Abstract
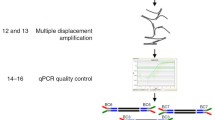
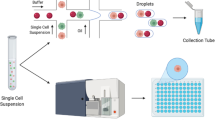
Single-cell RNA sequencing has evolved into a benchmark application to study cellular heterogeneity, advancing our understanding of cellular differentiation, disease progression, and gene regulation in a multitude of research areas. The generation of high-quality cDNA, an important step in the experimental workflow when generating sequence-ready libraries, is critical to maximizing data quality. Here we describe a strategy that uses a microfluidic device (i.e., the C1™ IFC) to synthesize full-length cDNA from single cells in a fully automated, nanoliter-scale format. The device also facilitates confirmation of the presence of a single, viable cell and recording of phenotypic information, quality control measures that are crucial for streamlining downstream data processing and enhancing overall data validity.
Similar content being viewed by others
References
Tang F, Barbacioru C, Wang Y, Nordman E, Lee C, Xu N, Wang X, Bodeau J, Tuch BB, Siddiqui A, Lao K, Surani MA (2009) mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods 6(5):377–382. https://doi.org/10.1038/nmeth.1315
Kolodziejczyk AA, Kim JK, Tsang JC, Ilicic T, Henriksson J, Natarajan KN, Tuck AC, Gao X, Buhler M, Liu P, Marioni JC, Teichmann SA (2015) Single cell RNA-sequencing of pluripotent states unlocks modular transcriptional variation. Cell Stem Cell 17(4):471–485. https://doi.org/10.1016/j.stem.2015.09.011
Zeisel A, Munoz-Manchado AB, Codeluppi S, Lonnerberg P, La Manno G, Jureus A, Marques S, Munguba H, He L, Betsholtz C, Rolny C, Castelo-Branco G, Hjerling-Leffler J, Linnarsson S (2015) Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347(6226):1138–1142. https://doi.org/10.1126/science.aaa1934
Stubbington MJ, Lonnberg T, Proserpio V, Clare S, Speak AO, Dougan G, Teichmann SA (2016) T cell fate and clonality inference from single-cell transcriptomes. Nat Methods 13(4):329–332. https://doi.org/10.1038/nmeth.3800
Grun D, van Oudenaarden A (2015) Design and analysis of single-cell sequencing experiments. Cell 163(4):799–810. https://doi.org/10.1016/j.cell.2015.10.039
Baker SC, Bauer SR, Beyer RP, Brenton JD, Bromley B, Burrill J, Causton H, Conley MP, Elespuru R, Fero M, Foy C, Fuscoe J, Gao X, Gerhold DL, Gilles P, Goodsaid F, Guo X, Hackett J, Hockett RD, Ikonomi P, Irizarry RA, Kawasaki ES, Kaysser-Kranich T, Kerr K, Kiser G, Koch WH, Lee KY, Liu C, Liu ZL, Lucas A, Manohar CF, Miyada G, Modrusan Z, Parkes H, Puri RK, Reid L, Ryder TB, Salit M, Samaha RR, Scherf U, Sendera TJ, Setterquist RA, Shi L, Shippy R, Soriano JV, Wagar EA, Warrington JA, Williams M, Wilmer F, Wilson M, Wolber PK, Wu X, Zadro R, External RNACC (2005) The external RNA controls consortium: a progress report. Nat Methods 2(10):731–734. https://doi.org/10.1038/nmeth1005-731
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Durruthy-Durruthy, R., Ray, M. (2018). Using Fluidigm C1 to Generate Single-Cell Full-Length cDNA Libraries for mRNA Sequencing. In: DiStefano, J. (eds) Disease Gene Identification. Methods in Molecular Biology, vol 1706. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7471-9_11
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7471-9_11
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7470-2
Online ISBN: 978-1-4939-7471-9
eBook Packages: Springer Protocols