Abstract
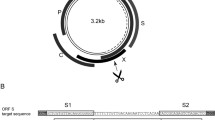
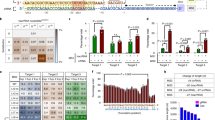
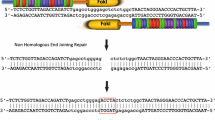
Gene editing using designer nucleases is now widely used in many fields of molecular biology. The technology is being developed for the treatment of viral infections such as persistant hepatitis B virus (HBV). The replication intermediate of HBV comprising covalently closed circular DNA (cccDNA) is stable and resistant to available licensed antiviral agents. Advancing gene editing as a means of introducing targeted mutations into cccDNA thus potentially offers the means to cure infection by the virus. Essentially, targeted mutations are initiated by intracellular DNA cleavage, then error-prone nonhomologous end joining results in insertions and deletions (indels) at intended sites. Characterization of these mutations is crucial to confirm activity of potentially therapeutic nucleases. A convenient tool for evaluation of the efficiency of target cleavage is the single strand-specific endonuclease, T7EI. Assays employing this enzyme entail initial amplification of DNA encompassing the targeted region. Thereafter the amplicons are denatured and reannealed to allow hybridization between indel-containing and wild-type sequences. Heteroduplexes that contain mismatched regions are susceptible to action by T7EI and cleavage of the hybrid amplicons may be used as an indicator of efficiency of designer nucleases. The protocol described here provides a method of isolating cccDNA from transfected HepG2.2.15 cells and evaluation of the efficiency of mutation by a transcription activator-like effector nuclease that targets the surface open reading frame of HBV.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Carroll D (2014) Genome engineering with targetable nucleases. Annu Rev Biochem 83:409–439
Mehta A, Haber JE (2014) Sources of DNA double-strand breaks and models of recombinational DNA repair. Cold Spring Harb Perspect Biol 6(9):a016428
Schiffer JT, Aubert M, Weber ND, Mintzer E, Stone D, Jerome KR (2012) Targeted DNA mutagenesis for the cure of chronic viral infections. J Virol 86(17):8920–8936
Bloom K, Ely A, Mussolino C, Cathomen T, Arbuthnot P (2013) Inactivation of hepatitis B virus replication in cultured cells and in vivo with engineered transcription activator-like effector nucleases. Mol Ther 21(10):1889–1897
Dong C, Qu L, Wang H, Wei L, Dong Y, Xiong S (2015) Targeting hepatitis B virus cccDNA by CRISPR/Cas9 nuclease efficiently inhibits viral replication. Antiviral Res 118:110–117
Cradick TJ, Keck K, Bradshaw S, Jamieson AC, McCaffrey AP (2010) Zinc-finger nucleases as a novel therapeutic strategy for targeting hepatitis B virus DNAs. Mol Ther 18(5):947–954
Chen J, Zhang W, Lin J, Wang F, Wu M, Chen C, Zheng Y, Peng X, Li J, Yuan Z (2014) An efficient antiviral strategy for targeting hepatitis B virus genome using transcription activator-like effector nucleases. Mol Ther 22(2):303–311
Levrero M, Pollicino T, Petersen J, Belloni L, Raimondo G, Dandri M (2009) Control of cccDNA function in hepatitis B virus infection. J Hepatol 51(3):581–592
Newbold JE, Xin H, Tencza M, Sherman G, Dean J, Bowden S, Locarnini S (1995) The covalently closed duplex form of the hepadnavirus genome exists in situ as a heterogeneous population of viral minichromosomes. J Virol 69(6):3350–3357
Tuttleman JS, Pourcel C, Summers J (1986) Formation of the pool of covalently closed circular viral DNA in hepadnavirus-infected cells. Cell 47(3):451–460
Pollicino T, Belloni L, Raffa G, Pediconi N, Squadrito G, Raimondo G, Levrero M (2006) Hepatitis B virus replication is regulated by the acetylation status of hepatitis B virus cccDNA-bound H3 and H4 histones. Gastroenterology 130(3):823–837
Caruntu FA, Molagic V (2005) CccDNA persistence during natural evolution of chronic VHB infection. Rom J Gastroenterol 14(4):373–377
Zoulim F (2004) Antiviral therapy of chronic hepatitis B: can we clear the virus and prevent drug resistance? Antivir Chem Chemother 15(6):299–305
Mussolino C, Morbitzer R, Lutge F, Dannemann N, Lahaye T, Cathomen T (2011) A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic Acids Res 39(21):9283–9293
Sells MA, Chen ML, Acs G (1987) Production of hepatitis B virus particles in Hep G2 cells transfected with cloned hepatitis B virus DNA. Proc Natl Acad Sci U S A 84(4):1005–1009
Abramoff MD, Magelhaes PJ, Ram SJ (2004) Image processing with ImageJ. Biophoton Int 11(12):36–42
Wong DK, Yuen MF, Yuan H, Sum SS, Hui CK, Hall J, Lai CL (2004) Quantitation of covalently closed circular hepatitis B virus DNA in chronic hepatitis B patients. Hepatology 40(3):727–737
Guschin DY, Waite AJ, Katibah GE, Miller JC, Holmes MC, Rebar EJ (2010) A rapid and general assay for monitoring endogenous gene modification. Methods Mol Biol 649:247–256
Doyon Y, Choi VM, Xia DF, Vo TD, Gregory PD, Holmes MC (2010) Transient cold shock enhances zinc-finger nuclease-mediated gene disruption. Nat Methods 7(6):459–460
Guo Y, Guo H, Zhang L, Xie H, Zhao X, Wang F, Li Z, Wang Y, Ma S, Tao J, Wang W, Zhou Y, Yang W, Cheng J (2005) Genomic analysis of anti-hepatitis B virus (HBV) activity by small interfering RNA and lamivudine in stable HBV-producing cells. J Virol 79(22):14392–14403
Reyon D, Tsai SQ, Khayter C, Foden JA, Sander JD, Joung JK (2012) FLASH assembly of TALENs for high-throughput genome editing. Nat Biotechnol 30(5):460–465
Acknowledgments
Work in the author’s laboratory is generously supported by funding from the National Research Foundation (NRF, GUIS 81768, 81692, 68339, 85981, and 77954) of South Africa, South African Medical Research Council (MRC), Poliomyelitis Research Foundation (PRF) and from the German Research Foundation (DFG).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Bloom, K., Ely, A., Arbuthnot, P. (2017). A T7 Endonuclease I Assay to Detect Talen-Mediated Targeted Mutation of HBV cccDNA. In: Guo, H., Cuconati, A. (eds) Hepatitis B Virus. Methods in Molecular Biology, vol 1540. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6700-1_8
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6700-1_8
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6698-1
Online ISBN: 978-1-4939-6700-1
eBook Packages: Springer Protocols