\theta}}_{\mathrm{M}}\right) \) is the distribution that minimizes the statistical deviance between P M(θ M) and \( {P}_{\mathrm{S}}\left({\widehat{\theta}}_{\mathrm{S}}\right) \). Deviance is defined as twice the difference between the maximum log-likelihood (LL) of the data under the saturated model and the maximum log-likelihood of the data under M:
A key feature of deviance is that it always decreases as more parameters are added to the model, corresponding to an increase in the probability of the data under that model. For example, suppose {P M2(θ M2)∣θ M2 ∈ ΩM2} is the same family of distributions as {P M1(θ M1)∣θ M1 ∈ ΩM1} but for the inclusion of one additional parameter ψ, so that θ M2 = (θ M1, ψ). The improvement in the probability of the data under \( {P}_{\mathrm{M}2}\left({\widehat{\theta}}_{\mathrm{M}2}\right) \) over its probability under \( {P}_{\mathrm{M}1}\left({\widehat{\theta}}_{\mathrm{M}1}\right) \) is assessed by the size of the reduction in deviance induced by ψ:
Equation 3 is just the familiar log-likelihood ratio (LLR) used to compare nested models under the maximum likelihood framework.
Given this measure of model improvement, the de facto objective of model building is not to provide a mechanistic explanation of the data that more accurately represents the true generating process, but only to move closer to the site-pattern distribution of the fitted saturated model. Real alignments are limited in size, so there will always be some distance between \( {P}_{\mathrm{S}}\left({\widehat{\theta}}_{\mathrm{S}}\right) \) and P GP due to sampling error (as represented in Fig. 2). But even with an infinite number of codon sites, when \( {P}_{\mathrm{S}}\left({\widehat{\theta}}_{\mathrm{S}}\right) \) converges to P GP, the criterion of minimizing deviance does not inevitably lead to a better explanation of the data because of the possibility of confounding. Two processes are said to be confounded if they can produce similar patterns in the data. Hence, if ψ represents a process E that did not actually occur when the data was generated, and if E is confounded with another process that did occur, the LLR in Eq. 3 can still be significant. Under this scenario, the addition of ψ to M1 would engender movement toward \( {P}_{\mathrm{S}}\left({\widehat{\theta}}_{\mathrm{S}}\right) \) and P GP, but the new model M2 would also provide a worse mechanistic explanation of the data because it would falsely indicate that E occurred. The possibility of confounding and its impact on inference is demonstrated in Case Study D.
3 Phase I: Pioneering CSMs
The first effort to detect positive selection at the molecular level [24] relied on heuristic counting methods [43]. Phase I of CSM development followed with the introduction of formal statistical approaches based on ML [16, 42]. The first CSMs were used to infer whether the estimate \( \widehat{\omega} \) of a single nonsynonymous to synonymous substitution rate ratio averaged over all sites and branches was significantly greater than one. Such CSMs were found to have low power due to the pervasiveness of synonymous substitutions at most sites within a typical gene [76]. An early attempt to increase the statistical power to infer positive selection was the CSM designed to detect \( \widehat{\omega}>1 \) on specific branches [78]. Models accounting for variations in ω across sites were subsequently developed, the most prominent of which are the M-series models [78, 81]. These were accompanied by methods to identify individual sites under positive selection. The quest for power culminated in the development of models that account for variations in the rate ratio across both sites and branches. The appearance of various branch-site models (e.g., [4, 10, 79, 86]) marks the end of Phase I of CSM development.
Two case studies are employed in this section to illustrate some of the inferential challenges associated with Phase I models. We use Case Study A to examine the impact of low information content on the inference of positive selection at individual codon sites. The subject of this study is the M1a vs M2a model contrast applied to the tax gene of the human T-cell lymphotropic virus type I (HTLV-I; [63, 82]). We use Case Study B to illustrate how model misspecification (i.e., differences between the fitted model and the generating process) can lead to false inferences. The subject of this study is the Yang–Nielsen branch-site model (YN-BSM; [79]) applied to simulated data.
3.1 Case Study A: Low Information Content
To study the impact of low information content on inference, we use a pair of nested M-series models known as M1a and M2a [70, 82]. Under M1a, sites are partitioned into two rate-ratio categories, 0 < ω 0 < 1 and ω 1 = 1 in proportions p 0 and p 1 = 1 − p 0. M2a includes an additional category for the proportion of sites p 2 = 1 − p 0 − p 2 that evolved under positive selection with ω 2 > 1. The use of multiple categories permits two levels of inference. The first is an omnibus likelihood ratio test (LRT) for evidence of positive selection somewhere in the gene, which is conducted by contrasting a pair of nested models. For example, the contrast of M1a vs M2a is made by computing the distance \( \mathrm{LLR}=\Delta D\left({\widehat{\theta}}_{\mathrm{M}1\mathrm{a}},{\widehat{\theta}}_{\mathrm{M}2\mathrm{a}}\right) \) between the two models and comparing the result to the limiting distribution of the LLR under the null model. In this case, the limiting distribution of LLR is often taken to be \( {\chi}_2^2 \) [75], which would be correct under regular likelihood theory because the models differ by two parameters. The second level of inference is used to identify individual sites that underwent positive selection. This is conducted only if positive selection is inferred by the omnibus test (e.g., if LLR > 5.99 for the M1a vs M2a contrast at the 5% level of significance). Let c 0, c 1, and c 2 represent the event that a given site pattern x falls into the stringent (\( 0<{\widehat{\omega}}_0<1 \)), neutral (\( {\widehat{\omega}}_1=1 \)), or positive (\( {\widehat{\omega}}_2>1 \)) selection category, respectively. Applying Bayes’ rule:
Sites with a sufficiently high posterior probability (e.g., \( \Pr \left({c}_2\mid x,{\widehat{\theta}}_{\mathrm{M}2\mathrm{a}}\right)>0.95 \)) are inferred to have undergone positive selection. Equation 4 is representative of the naive empirical Bayes (NEB) approach under which MLEs (\( {\widehat{\theta}}_{\mathrm{M}2\mathrm{a}} \)) are used to compute posterior probabilities.
The NEB approach ignores potential errors in parameter estimates that can lead to false inference of positive selection at a site (i.e., a false positive). The resulting false positive rate can be especially high for alignments with low information content. An example setting with low information content arises when there are a substantial number of invariant sites, since these provide little information about the substitution process. The issue of low information content is well illustrated by the extreme case of the tax gene, HTLV-I [63]. The alignment consists of 20 sequences with 181 codon sites, 158 of which are invariant. The 23 variable sites have only one substitution each: 2 are synonymous and 21 are nonsynonymous. The high ratio of nonsynonymous-to-synonymous substitutions suggests that the gene underwent positive selection. This hypothesis was supported by analytic results: the LLR for the M1a vs M2a contrast was 6.96 corresponding to a p-value of approximately 0.03 [82]. The omnibus test therefore supported the conclusion that the gene underwent positive selection. However, the MLE for p 2 under M2a was \( {\widehat{p}}_2=1 \). Using this value in Eq. 4 gives \( \Pr \left({c}_2\mid x,{\widehat{\theta}}_{\mathrm{M}2\mathrm{a}}\right)=1 \) for all sites, including the 158 invariable sites. Such an unreasonable result can occur under NEB because, despite the possibility of large sampling errors in MLEs due to low information, \( {\widehat{\theta}}_{\mathrm{M}2\mathrm{a}} \) is treated as a known value in Eq. 4.
Bayes empirical Bayes (BEB; [82]), a partial Bayesian approach under which rate ratios and their corresponding proportions are assigned discrete prior distributions (cf. [21]), was proposed as an alternative to NEB. Numerical integration over the assumed priors tends to provide better estimates of posterior probabilities, particularly in cases where information content is low. Using BEB in the analysis of the tax gene, for example, the posterior probability was \( 0.91<\Pr \left({c}_2\mid x,{\widehat{\theta}}_{\mathrm{M}2\mathrm{a}}\right)<0.93 \) for the 21 sites with a single nonsynonymous change and \( 0.55<\Pr \left({c}_2\mid x,{\widehat{\theta}}_{\mathrm{M}2\mathrm{a}}\right)<0.61 \) for the remaining sites [82]. Hence, the BEB approach mitigated the problem of low information content, as the posterior probability of positive selection at invariant sites was reduced. An alternative to BEB is called smoothed bootstrap aggregation (SBA) [38]. SBA entails drawing site patterns from X with replacement (i.e., bootstrap) to generate a set of alignments {X 1, …, X m} with similar information content as X. The MLEs \( {\left\{{\widehat{\theta}}_i\right\}}_{i=1}^m \) for the vector of model parameters θ are then estimated by fitting the CSM to each X i ∈{X 1, …, X m}. A kernel smoother is applied to these values to reduce sampling errors. The mean value of the resulting smoothed \( {\left\{{\widehat{\theta}}_i\right\}}_{i=1}^m \) is then used in Eq. 4 in place of the MLE for θ obtained from the original alignment to estimate posterior probabilities. This approach was shown to balance power and accuracy at least as well as BEB. But, SBA has the advantage that it can accommodate the uncertainty of all parameter estimates (not just those of the ω distribution, as in BEB) and is much easier to implement. When SBA was applied to the tax gene, the posterior probabilities for positive selection were further reduced: \( 0.87<\Pr \left({c}_2\mid x,{\widehat{\theta}}_{\mathrm{M}2\mathrm{a}}\right)<0.89 \) for the 21 sites with a single nonsynonymous change, and \( 0.55<\Pr \left({c}_2\mid x,{\widehat{\theta}}_{\mathrm{M}2\mathrm{a}}\right)<0.60 \) for the remaining sites [38].
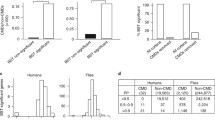
The problem of low information content was fairly obvious in the case of the tax gene, as 158 of the 181 codon sites within that dataset were invariant. However, it can sometimes be unclear whether there is enough variation in an alignment to ensure reliable inferences. It would be useful to have a method to determine whether a given data set might be problematic. An MLE \( \widehat{\theta} \) will always converge to a normal distribution centered at the true parameter value θ with variance proportional to 1∕n as the sample size n (a proxy for information content) gets larger, provided that the CSM satisfies certain “regularity” conditions (a set of technical conditions that must hold to guarantee that MLEs will converge in distribution to a normal, and that the LLR for any pair of nested models will converge to its expected chi-squared distribution). This expectation makes it possible to assess whether an alignment is sufficiently informative to obtain the benefits of regularity. The first step is to generate a set of bootstrap alignments {X 1, …, X m}. The CSM can then be fitted to these to produce a sample distribution \( {\left\{{\widehat{\theta}}_i\right\}}_{i=1}^m \) for the MLE of any model parameter θ. If the alignment is sufficiently informative with respect θ, then a histogram of \( {\left\{{\widehat{\theta}}_i\right\}}_{i=1}^m \) should be approximately normal in distribution. Serious departures from normality (e.g., a bimodal distribution) indicate unstable MLEs, which are a sign of insufficient information or an irregular modeling scenario. Mingrone et al. [38] recommend using this technique with real data as a means of gaining insight into potential difficulties of parameter estimation using a given CSM.
3.1.1 Irregularity and Penalized Likelihood
Issues associated with low information content can be made worse by violations of certain regularity conditions. For example, M2a is the same as M1a but for two extra parameters, p 2 and ω 2. Usual likelihood theory would therefore predict that the limiting distribution of the LLR is \( {\chi}_2^2 \). However, this result is valid only if the regularity conditions hold. Among these conditions is that the null model is not obtained by placing parameters of the alternate model on the boundary of parameter space. Since M1a is the same as M2a but with p 2 = 0, this condition is violated. The same can be said for many nested pairs of Phase I CSMs, such as M7 vs M8 [81] or M1 vs branch-site Model A [79]. Although the theoretical limiting distribution of the LLR under some irregular conditions has been determined by Self and Liang [54], those results do not include cases where one of the model parameters is unidentifiable under the null [2]. Since M1a is M2a with p 2 = 0, the likelihood under M1a is the same for any value of ω 2. This makes ω 2 unidentifiable under the null. The limiting distribution for the M1a vs M2a contrast is therefore unknown [74].
A penalized likelihood ratio test (PLRT; [39]) has been proposed to mitigate problems associated with unidentifiable parameters. Under this method, the likelihood function for the alternate model (e.g., M2a) is modified so that values of p 2 closer to zero are penalized. This has the effect of drawing the MLE for p 2 away from the boundary, and can interpreted as a way to “regularize” the model. PLRT seems to be more useful in cases where the analysis of a real alignment produces a small value of \( {\widehat{p}}_2 \) accompanied by an unrealistically large value of \( {\widehat{\omega}}_2 \). This can happen because \( {\widehat{\omega}}_2 \) is influenced by fewer and fewer site patterns as \( {\widehat{p}}_2 \) approaches zero, and is therefore subject to larger and larger sampling errors. In addition, \( {\widehat{\omega}}_2 \) and \( {\widehat{p}}_2 \) tend to be negatively correlated, which further contributes to the large sampling errors. For example, Mingrone et al. [39] found that M2a fitted to a 5-taxon alignment with 198 codon sites without penalization gave \( \left({\widehat{p}}_2,{\widehat{\omega}}_2\right)=\left(0.01,34.70\right) \). These MLEs, if taken at face value, suggest that a small number of sites in the gene underwent positive selection. However, such a large rate ratio is difficult to believe given that its estimate is consistent with only approximately 2 codon sites (e.g., an estimated 1% of the 198 sites or ≈2 sites). Using the PLRT, the MLEs were \( \left({\widehat{p}}_2,{\widehat{\omega}}_2\right)=\left(0.09,1.00\right) \). These suggest that selection pressure was nearly neutral at a significant proportion of sites in the gene. In this case, the rate ratio is consistent with 9% of the 198 sites or ≈18 sites and is therefore less likely to be an artifact of sampling error. We expect this approach to be useful in a wide variety of evolutionary applications that rely on mixture models to make inferences (e.g., [13, 34, 47, 66]).
Other approaches for dealing with low information content in the data for an individual gene include the empirical Bayes approach of Kosiol et al. [33] and the parametric bootstrapping methods of Gibbs [14]. Both methods exploit the additional information content available from other genes. Kosiol et al. [33] adopted an empirical Bayes approach, where ω values varied over edges and genes according to a distribution. Because empirical posterior distributions are used, the approach is more akin to detecting sites under positive selection (e.g., using NEB) than formal testing. By contrast, Gibbs [14] adopted a test-based approach and utilized parametric bootstrapping [15] to approximate the distribution of the likelihood ratio statistic using data from other genes to obtain parameter sets to use in the bootstrap. Whereas this approach can attenuate issues associated with low information content, it can also be computationally expensive, especially when applied to large alignments.
3.2 Case Study B: Model Misspecification
The mechanisms that give rise to the diversity of site patterns in a set of homologous genes are highly complex and not fully understood. CSMs are therefore necessarily simplified representations of the true generating process, and are in this sense misspecified. The extent to which misspecification might cause an omnibus LRT to falsely detect positive selection was of primary concern during Phase I of model development. We use a particular form of the YN-BSM called Model A [79] to illustrate this issue. In its original form, the omnibus LRT assumes a null under which a proportion p 0 of sites evolved under stringent selection with ω 0 = 0 and the remaining sites evolved under a neutral regime with ω 1 = 1 on all branches of the tree (i.e., model M1 in [44]). This is contrasted with Model A, which is the same as M1 except that it assumes that some stringent sites and some neutral sites evolved under positive selection with ω 2 > 1 on a prespecified branch called the foreground branch. The omnibus test contrasting M1 with Model A was therefore designed to detect a subset of sites that evolved adaptively on the same branch of the tree.
During this period of model development, the standard method to test the impact of misspecification on the reliability of an omnibus LRT was to generate alignments in silico using a more complex version of the CSM to be tested as the generating model. This usually involved adding more variability in ω across sites and/or branches than assumed by the fitted CSM while leaving all other aspects of the generating model the same. In Zhang [85], for example, alignments were generated using site-specific rate matrices, as in Eq. 1, with rate ratios ω specified by predetermined selection regimes, two of which are shown in Table 1. In one simulation, 200 alignments were generated using regime Z on a single foreground branch and regime X on all of the remaining branches of a 10 or 16 taxon tree. The gene therefore underwent a mixture of stringent selection and neutral evolution over most of the tree (regime X), but with complete relaxation of selection pressure on the foreground branch (regime Z). Positive selection did not occur at any of the sites. Nevertheless, the M1 vs Model A contrast inferred positive selection in 20–55% of the alignments, depending on the location of the foreground branch. Such a high rate of false positives was attributed to the mismatch between the process used to generate the data compared to the process assumed by the null model M1 [85].
The branch-site model was subsequently modified to allow 0 < ω 0 < 1 instead of ω 0 = 0 (Modified Model A in [86]). Furthermore, the new null model is specified under the assumption that some proportion p 0 of sites (the stringent sites) evolved under stringent selection with 0 < ω 0 < 1 everywhere in the tree except on the foreground branch, where those same sites evolved neutrally with ω 2 = 1. All other sites in the alignment (the neutral sites) are assumed to have evolved neutrally with ω 1 = 1 everywhere in the tree. This is contrasted with the Modified Model A, which assumes that some of the stringent sites and some of the neutral sites evolved under positive selection with ω 2 > 1 on the foreground. Hence, unlike the original omnibus test that contrasts M1 with Model A, the new test contrasts Modified Model A with ω 2 = 1 against Modified Model A with ω 2 > 1. These changes to the YN-BSM were shown to mitigate the problem of false inference. For example, using the same generating model with regimes X and Z, the modified omnibus test falsely inferred positive selection in only 1–7.5% of the alignments, consistent with the 5% level of significance of the test [86].
This case study demonstrates how problems associated with model misspecification were traditionally identified, and how they could be completely corrected through relatively minor changes to the model. However, the generating methods employed by studies such as Zhang [85] and Zhang et al. [86], although sophisticated for their time, produced alignments that were highly unrealistic compared to real data. For example, it was recently shown that a substantial proportion of variation in many real alignments might be due to selection effects associated with shifting balance over static site-specific fitness landscapes [25, 26]. This process results in random changes in site-specific rate ratios, or heterotachy, that cannot be replicated using traditional CSMs as the generating model. While the mitigation of statistical pathologies due to low information content (e.g., using BEB or SBA) or model misspecification (e.g., by altering the null and alternative hypotheses or the use of penalized likelihood) were critical advancements during Phase I of CSM development, other statistical pathologies went unrecognized due to reliance on unrealistic simulation methods. This issue is taken up in the next section.
4 Phase II: Advanced CSMs
A typical protein-coding gene evolves adaptively only episodically [59]. The evidence of adaptive evolution of this type can be very difficult to detect. For example, it is assumed under the YN-BSM that a random subset of sites switched from a stringent or neutral selection regime to positive selection together on the same set of foreground branches. The power to detect a signal of this kind can be very low when the proportion of sites that switched together is small [77]. Perhaps encouraged by the reliability of Phase I models demonstrated by extensive simulation studies [2, 3, 29, 31, 37, 70, 77, 82, 85, 86], combined with experimental validation of results obtained from their application to real data [1, 71, 76], investigators began to formulate increasingly complex and parameter-rich CSMs [31, 41, 48, 50, 55, 64, 65]. The hope was that carefully selected increases in model complexity would yield greater power to detect subtle signatures of positive selection overlooked by Phase I models. The introduction of such CSMs marks the beginning of Phase II of their historical development.
Phase II models fall into three broad categories:
-
1.
The first consists of Phase I CSMs modified to account for more variability in selection effects across sites and branches than previously assumed, with the aim of increasing the power to detect subtle signatures of positive selection (e.g., the branch-site random effects likelihood model, BSREL; [31]).
-
2.
The second category includes Phase I CSMs modified to contain parameters for mechanistic processes not directly associated with selection effects. Many such models have been motivated by a particular interest in the added mechanism (e.g., the fixation of double and triple mutations; [26, 40, 83]), or by the notion that increasing the mechanistic content of a CSM can only improve inferences about selection effects (e.g., by accounting for variations in the synonymous substitution rate; [30, 51]).
-
3.
The third category of models abandons the traditional formulation of Eq. 1 in favor of a substitution process expressed in terms of explicit population genetic parameters, such as population size and selection coefficients [45, 48,49,50, 64, 65].
An example of the first category of models is BSREL, which accounts for variations in selection effects across sites and over branches by assuming a different rate-ratio distribution \( \left\{\left({\omega}_i^b,{p}_i^b\right):i=1,\dots, {k}_b\right)\Big\} \) for each branch b of a tree [31]. BSREL was later found to be more complex than necessary, so an adaptive version was formulated to allow the number of components k b on a given branch to adjust to the apparent complexity of selection effects on that branch (aBSREL; [55]). A further reduction in model complexity led to the formulation of the test known as BUSTED (for branch-site unrestricted statistical test for episodic diversification; [41]), which we use to illustrate the problem of confounding in Case Study C. An example of the second category of models is the addition of parameters for the rate of double and triple mutations to traditional CSMs, the most sophisticated version of which is RaMoSSwDT (for Random Mixture of Static and Switching sites with fixation of Double and Triple mutations; [26]). This model is used in Case Study D to illustrate the problem of phenomenological load.
Models in the third category are the most ambitious CSMs currently in use, and are far more challenging to fit to real alignments than traditional models. One of the most impressive examples of their application is the site-wise mutation-selection model (swMutSel; [64, 65]) fitted to a concatenated alignment of 12 mitochondrial genes (3598 codon sites) from 244 mammalian species. Based on the mutation-selection framework of Halpern and Bruno [19], swMutSel estimates a vector of selection coefficients for each site in an alignment. This and similar models (e.g., [48,49,50]) appear to be reliable [58], but require a very large number of taxa (e.g., hundreds). Phase II models of this category are therefore impractical for the majority of empirical datasets. Here, we utilize MutSel as an effective means to generate realistic alignments with plausible levels of variation in selection effects across sites and over time rather than as a tool of inference.
4.1 Case Study C: Confounding
By expressing the codon substitution process in terms of explicit population genetic parameters, the MutSel framework facilitates the investigation of complex evolutionary dynamics, such as shifting balance on a fixed fitness landscape or adaptation to a change in selective constraints (i.e., a peak shift; [6, 25]) that are missing from alignments generated using traditional methods. Specifically, by assigning a different vector of fitness coefficients for the 20 amino acids to each site, MutSel can generate more variation in rate ratio across sites and over time than has been realized in the past simulation studies (e.g., Table 1). In this way, MutSel provides the basis of a generating model that can be adjusted to produce alignments that closely mimic real data [26]. MutSel therefore serves to connect demonstrably plausible evolutionary dynamics to the pathology we refer to as confounding.
Under MutSel, the dynamic regime at the hth codon site (e.g., shifting balance, neutral, nearly neutral, or adaptive evolution) is uniquely specified by a vector of fitness coefficients \( {\mathbf{f}}^h=<{f}_1^h,\dots, {f}_m^h> \). It is generally assumed that mutation to any of the three stop codons is lethal, so m = 61 for nuclear genes and m = 60 for mitochondrial genes. And, although it is not a requirement, it is typical to assume that the \( {f}_j^h \) are constant across synonymous codons [25, 57]. Given f h, the elements of a site-specific instantaneous rate matrix A h can be defined as follows for all i ≠ j (cf. Eq. 1):
where μ ij is the rate at which codon i mutates to codon j and \( {s}_{ij}^h=2{N}_e\left({f}_j^h-{f}_i^h\right) \) is the scaled selection coefficient for a population of haploids with effective population size N e. The probability that the new mutant j is fixed is approximated by \( {s}_{ij}^h/ \left\{1-\exp \left(-{s}_{ij}^h\right)\right\} \) [9, 28].
The rate matrix A h defines the dynamic regime for the site as illustrated in Fig. 3. The bar plot shows codon frequencies \( {\boldsymbol{\pi}}^h=<{\pi}_1^h,\dots, {\pi}_m^h> \) sorted in descending order. A site spends most of its time occupied by codons to the left or near the “peak” of its landscape. The codon-specific rate ratio for the site (\( d{N}_i^h/ d{S}_i^h \) for codon i) is low near the peak (red line plot in Fig. 3) since mutations away from the peak are seldom fixed. However, if selection is not too stringent, the site will occasionally drift to the right into the “tail” of its landscape. When this occurs, the codon-specific rate ratio will be elevated for a time until a combination of drift and positive selection moves the site back to its peak. This dynamic between selection and drift is reminiscent of Wright’s shifting balance. It implies that, when a population is evolving on a fixed fitness landscape (i.e., with no adaptive evolution), its gene sequences can nevertheless contain signatures of temporal changes in site-specific rate ratios (heterotachy), and that these might include evidence of transient elevation to values greater than one (i.e., positive selection). Such signatures of positive selection due to shifting balance can be detected by Phase II CSMs [25].
Fitness coefficients for the 20 amino acids were drawn from a normal distribution centered at zero and with standard deviation σ = 0.001. Bars show the resulting stationary frequencies (a proxy for fitness) sorted from largest to smallest. They compose a metaphorical site-specific landscape over which the site is imagined to move. The solid red line shows the codon-specific rate ratio \( d{N}_i^h/ d{S}_i^h \) for the sorted codons. This varies depending on the codon currently occupying the site, and can be greater than one following a chance substitution into the tail (to the right) of the landscape. In this case, the codon-specific rate ratio for the site ranged from 0.21 to 4.94 with a temporally averaged site-specific rate ratio of dN h∕dS h = 0.52
For example, BUSTED [41] was developed as an omnibus test for episodic adaptive evolution. The underlying CSM was formulated to account for variations in the intensity of selection over both sites and time modeled as a random effect. This is in contrast to the YN-BSM, which treats temporal changes in rate ratio as a fixed effect that occurs on a prespecified foreground branch (although the sites under positive selection are still a random effect). We therefore refer to the CSM underlying BUSTED as the random effects branch-site model (RE-BSM) to serve as a reminder of this important distinction. Under RE-BSM, the rate ratio at each site and branch combination is assumed to be an independent draw from the distribution \( \left\{\left({\omega}_0,{p}_0\right),\Big({\omega}_1,{p}_1\Big),\Big({\omega}_2,{p}_2\Big)\right\} \). In this way, the model accounts for variations in selection effects both across sites and over time. BUSTED contrasts the null hypothesis that ω 0 ≤ ω 1 ≤ ω 2 = 1 with the alternative that ω 0 ≤ ω 1 ≤ 1 ≤ ω 2. When applied to real data, rejection of the null is interpreted as evidence of episodic adaptive evolution.
Unlike the YN-BSM that aims to detect a subset of sites that underwent adaptive evolution together on the same foreground branches (i.e., coherently), BUSTED was designed to detect heterotachy similar to the type predicted by the mutation-selection framework: shifting balance on a static fitness landscape. Jones et al. [25] recently demonstrated that plausible levels of shifting balance can produce signatures of episodic positive selection that can be detected. BUSTED inferred episodic positive selection in as many as 40% of alignments generated using the MutSel framework. Significantly, BUSTED was correct to identify episodic positive selection in these trials. Even though the generating process assumed fixed site-specific landscapes (so there was no episodic adaptive evolution), and the long-run average rate ratio at each site was necessarily less than one [57], positive selection nevertheless did sometimes occur by shifting balance. This illustrates the general problem of confounding. Two processes are said to be confounded if they can produce the same or similar patterns in the data. In this case, episodic adaptive evolution (i.e., the evolutionary response to changes in site-specific landscapes) and shifting balance (i.e., evolution on a static fitness landscape) are confounded because they can both produce rate-ratio distributions that indicate episodic positive selection. The possibility of confounding underlines the fact that there are limitations in what can be inferred about evolutionary processes based on an alignment alone.
4.2 Case Study D: Phenomenological Load
Phenomenological load (PL) is a statistical pathology related to both model misspecification (Case Study B) and confounding (Case Study C) that was not recognized during Phase I of CSM development. When a model parameter that represents a process that played no role in the generation of an alignment (i.e., a misspecified process) nevertheless absorbs a significant amount of variation, its MLE is said to carry PL [26]. This is more likely to occur when the misspecified process is confounded with one or more other processes that did play a role in the generation of the data, and when a substantial proportion of the total variation in the data is unaccommodated by the null model [26]. PL increases the probability that a hypothesis test designed to detect the misspecified process will be statistically significant (as indicated by a large LLR) and can therefore lead to the incorrect conclusion that the misspecified process occurred. Critically, Jones et al. [26] showed that PL was only detected when model contrasts were fitted to data generated with realistic evolutionary dynamics using the MutSel model framework.
To illustrate the impact of PL, we consider the case of CSMs modified to detect the fixation of codons following simultaneous double and triple (DT) nucleotide mutations. The majority of CSMs currently in use assume that codons evolve by a series of single-nucleotide substitutions, with the probability for DT changes set to zero. However, recent model-based analyses have uncovered evidence for DT mutations [32, 68, 83]. Early estimates of the percentage of fixed mutations that are DT were perhaps unrealistically high. Kosiol et al. [32], for example, estimated a value close to 25% in an analysis of over 7000 protein families from the Pandit database [69]. Alternatively, when estimates were derived from a more realistic site-wise mutation-selection model, DT changes comprised less than 1% of all fixed mutations [64]. More recent studies suggest modest rates of between 1% and 3% [5, 20, 27, 53]. Whatever the true rate, several authors have argued that it would be beneficial to introduce a few extra parameters into a standard CSM to account for DT mutations (e.g., [40, 83]). The problem with this suggestion is that episodic fixation of DT mutations can produce signatures of heterotachy consistent with shifting balance.
Recall the comparison of M1, a CSM containing parameters represented by the vector θ 1, and M2, the same model but for the inclusion of one additional parameter ψ, so that θ 2 = (θ 1, ψ). The parameter ψ will reduce the deviance of M2 compared to M1 by some proportion of the baseline deviance between the simplest CSM (M0) and the saturated model \( {P}_{\mathrm{S}}\left({\widehat{\theta}}_{\mathrm{S}}\right) \). We call this the percent reduction in deviance (PRD) attributed to \( \widehat{\psi} \):
Suppose M1 and M2 were fitted to an alignment and that the \( \mathrm{LLR}=\Delta D\left({\widehat{\theta}}_{\mathrm{M}1},{\widehat{\theta}}_{\mathrm{M}2}\right) \) was found to be statistically significant. This would lead an analyst to attribute the \( \mathrm{PRD}\left(\widehat{\psi}\right) \) to real signal for the process ψ was meant to represent, possibly combined with some PL and noise. Now, consider the case in which the process represented by ψ did not actually occur (i.e., it was not a component of the true generating process). Under this scenario, \( \mathrm{PRD}\left(\widehat{\psi}\right) \) would contain no signal, but would be entirely due to PL plus noise. When this is known to be the case, we set \( \mathrm{PRD}\left(\widehat{\psi}\right)=\mathrm{PL}\left(\widehat{\psi}\right) \). As illustrated below, \( \mathrm{PL}\left(\widehat{\psi}\right) \) can be large enough to result in rejection of the null, and therefore lead to a false conclusion about the data generating process.
We illustrate PL by contrasting the model RaMoSS with a companion model RaMoSSwDT that accounts for the fixation of DT mutations via two rate parameters, α (the double mutation rate) and β (the triple mutation rate) [26]. RaMoSS combines the standard M-series model M3 with the covarion-like model CLM3 (cf., [12, 18]). Specifically, RaMoSS mixes (with proportion p M3) one model with two rate-ratio categories ω 0 < ω 1 that are constant over the entire tree with a second model (with proportion p CLM3 = 1 − p M3) under which sites switch randomly in time between \( {\omega}_0^{\prime }<{\omega}_1^{\prime } \) at an average rate of δ switches per unit branch length. Fifty alignments were simulated to mimic a real alignment of 12 concatenated H-strand mitochondrial DNA sequences (3331 codon sites) from 20 mammalian species as distributed in the PAML package [73]. The generating model, MutSel-mmtDNA [26], was based on the mutation-selection framework and produced alignments with single-nucleotide mutations only. Since DT mutations are not fixed under MutSel-mmtDNA, the PRD carried by \( \left(\widehat{\alpha},\widehat{\beta}\right) \) in each trial can be equated to PL (plus noise). The resulting distribution of \( \mathrm{PL}\left(\widehat{\alpha},\widehat{\beta}\right) \) is shown as a boxplot in Fig. 4.
The box plot depicts the distribution of the phenomenological load (PL) carried by \( \left(\hat{\alpha},\hat{\beta}\right) \) produced by fitting the RaMoSS vs RaMoSSwDT contrast to 50 alignments generated under MutSel-mmtDNA: the circles represent outliers of this distribution. The diamond is the percent reduction in deviance for the same parameters estimated by fitting RaMoSS vs RaMoSSwDT to the real mtDNA alignment
Although DT mutations were not fixed when the data was generated, shifting balance on a static landscape can produce similar site patterns as a process that includes rare fixation of DT mutations (site patterns exhibiting both synonymous and nonsynonymous substitutions; [26]).Footnote 3 DT and shifting balance are therefore confounded. And since shifting balance tends to occur at a substantial proportion (approximately 20%) of sites when an alignment is generated under MutSel-mmtDNA, DT mutations were falsely inferred by the LRT in 48 of 50 trials at the 5% level of significance (assuming LLR \( \approx {\chi}_2^2 \) for the two extra parameters α and β in RaMoSSwDT compared to RaMoSS). The PRD\( \left(\widehat{\alpha},\widehat{\beta}\right) \) when RaMoSS vs RaMoSSwDT was fitted to the real mmtDNA is shown as a diamond in the same plot. Although \( \left(\widehat{\alpha},\widehat{\beta}\right) \) estimated from the real mmtDNA were found to be highly significant (LLR = 84, p-value << 0.001), the \( \mathrm{PRD}\left(\widehat{\alpha},\widehat{\beta}\right) \) was found to be just under the 95th percentile of \( \mathrm{PL}\left(\widehat{\alpha},\widehat{\beta}\right) \) (PRD = 0.060% compared to the 95th percentile of PL = 0.061). The evidence for DT mutations in the real data is therefore only marginal, and it is reasonable to suspect that its \( \mathrm{PRD}\left(\widehat{\alpha},\widehat{\beta}\right) \), if not entirely the result of PL, is at least partially caused by PL.
5 Discussion
CSMs have been subjected to a certain degree of censure, particularly during Phase I of their development [11, 22, 23, 46, 60,61,62,63, 85]. We maintain that it is not the model in and of itself, or the maximum likelihood framework it is based on, that gives rise to statistical pathologies, but the relationship between model and data. This principle was illustrated by our analysis of the history of CSM development, which we divided into two phases. Phase I was characterized by the formulation of models to account for differences in selection effects across sites and over time that comprise the major component of variation in an alignment. Starting with M0, such models represent large steps toward the fitted saturated model in Fig. 2, and also provide a better representation of the true generating process. The main criticism of Phase I models was the possibility of falsely inferring positive selection in a gene or at an individual codon site [62, 63, 85]. But, the most compelling empirical case of false positives was shown to be the result of inappropriate application of a complex model to a sparse alignment [63]. Methods for identifying (bootstrap) and dealing with (BEB, SBA, and PLRT) low information content were illustrated in Case Study A.
The other big concern that arose during Phase I development was the possibility of pathologies associated with model misspecification. The method used to identify such problems was to fit a model to alignments generated under a scenario contrived to be challenging, as illustrated in Case Study B. There, the omnibus test based on Model A of the YN-BSM was shown to result in an excess of false positives when fitted to alignments simulated using the implausible but difficult “XZ” generating scenario (e.g., with complete relaxation of selection pressure at all sites on one branch of the tree; Table 1). Subsequent modifications to the test reduced the false positive rate to acceptable values. Hence, Case Study B underlines the importance of the model–data relationship. However, it is not clear whether a model adjusted to suit an unrealistic data-generating process is necessarily more reliable when fitted to a real alignment. This difficulty highlights the need to find ways, for the purpose of model testing and adjustment, to generate alignments that mimic real data as closely as possible.
Confidence in the CSM approach, combined with the exponential increase in the volume of genetic data and the growth of computational power, spurred the formulation CSMs of ever-increasing complexity during Phase II. The main issue with these models, which has not been widely appreciated, is confounding. Two processes are confounded if they can produce the same or similar patterns in the data. It is not possible to identify such processes when viewed through the narrow lens of an alignment (i.e., site patterns) alone. This was illustrated by Case Study C, where shifting balance on a static landscape was shown to be confounded with episodic adaptive evolution [7, 25]. Confounding can lead to what we call phenomenological load, as demonstrated in Case Study D. In that analysis, the parameters (α, β) were assigned a specific mechanistic interpretation, the rate at which double and triple mutations arise. It was shown that (α, β) can absorb variations in the data caused by shifting balance; hence, the MLEs \( \left(\widehat{\alpha},\widehat{\beta}\right) \) resulted in a significant reduction in deviance in 48/50 trials (Fig. 4), and therefore improved the fit of the model to the data. However, the absence of DT mutations in the generating process invalidated the intended interpretation of \( \left(\widehat{\alpha},\widehat{\beta}\right) \). This result underlines that a better fit does not imply a better mechanistic representation of the true generating process.
It is natural to assume that a better mechanistic representation of the true generating process can be achieved by adding parameters to our models to account for more of the processes believed to occur. The problem with this assumption is that the metric of model improvement under ML (reduction in deviance) is independent of mechanism. A parameter assigned a specific mechanistic interpretation is consequently vulnerable to confounding with other processes that can produce the same distribution of site patterns. As CSMs become more complex, it seems likely that the opportunity for confounding will only increase. It would therefore be desirable to assess each new model parameter for this possibility using something like the method shown in Fig. 4 whenever possible. The idea is to generate alignments using MutSel or some other plausible generating process in such a way as to mimic the real data as closely as possible, but with the new parameter set to its null value. To provide a second example, consider the test for changes in selection intensity in one clade compared to the remainder of the tree known as RELAX [67]. Under this model, it is assumed that each site evolved under a rate ratio randomly drawn from ω R = {ω 1, …, ω k} on a set of prespecified reference branches, and from a modified set of rate ratios \( {\boldsymbol{\omega}}_T=\left\{{\omega}_1^m,\dots, {\omega}_k^m\right\} \) on test branches, where m is an exponent. A value 0 < m < 1 moves the rate ratios in ω T closer to one compared to their corresponding values in ω R, consistent with relaxation of selection pressure at all sites on the test branches. Relaxation is indicated when the contrast of the null hypothesis that m = 1 versus the alternative that m < 1 is statistically significant. The distribution of PL(\( \widehat{m} \)) can be estimated from alignments generated with m = 1. The PRD(\( \widehat{m} \)) estimated from the real data can then be compared to this to assess the impact of PL (cf. Fig. 4). This approach is predicated on the existence of a generating model that could have plausibly produced the site patterns in the real data. Jones et al. [26] present a variety of methods for assessing the realism of a simulated alignment, although further development of such methods is warranted. Software based on MutSel is currently available for generating data that mimic large alignments of 100-plus taxa (Pyvolve; [56]). Other methods have been developed to mimic smaller alignments of certain types of genes (e.g., MutSel-mmtDNA; [25]). It is only by the use of these or other realistic simulation methods that the relationship between a given model and an alignment can be properly understood.
Notes
- 1.
The original CSM proposed by Goldman and Yang [16] was in fact quite complex in that it adjusted substitution rates between nonsynonymous codons to account for differences in physicochemical properties using the Grantham matrix [17]. This approach was later abandoned in favor of the simpler formulation now known as M0 [44], e.g., the first M-series model [81].
- 2.
Single-nucleotide (SN) mutations that are nonsynonymous occur more frequently than those that are synonymous due to idiosyncrasies in the genetic code. This is accounted for in the formulation of dN and dS, so that dN can be interpreted as the proportion of nonsynonymous SN mutations that are fixed. Likewise, dS is the proportion of synonymous SN mutations that are fixed. See Jones et al. [25] for a discussion of various interpretations of dN∕dS.
- 3.
It has previously been noted that the rapid fixation of compensatory mutations following substitution to an unstable base pair (e.g., AT→GT→GC) can also produce site patterns that suggest fixation of DT mutations [74, p. 46].
References
Anisimova M, Kosiol C (2009) Investigating protein-coding sequence evolution with probabilistic codon substitution models. Mol Biol Evol 26:255–271
Anisimova M, Bielawski JP, Yang ZH (2001) Accuracy and power of the likelihood ratio test in detecting adaptive molecular evolution. Mol Biol Evol 18:1585–1592
Anisimova M, Bielawski JP, Yang ZH (2002) Accuracy and power of Bayes prediction of amino acid sites under positive selection. Mol Biol Evol 19:950–958
Bielawski JP, Yang ZH (2004) A maximum likelihood method for detecting functional divergence at individual codon sites, with application to gene family evolution. J Mol Evol 59:121–132
De Maio N, Holmes I, Schlötterer C, Kosiol C (2013) Estimating empirical codon hidden Markov models. Mol Biol Evol 30:725–736
dos Reis M (2013). http://arxiv:1311.6682v1. Last accessed 26 Nov 2013
dos Reis M (2015) How to calculate the non-synonymous to synonymous rate ratio protein-coding genes under the Fisher-Wright mutation-selection framework. Biol Lett 11:1–4.
Field SF, Bulina MY, Kelmanson IV, Bielawski JP, Matz MV (2006) Adaptive evolution of multicolored fluorescent proteins in reef-building corals. J Mol Evol 62:332–339
Fisher R (1930) The distribution of gene ratios for rare mutations. Proc R Soc Edinb 50:205–220
Forsberg R, Christiansen FB (2003) A codon-based model of host-specific selection in parasites, with an application to the influenza a virus. Mol Biol Evol 20:1252–1259
Friedman R, Hughes AL (2007) Likelihood-ratio tests for positive selection of human and mouse duplicate genes reveal nonconservative and anomalous properties of widely used methods. Mol Phylogenet Evol 542:388–393
Galtier N (2001) Maximum-likelihood phylogenetic analysis under a covarion-like model. Mol Biol Evol 18:866–873
Gaston D, Susko E, Roger AJ (2011) A phylogenetic mixture model for the identification of functionally divergent protein residues. Bioinformatics 27:2655–2663
Gibbs RA (2007) Evolutionary and biomedical insights from the Rhesus macaque genome. Science 316:222–234
Goldman N (1993) Statistical tests of models of DNA substitution. J Mol Evol 36:182–198
Goldman N, Yang ZH (1994) Codon-based model of nucleotide substitution for protein-coding DNA-sequences. Mol Biol Evol 11:725–736
Grantham R (1974) Amino acid difference formula to help explain protein evolution. Science 862–864
Guindon S, Rodrigo AG, Dyer KA, Huelsenbeck JP (2004) Modeling the site-specific variation of selection patterns along lineages. Proc Natl Acad Sci USA 101:12957–12962
Halpern AL, Bruno WJ (1998) Evolutionary distances for protein-coding sequences: modeling site-specific residue frequencies. Mol Biol Evol 15:910–917
Harris K, Nielsen R (2014) Error-prone polymerase activity causes multinucleotide mutations in humans. Genome Res 9:1445–1554
Huelsenbeck JP, Dyer KA (2004) Bayesian estimation of positively selected sites. J Mol Evol 58:661–672
Hughes AL (2007) Looking for Darwin in all the wrong places: the misguided quest for positive selection at the nucleotide sequence level. Heredity 99:364–373
Hughes AL, Friedman R (2008) Codon-based tests of positive selection, branch lengths, and the evolution of mammalian immune system genes. Immunogenetics 60:495–506
Hughes AL, Nei M (1988) Pattern of nucleotide substitution at major histocompatibility complex class-1 loci reveals overdominant selection. Nature 335:167–170
Jones CT, Youssef N, Susko E, Bielawski JP (2017) Shifting balance on a static mutation-selection landscape: a novel scenario of positive selection. Mol Biol Evol 34:391–407
Jones CT, Youssef N, Susko E, Bielawski JP (2018) Phenomenological load on model parameters can lead to false biological conclusions. Mol Biol Evol 35:1473–1488
Keightley P, Trivedi U, Thomson M, Oliver F, Kumar S, Blaxter M (2009) Analysis of the genome sequences of three Drosophila melanogaster spontaneous mutation accumulation lines. Genet Res 19:1195–1201
Kimura M (1962) On the probability of fixation of mutant genes in a population. Genetics 47:713–719
Kosakovsky Pond SL, Frost SDW (2005) Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol Biol Evol 22:1208–1222
Kosakovsky Pond SL, Muse SV (2007) Site-to-site variations of synonymous substitution rates. Mol Biol Evol 22:2375–2385
Kosakovsky Pond SL, Murrell B, Fourment M, Frost SDW, Delport W, Scheffler K (2011) A random effects branch-site model for detecting episodic diversifying selection. Mol Biol Evol 28:3033–3043
Kosiol C, Holmes I, Goldman N (2007) An empirical codon model for protein sequence evolution. Mol Biol Evol 24:1464–1479
Kosiol C, Vinař T, daFonseca RR, Hubisz MJ, Bustamante CD, Nielsen R, Siepel A (2008) Patterns of positive selection in six mammalian genomes. PLoS Genet 4:1–17
Lartillot N, Philippe H (2004) A Bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Mol Biol Evol 21:1095–1109
Liberles DA, Teufel AI, Liu L, Stadler T (2013) On the need for mechanistic models in computational genomics and metagenomics. Genome Biol Evol 5:2008–2018
Lopez P, Casane D, Phillipe H (2002) Heterotachy, and important process of protein evolution. Mol Biol Evol 19:1–7
Lu A, Guindon S (2013) Performance of standard and stochastic branch-site models for detecting positive selection among coding sequences. Mol Biol Evol 31:484–495
Mingrone J, Susko E, Bielwaski JP (2016) Smoothed bootstrap aggregation for assessing selection pressure at amino acid sites. Mol Biol Evol 33:2976–2989
Mingrone J, Susko E, Bielwaski JP (2018) Modified likelihood ratio tests for positive selection (submitted). Bioinformatics, Advance Access https://doi.org/10.1093/bioinformatics/bty1019
Miyazawa S (2011) Advantages of a mechanistic codon substitution model for evolutionary analysis of protein-coding sequences. PLoS ONE 6:20
Murrell B, Weaver S, Smith MD, Wertheim JO, Murrell S, Aylward A, Eren K, Pollner T, Martin DP, Smith DM, Scheffler K, Pond SLK (2015) Gene-wide identification of episodic selection. Mol Biol Evol 32:1365–1371
Muse SV, Gaut BS (1994) A likelihood approach for comparing synonymous and nonsynonymous nucleotide substitution rates, with applications to the chloroplast genome. Mol Biol Evol 11:715–724
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Nielsen R, Yang ZH (1998) Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 148:929–936
Nielsen R, Yang Z (2003) Estimating the distribution of selection coefficients from phylogenetic data with applications to mitochondrial and viral DNA. Mol Biol Evol 20:1231–1239
Nozawa M, Suzuki Y, Nei M (2009) Reliabilities of identifying positive selection by the branch-site and the site-prediction methods. Proc Natl Acad Sci USA 106:6700–6705
Pagel M, Meade A (2004) A phylogenetic mixture model for detecting pattern-heterogeneity in gene sequence or character-state data. Syst Biol 53:571–581
Rodrigue N, Lartillot N (2014) Site-heterogeneous mutation-selection models with the PhyloBayes-MPI package. Bioinformatics 30:1020–1021
Rodrigue N, Lartillot N (2016) Detection of adaptation in protein-coding genes using a Bayesian site-heterogeneous mutation-selection codon substitution model. Mol Biol Evol 34:204–214
Rodrigue N, Philippe H, Lartillot N (2010) Mutation-selection models of coding sequence evolution with site-heterogeneous amino acid fitness profiles. Proc Natl Acad Sci USA 107:4629–4634
Rubinstein ND, Doron-Faigenboim A, Mayrose I, Pupko T (2011) Evolutionary model accounting for layers of selection in protein-coding genes and their impact on the inference of positive selection. Mol Biol Evol 28:3297–3308
Sawyer SL, Emerman M, Malik HS (2007) Discordant evolution of the adjacent antiretroviral genes trim22 and trim5 in mammals. PLoS Pathog 3:e197
Schrider D, Hourmozdi J, Hahn M (2014) Pervasive multinucleotide mutational events in eukaryotes. Curr Biol 21:1051–1054
Self SG, Liang KY (1987) Asymptotic properties of maximum likelihood estimators and likelihood ratio test under nonstandard conditions. J Am Stat Assoc 82:605–610
Smith MD, Wertheim JO, Weaver S, Murrell B, Scheffler K, Pond SLK (2015) Less is more: an adaptive branch-site random effects model for efficient detection of episodic diversifying selection. Mol Biol Evol 32:1342–1353
Spielman S, Wilke CO (2015) Pyvolve: a flexible Python module for simulating sequences along phylogenies. PLoS ONE 10:1–7
Spielman S, Wilke CO (2015) The relationship between dN/dS and scaled selection coefficients. Mol Biol Evol 34:1097–1108
Spielman S, Wilke CO (2016) Extensively parameterized mutation-selection models reliably capture site-specific selective constraints. Mol Biol Evol 33:2990–3001
Struder RA, Robinson-Rechavi M (2009) Evidence for an episodic model of protein sequence evolution. Biochem Soc Trans 37:783–786
Suzuki Y (2008) False-positive results obtained from the branch-site test of positive selection. Genes Genet Syst 83:331–338
Suzuki Y, Nei M (2001) Reliabilities of parsimony-based and likelihood-based methods for detecting positive selection at single amino acid sites. Mol Biol Evol 18:2179–2185
Suzuki Y, Nei M (2002) Simulation study of the reliability and robustness of the statistical methods for detecting positive selection at single amino acid sites. Mol Biol Evol 19:1865–1869
Suzuki Y, Nei M (2004) False-positive selection identified by ML-based methods: examples from the Sig1 gene of the diatom Thalassiosira weissflogii and the tax gene of the human T-cell lymphotropic virus. Mol Biol Evol 21:914–921
Tamuri AU, dos Reis M, Goldstein RA (2012) Estimating the distribution of selection coefficients from phylogenetic data using sitewise mutation-selection models. Genetics 190:1101–1115
Tamuri AU, Goldman N, dos Reis M (2014) A penalized-likelihood method to estimate the distribution of selection coefficients from phylogenetic data. Genetics 197:257–271
Wang H, Li K, Susko E, Rodger AJ (2008) A class frequency mixture model that adjusts for site-specific amino acid frequencies and improves inference of protein phylogeny. BMC Evol Biol 8:1–13
Wertheim JO, Murrell B, Smith MD, Pond SLK, Scheffler K (2014) Relax: detecting relaxed selection in a phylogenetic framework. Mol Biol Evol 32:820–832
Whelan S, Goldman N (2004) Estimating the frequency of events that cause multiple-nucleotide changes. Genetics 167:2027–2043
Whelan S, de Bakker PIW, Quevillon E, Rodriguez N, Goldman N (2006) Pandit: an evolution-centric database of protein and associated nucleotide domains with inferred trees. Nucleic Acids Res 34(Database issue):D327–D331
Wong WSW, Yang ZH, Goldman N, Nielsen R (2004) Accuracy and power of statistical methods for detecting adaptive evolution in protein coding sequences and for identifying positively selected sites. Genetics 168:1041–1051
Yang ZH (2005) The power of phylogenetic comparison in revealing protein function. Proc Natl Acad Sci USA 102:3179–3180
Yang ZH (2006) On the varied pattern of evolution in 2 fungal genomes: a critique of Hughes and Friedman. Mol Biol Evol 23:2279–2282
Yang ZH (2007) PAML4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang ZH (2014) Molecular evolution: a statistical approach. Oxford University Press, Oxford
Yang ZH (2017) PAML: phylogenetic analysis by maximum likelihood. http://abacus.gene.ucl.ac.uk/software/pamlDOC.pdf
Yang ZH, Bielawski JP (2000) Statistical methods for detecting molecular adaptation. Trends Ecol Evol 15:496–503
Yang ZH, dos Reis M (2011) Statistical properties of the branch-site test of positive selection. Mol Biol Evol 28:1217–1228
Yang ZH, Nielsen R (1998) Synonymous and nonsynonymous rate variation in nuclear genes of mammals. J Mol Evol 46: 409–418
Yang ZH, Nielsen R (2002) Codon-substitution models for detecting molecular adaptation at individual sites along specific lineages. Mol Biol Evol 19:908–917
Yang ZH, Nielsen R (2007) Mutation-selection models of codon substitution and their use to estimate selective strengths on codon usage. Mol Biol Evol 25:568–579
Yang ZH, Nielsen R, Goldman N, Pedersen AMK (2000) Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155:431–449
Yang ZH, Wong SWS, Nielsen R (2005) Bayes empirical Bayes inference of amino acid sites under positive selection. Mol Biol Evol 22:1107–1118
Zaheri M, Dib L, Salamin N. (2014) A generalized mechanistic codon model. Mol Biol Evol 31:2528–2541
Zhai W, Nielsen R, Goldman N, Yang ZH (2012) Looking for Darwin in genomic sequences – validity and success of statistical methods. Mol Biol Evol 20:2889–2893
Zhang J (2004) Frequent false detection of positive selection by the likelihood method with branch-site models. Mol Biol Evol 21:1332–1339
Zhang J, Nielsen R, Yang ZH (2005) Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol Biol Evol 22:2472–2479
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2019 The Author(s)
About this protocol
Cite this protocol
Jones, C.T., Susko, E., Bielawski, J.P. (2019). Looking for Darwin in Genomic Sequences: Validity and Success Depends on the Relationship Between Model and Data. In: Anisimova, M. (eds) Evolutionary Genomics. Methods in Molecular Biology, vol 1910. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9074-0_13
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9074-0_13
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9073-3
Online ISBN: 978-1-4939-9074-0
eBook Packages: Springer Protocols