Abstract
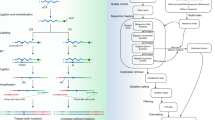
The base-modified nucleotide, N 6-methyladenosine, is a relatively abundant modification found in the mRNA of most higher eukaryotes. Methylation levels can change dependent upon environmental conditions, cell differentiation state, or following knockdown of members of the methylase complex, and it is often useful to directly measure and compare N 6-methyladenosine levels between samples. Two dimensional chromatography of radiolabeled nucleotides, following specific nuclease treatments, provides a robust, sensitive, and reproducible assay for this modification.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Wang Y, Li Y, Toth JI, Petroski MD, Zhang Z, Zhao JC (2014) N 6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nat Cell Biol 16:191–198. doi:10.1038/ncb2902
Wang X, Lu Z, Gomez A, Hon GC, Yue Y et al (2014) N 6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505:117–120 /nature12730
Bodi Z, Bottley A, Archer N, May ST, Fray RG (2015) Yeast m6A methylated mRNAs are enriched on translating ribosomes during meiosis, and under rapamycin treatment. PLoS One 10(7):e0132090. doi:10.1371/journal.pone.0132090
Meyer KD, Patil DP, Zhou J, Zinoviev A, Skabkin M, Elemento O et al (2015) 5′UTR m6A promotes cap-independent translation. Cell 163:999–1010
Horowitz S, Horowitz A, Nilsen TW, Munns TW, Rottman FM (1984) Mapping of N 6-methyladenosine residues in bovine prolactin mRNA. Proc Natl Acad Sci U S A 81:5667–5671
Kane SE, Beemon K (1985) Precise localization of m6A in Rous sarcoma virus RNA reveals clustering of methylation sites: Implications for RNA processing. Mol Cell Biol 5:2298–2306
Harper JE, Miceli SM, Roberts RJ, Manley JL (1990) Sequence specificity of the human mRNA N 6-adenosine methylase in vitro. Nucleic Acids Res 18:5735–5741
Schwartz S, Mumbach MR, Jovanovic M, Wang T, Maciag K et al (2014) Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5′ sites. Cell Rep 8:284–296. doi:10.1016/j.celrep.2014.05.048
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR (2012) Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149:1635–1646
Zhong S, Hongying L, Bodi Z, Button JD, Vespa L, Herzog M, Fray RG (2008) MTA is an Arabidopsis messenger RNA adenosine methylase and interacts with a homolog of a sex-specific splicing factor. Plant Cell 20:1278–1288
Sambrook J, Russel DW (2001) Molecular cloning : a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, pp. 7.13–7.17
Conrad RC, AMBION, INC. High efficiency mRNA isolation. United States Patent Application Publication, Pub. No.: US 2004/0230048 A1 Conrad Pub. Date: 18 Nov 2004.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Bodi, Z., Fray, R.G. (2017). Detection and Quantification of N 6-Methyladenosine in Messenger RNA by TLC. In: Lusser, A. (eds) RNA Methylation. Methods in Molecular Biology, vol 1562. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6807-7_6
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6807-7_6
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6805-3
Online ISBN: 978-1-4939-6807-7
eBook Packages: Springer Protocols