Abstract
In archaea and bacteria the major classes of RNAs are synthesized by one DNA-dependent RNA polymerase (RNAP). In contrast, most eukaryotes have three highly specialized RNAPs to transcribe the nuclear genome. RNAP I synthesizes almost exclusively ribosomal (r)RNA, RNAP II synthesizes mRNA as well as many noncoding RNAs involved in RNA processing or RNA silencing pathways and RNAP III synthesizes mainly tRNA and 5S rRNA. This review discusses functional differences of the three nuclear core RNAPs in the yeast S. cerevisiae with a particular focus on RNAP I transcription of nucleolar ribosomal (r)DNA chromatin.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
- RNA polymerase I
- RNA polymerase II
- RNA polymerase III
- Transcription
- Chromatin
- Nucleosomes
- Ribosomal RNA genes
- Transcription factors
- Gene expression
- Yeast
- Saccharomyces cerevisiae
1 RNA Polymerase I Has Only One Essential Genomic Target
Nuclear yeast RNAPs are protein complexes consisting of 12 (RNAP II ), 14 (RNAP I ) and 17 (RNAP III ) subunits. They all share a conserved architecture of the RNAP core, which catalyzes the highly accurate polymerization of RNA from single NTP molecules [1,2,3] (see review by Pilsl and Engel in this issue). Based on structural analyses and on functional in vitro assays, on the one hand the molecular mechanisms driving the RNA polymerization appear to be similar in all RNAPs. On the other hand, all three enzymes have special features, supporting their specific in vivo tasks [1, 2]. Thus, RNAP II has to transcribe thousands of different genes and produces transcripts from a few hundred to several thousand of bases in length. To account for dynamic changes in cellular gene expression RNAP II has (a) to recognize many promoters, (b) to access differently modified chromatin templates, which probably requires to adjust its elongation and termination properties. To fulfill these multiple tasks, RNAP II interacts with many different factors at each step of the transcription process [4, 5]. In contrast, RNAP III recognizes only three distinct classes of promoters and has probably a distinct transcription termination mechanism [6]. RNAP III synthesizes mainly short noncoding RNAs (typically <200 bp) with high efficiency in rapidly growing cells. Accordingly, the RNAP III transcription machinery is rather well defined [7,8,9,10]. Finally, the yeast RNAP I transcription machinery has only one known genomic target, the multicopy 9.1 kb 35S rRNA genes [11]. The rRNA genes are transcribed at very high rates accounting for up to 60% of RNA synthesis upon cellular growth [12]. Accordingly, electron micrographs of chromatin spreads show an extremely high density of RNAP I molecules at rRNA genes, whereas most RNAP II-dependent genes are only sparsely covered with polymerases [13,14,15]. Furthermore, RNAP I and RNAP III-dependent genes are constitutively transcribed in actively dividing cells and—as opposed to the majority of RNAP II transcribed genes—apparently devoid of nucleosomes (see as review [16,17,18] and Schächner et al., within this issue).
2 RNA Polymerase I Contains Additional Subunits Resembling Transcription Factors of RNAP II
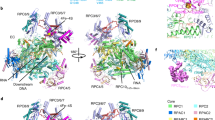
RNAPs I, II, and III contain ten conserved subunits which form the catalytic core . In addition, all three enzymes contain a 2-subunit stalk structure, which is distantly related and consists of subunits A14/A43, Rpb4/Rpb7 and C17/C25 in RNAPs I, II and III, respectively. In RNAP III an additional heterotrimer C82/C34/C3 connects the stalk with the RNAP III clamp , which probably helps to open the DNA duplex [19]. The overall architecture of the three RNAPs differs mainly in vicinity of the lobe structure, which is formed by the second largest subunits Rpa135 , Rpb2 and Rpc128 , respectively. Only one subunit—Rpb9 —is bound to the RNAP II lobe, whereas the heterodimer A34.5/49 and subunit A12.2 bind to the lobe of RNAP I , and the homologous C17/C25 and C11 subunits to the lobe of RNAP III . RNAP I subunits A34.5 and A49 consist of three subdomains: a dimerization module formed by A34.5 and the N-terminal part of A49 (full length A34.5 and aa 1–110 of A49); the A49 linker (aa 105–187 of A49); and the C-terminal part of A49 (aa 187–415). The dimerization module binds to the “lobe” and “external” domains of the second largest Pol I subunit A135 on the core module side [20,21,22]. In contrast, the C-terminal part of A49 which contains a tandem winged helix can be detected at the upstream face of the clamp core in several states of transcriptional active RNAP I molecules and seems to be flexible attached [20, 21, 23,24,25,26,27,28,29,30,31]. Biochemical and cell biological experiments showed that A34.5/A49 support transcription initiation , enhance RNAP I elongation and stimulate the intrinsic RNAP I RNA cleavage activity [26, 28, 32,33,34,35,36]. RNA cleavage activity depends on the dimerization module which is located in close proximity to A12.2 whose C-terminal part is also important for efficient RNA cleavage [26]. Deletion of A12.2 results in growth inhibition at elevated temperature, sensitivity to nucleotide-reducing drugs, and inefficient transcription termination ; hampers the assembly of the RNAP I enzyme; and leads to incorporation of wrong NTPs [37,38,39,40]. The lack of A12.2 may also lead to the loss of A34.5/A49, and might influence the intrinsic stability of elongation and termination complexes [40, 41].
Based on amino acid sequence similarities, position on the enzyme and function, the heterodimer formed by A34.5 and the N-terminus of A49 was suggested to be homologous to the RNAP II transcription factor TFIIF [26, 33]. TFIIF is predominantly found at promoter-proximal regions suggesting a crucial role in transcription initiation [42, 43]. On the other hand, TFIIF was suggested to leave the promoter—at least transiently—in complex with RNAP II , likely supporting early elongation [42,43,44,45]. A role of TFIIF in RNAP II elongation is further corroborated by in vitro studies, where it increases transcription rates by suppressing RNAP II pausing [46,47,48]. Several studies propose that TFIIF promotes transcription elongation in concert with the RNA cleavage supporting factor TFIIS , which structurally resembles the C-terminus of the RNAP I subunit A12.2 (reviewed in [3, 48,49,50,51]. In the RNAP II system, TFIIF and TFIIS are independently capable to release arrested RNAP II to resume productive elongation . However, these factors may synergistically enhance resumption of RNAP II transcription especially in conditions when the paused enzyme has additionally backtracked on the template [49]. Backtracked RNAP I requires the C-terminal, RNA cleavage activating part of A12.2 to resume elongation [52]. It is, however, unknown if the heterodimer A34.5/A49 participates in this process.
Finally, the C-terminus of A49 structurally and functionally resembles the tandem winged helix of TFIIE [33]. As TFIIE in RNAP II transcription , the C-terminal domain of A49 binds DNA and supports promoter-dependent transcription initiation in vitro [28] and RNAP I promoter recruitment in vivo [32]. In contrast to TFIIE, the A49 subunit stays associated with the enzyme after promoter clearance in vivo [32], and supports elongation of RNA from a DNA/RNA scaffold in vitro [33]. Recent studies suggest a more dynamic association of the heterodimer A34.5/A49 to the RNAP I lobe, since the heterodimer was absent from the core enzyme and A12.2 C-terminus was rearranged when RNAP I elongation was artificially blocked by addition of a nonhydrolyzable nucleotide [53].
A specific challenge for all elongating RNAPs is the transcription of chromatin templates. Since RNAP I and III transcribe nucleosome-depleted chromatin templates ( [13,14,15] see short review of Schächner et al. this issue). This indicates that nucleosomes are displaced from the chromatin template in the initial round of transcription , and it is possible that the lobe associated subunits of RNAP I and III may be involved in the process of nucleosome depletion.
3 Nuclear RNAPs Transcribe Chromatin Templates
In eukaryotic cells, nuclear DNA is assembled into repeated units called nucleosomes consisting of 146 bp of DNA wrapped around an octameric complex of histone proteins [54]. Nucleosomes generally provide a strong barrier for elongating RNAP II in vitro [46, 55, 56]. DNA attached to nucleosomes recoils on the octamer, locking the enzyme in an arrested state [57] thereby providing four major superhelical pausing sites [58]. Additional factors are required for passage of RNAP II through this barrier (see below). The mechanism how purified RNAP II complexes passes nucleosomes in vitro was thoroughly studied [59, 60]. Whether assembled nucleosomes stay associated or are evicted during RNAP II transcription depends on the formation of a small intranucleosomal DNA loop and on the transcription efficiency (rate) [61]. Accordingly, various RNAP II complexes can remodel chromatin to a different extent [59, 60].
It was suggested, that RNAP II can only pass nucleosomes if uncoiling of the DNA from the surface of the octamer is facilitated and if transcription elongation factors keep the polymerase in a transcriptionally competent state [62]. TFIIF and TFIIS may prevent the release of RNAP II at nucleosomal barriers and thereby support transcription through a nucleosome in a synergistic manner [63]. Passage of purified RNAP II through in vitro assembled nucleosomes was also supported by elongation factor Spt4/Spt5 [64] and in the presence of TFIIS together with Spt4/5 and elongation factor Elf1 [65]. Insights in the molecular mechanism how Spt4/5 together with Elf1 facilitate progression of RNAP II through a nucleosome were recently obtained using high resolution structures by cryo-EM of different stalled elongation complexes [65]. Other factors that were reported to partially disassemble nucleosomes similar to Spt4/Spt5 are the histone chaperone FACT and Paf1c [66,67,68].
Much less is known about how RNAP I and RNAP III interact with nucleosomes in vitro. However, in vivo there is ample evidence that RNAP I and RNAP III genes are largely devoid of nucleosomes (reviewed in [16,17,18], see short review of Schächner et al. in this issue). It is an open question how RNAP I and RNAP III deal with nucleosomal genes in the initial round of transcription . In contrast to RNAP II which has only subunit Rpb9 associated to the lobe structure, yeast RNAP I and RNAP III have the TFIIF- and TFIIS-homologous subunits A34.5/A49, A12.2 (RNAP I ), and C37/C53, C11 (RNAP III ) tightly associated to the lobe. Similar to TFIIF and TFIIS in RNAP II transcription , the homologous Pol I subunits A34.5/A49 and A12.2 facilitate RNAP I passage through nucleosomes [35]. Depletion of either the heterodimeric subunits A34.5/A49 or the cleavage supporting activity of A12.2 resulted in both reduced Pol I processivity and impaired passage though nucleosomes [35]. It is tempting to speculate that the homologous RNAP III subunits could play a similar role in RNAP III chromatin transcription . Furthermore, additional factors like FACT , Spt4/Spt5, or Paf1c have all been suggested to support RNAP I elongation [69,70,71,72]. Appropriate in vitro transcription system using highly purified factors and defined nucleosome templates will be the key to elucidate details of the molecular mechanisms of RNAP I and RNAP III transcription in the context of chromatin (see chapter by Merkl et al. in this issue).
References
Engel C, Neyer S, Cramer P (2018) Distinct mechanisms of transcription initiation by RNA polymerases I and II. Annu Rev Biophys 47:425–446
Khatter H, Vorländer MK, Müller CW (2017) RNA polymerase I and III: similar yet unique. Curr Opin Struct Biol 47:88–94
Vannini A, Cramer P (2012) Conservation between the RNA polymerase I, II, and III transcription initiation machineries. Mol Cell 45:439–446
Chen FX, Smith ER, Shilatifard A (2018) Born to run: control of transcription elongation by RNA polymerase II. Nat Rev Mol Cell Biol 19:464–478
Haberle V, Stark A (2018) Eukaryotic core promoters and the functional basis of transcription initiation. Nat Rev Mol Cell Biol 19:621–637
Arimbasseri AG, Rijal K, Maraia RJ (2013) Transcription termination by the eukaryotic RNA polymerase III. Biochim Biophys Acta 1829:318–330
Arimbasseri AG, Rijal K, Maraia RJ (2014) Comparative overview of RNA polymerase II and III transcription cycles, with focus on RNA polymerase III termination and reinitiation. Transcription 5:e27639
Graczyk D, Cieśla M, Boguta M (2018) Regulation of tRNA synthesis by the general transcription factors of RNA polymerase III - TFIIIB and TFIIIC, and by the MAF1 protein. Biochim Biophys Acta Gene Regul Mech 1861:320–329
Moir RD, Willis IM (2013) Regulation of pol III transcription by nutrient and stress signaling pathways. Biochim Biophys Acta 1829:361–375
Willis IM, Moir RD (2018) Signaling to and from the RNA polymerase III transcription and processing machinery. Annu Rev Biochem 87:75–100
Nogi Y, Yano R, Nomura M (1991) Synthesis of large rRNAs by RNA polymerase II in mutants of Saccharomyces cerevisiae defective in RNA polymerase I. Proc Natl Acad Sci U S A 88:3962–3966
Warner JR (1999) The economics of ribosome biosynthesis in yeast. Trends Biochem Sci 24:437–440
French SL, Osheim YN, Cioci F, Nomura M, Beyer AL (2003) In exponentially growing Saccharomyces cerevisiae cells, rRNA synthesis is determined by the summed RNA polymerase I loading rate rather than by the number of active genes. Mol Cell Biol 23:1558–1568
French SL, Osheim YN, Schneider DA, Sikes ML, Fernandez CF, Copela LA, Misra VA, Nomura M, Wolin SL, Beyer AL (2008) Visual analysis of the yeast 5S rRNA gene transcriptome: regulation and role of La protein. Mol Cell Biol 28:4576–4587
Laird CD, Chooi WY (1976) Morphology of transcription units in Drosophila melanogaster. Chromosoma 58:193–218
Hamperl S, Wittner M, Babl V, Perez-Fernandez J, Tschochner H, Griesenbeck J (2013) Chromatin states at ribosomal DNA loci. Biochim Biophys Acta 1829:405–417
Morse RH, Roth SY, Simpson RT (1992) A transcriptionally active tRNA gene interferes with nucleosome positioning in vivo. Mol Cell Biol 12:4015–4025
Moss T, Mars J-C, Tremblay MG, Sabourin-Felix M (2019) The chromatin landscape of the ribosomal RNA genes in mouse and human. Chromosome Res 27(1-2):31–40. https://doi.org/10.1007/s10577-018-09603-9
Hoffmann NA, Jakobi AJ, Moreno-Morcillo M, Glatt S, Kosinski J, Hagen WJH, Sachse C, Müller CW (2015) Molecular structures of unbound and transcribing RNA polymerase III. Nature 528:231–236
Engel C, Sainsbury S, Cheung AC, Kostrewa D, Cramer P (2013) RNA polymerase I structure and transcription regulation. Nature 502:650–655
Fernández-Tornero C, Moreno-Morcillo M, Rashid UJ, Taylor NMI, Ruiz FM, Gruene T, Legrand P, Steuerwald U, Müller CW (2013) Crystal structure of the 14-subunit RNA polymerase I. Nature 502:644–649
Jennebach S, Herzog F, Aebersold R, Cramer P (2012) Crosslinking-MS analysis reveals RNA polymerase I domain architecture and basis of rRNA cleavage. Nucleic Acids Res 40:5591–5601
Engel C, Gubbey T, Neyer S, Sainsbury S, Oberthuer C, Baejen C, Bernecky C, Cramer P (2017) Structural basis of RNA polymerase I transcription initiation. Cell 169:120–131.e22
Han Y, Yan C, Nguyen THD, Jackobel AJ, Ivanov I, Knutson BA, He Y (2017) Structural mechanism of ATP-independent transcription initiation by RNA polymerase I. elife 6:e27414
Kostrewa D, Kuhn C-D, Engel C, Cramer P (2015) An alternative RNA polymerase I structure reveals a dimer hinge. Acta Crystallogr D Biol Crystallogr 71:1850–1855
Kuhn C-D, Geiger SR, Baumli S, Gartmann M, Gerber J, Jennebach S, Mielke T, Tschochner H, Beckmann R, Cramer P (2007) Functional architecture of RNA polymerase I. Cell 131:1260–1272
Neyer S, Kunz M, Geiss C, Hantsche M, Hodirnau V-V, Seybert A, Engel C, Scheffer MP, Cramer P, Frangakis AS (2016) Structure of RNA polymerase I transcribing ribosomal DNA genes. Nature 540(7634):607–610. https://doi.org/10.1038/nature20561
Pilsl M, Crucifix C, Papai G, Krupp F, Steinbauer R, Griesenbeck J, Milkereit P, Tschochner H, Schultz P (2016) Structure of the initiation-competent RNA polymerase I and its implication for transcription. Nat Commun 7:12126
Sadian Y, Tafur L, Kosinski J, Jakobi AJ, Wetzel R, Buczak K, Hagen WJ, Beck M, Sachse C, Müller CW (2017) Structural insights into transcription initiation by yeast RNA polymerase I. EMBO J 36:2698–2709
Sanz-Murillo M, Xu J, Belogurov GA, Calvo O, Gil-Carton D, Moreno-Morcillo M, Wang D, Fernández-Tornero C (2018) Structural basis of RNA polymerase I stalling at UV light-induced DNA damage. Proc Natl Acad Sci U S A 115:8972–8977
Tafur L, Sadian Y, Hoffmann NA, Jakobi AJ, Wetzel R, Hagen WJH, Sachse C, Müller CW (2016) Molecular structures of transcribing RNA polymerase I. Mol Cell 64:1135–1143
Beckouet F, Labarre-Mariotte S, Albert B, Imazawa Y, Werner M, Gadal O, Nogi Y, Thuriaux P (2008) Two RNA polymerase I subunits control the binding and release of Rrn3 during transcription. Mol Cell Biol 28:1596–1605
Geiger SR, Lorenzen K, Schreieck A, Hanecker P, Kostrewa D, Heck AJR, Cramer P (2010) RNA polymerase I contains a TFIIF-related DNA-binding subcomplex. Mol Cell 39:583–594
Liljelund P, Mariotte S, Buhler JM, Sentenac A (1992) Characterization and mutagenesis of the gene encoding the A49 subunit of RNA polymerase A in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 89:9302–9305
Merkl PE, Pilsl M, Fremter T, Schwank K, Engel C, Längst G, Milkereit P, Griesenbeck J, Tschochner H (2020) RNA polymerase I (Pol I) passage through nucleosomes depends on Pol I subunits binding its lobe structure. J Biol Chem 295:4782–4795
Knutson BA, McNamar R, Rothblum LI (2020) Dynamics of the RNA polymerase I TFIIF/TFIIE-like subcomplex: a mini-review. Biochem Soc Trans 48:1917–1927
Gout J-F, Li W, Fritsch C, Li A, Haroon S, Singh L, Hua D, Fazelinia H, Smith Z, Seeholzer S, Thomas K, Lynch M, Vermulst M (2017) The landscape of transcription errors in eukaryotic cells. Sci Adv 3:e1701484
Nogi Y, Yano R, Dodd J, Carles C, Nomura M (1993) Gene RRN4 in Saccharomyces cerevisiae encodes the A12.2 subunit of RNA polymerase I and is essential only at high temperatures. Mol Cell Biol 13:114–122
Prescott EM, Osheim YN, Jones HS, Alen CM, Roan JG, Reeder RH, Beyer AL, Proudfoot NJ (2004) Transcriptional termination by RNA polymerase I requires the small subunit Rpa12p. Proc Natl Acad Sci U S A 101:6068–6073
Van Mullem V, Landrieux E, Vandenhaute J, Thuriaux P (2002) Rpa12p, a conserved RNA polymerase I subunit with two functional domains. Mol Microbiol 43:1105–1113
Appling FD, Scull CE, Lucius AL, Schneider DA (2018) The A12.2 subunit is an intrinsic destabilizer of the RNA polymerase I elongation complex. Biophys J 114:2507–2515
Krogan NJ, Kim M, Ahn SH, Zhong G, Kobor MS, Cagney G, Emili A, Shilatifard A, Buratowski S, Greenblatt JF (2002) RNA polymerase II elongation factors of Saccharomyces cerevisiae: a targeted proteomics approach. Mol Cell Biol 22:6979–6992
Mayer A, Lidschreiber M, Siebert M, Leike K, Söding J, Cramer P (2010) Uniform transitions of the general RNA polymerase II transcription complex. Nat Struct Mol Biol 17:1272–1278
Cojocaru M, Jeronimo C, Forget D, Bouchard A, Bergeron D, Côte P, Poirier GG, Greenblatt J, Coulombe B (2008) Genomic location of the human RNA polymerase II general machinery: evidence for a role of TFIIF and Rpb7 at both early and late stages of transcription. Biochem J 409:139–147
Rani PG, Ranish JA, Hahn S (2004) RNA polymerase II (Pol II)-TFIIF and Pol II-mediator complexes: the major stable Pol II complexes and their activity in transcription initiation and reinitiation. Mol Cell Biol 24:1709–1720
Izban MG, Luse DS (1992) Factor-stimulated RNA polymerase II transcribes at physiological elongation rates on naked DNA but very poorly on chromatin templates. J Biol Chem 267:13647–13655
Renner DB, Yamaguchi Y, Wada T, Handa H, Price DH (2001) A highly purified RNA polymerase II elongation control system. J Biol Chem 276:42601–42609
Zhang C, Yan H, Burton ZF (2003) Combinatorial control of human RNA polymerase II (RNAP II) pausing and transcript cleavage by transcription factor IIF, hepatitis delta antigen, and stimulatory factor II. J Biol Chem 278:50101–50111
Schweikhard V, Meng C, Murakami K, Kaplan CD, Kornberg RD, Block SM (2014) Transcription factors TFIIF and TFIIS promote transcript elongation by RNA polymerase II by synergistic and independent mechanisms. Proc Natl Acad Sci U S A 111:6642–6647
Zhang C, Burton ZF (2004) Transcription factors IIF and IIS and nucleoside triphosphate substrates as dynamic probes of the human RNA polymerase II mechanism. J Mol Biol 342:1085–1099
Zhang C, Zobeck KL, Burton ZF (2005) Human RNA polymerase II elongation in slow motion: role of the TFIIF RAP74 alpha1 helix in nucleoside triphosphate-driven translocation. Mol Cell Biol 25:3583–3595
Lisica A, Engel C, Jahnel M, Roldán É, Galburt EA, Cramer P, Grill SW (2016) Mechanisms of backtrack recovery by RNA polymerases I and II. Proc Natl Acad Sci U S A 113:2946–2951
Tafur L, Sadian Y, Hanske J, Wetzel R, Weis F, Müller CW (2019) The cryo-EM structure of a 12-subunit variant of RNA polymerase I reveals dissociation of the A49-A34.5 heterodimer and rearrangement of subunit A12.2. eLife 8:e43204
Kornberg RD (1974) Chromatin structure: a repeating unit of histones and DNA. Science 184:868–871
Chang CH, Luse DS (1997) The H3/H4 tetramer blocks transcript elongation by RNA polymerase II in vitro. J Biol Chem 272:23427–23434
Izban MG, Luse DS (1991) Transcription on nucleosomal templates by RNA polymerase II in vitro: inhibition of elongation with enhancement of sequence-specific pausing. Genes Dev 5:683–696
Gaykalova DA, Kulaeva OI, Volokh O, Shaytan AK, Hsieh F-K, Kirpichnikov MP, Sokolova OS, Studitsky VM (2015) Structural analysis of nucleosomal barrier to transcription. Proc Natl Acad Sci U S A 112:E5787–E5795
Kujirai T, Ehara H, Fujino Y, Shirouzu M, Sekine S-I, Kurumizaka H (2018) Structural basis of the nucleosome transition during RNA polymerase II passage. Science 362:595–598
Dangkulwanich M, Ishibashi T, Bintu L, Bustamante C (2014) Molecular mechanisms of transcription through single-molecule experiments. Chem Rev 114:3203–3223
Kulaeva OI, Hsieh F-K, Chang H-W, Luse DS, Studitsky VM (2013) Mechanism of transcription through a nucleosome by RNA polymerase II. Biochim Biophys Acta 1829:76–83
Bintu L, Kopaczynska M, Hodges C, Lubkowska L, Kashlev M, Bustamante C (2011) The elongation rate of RNA polymerase determines the fate of transcribed nucleosomes. Nat Struct Mol Biol 18:1394–1399
Luse DS, Studitsky VM (2011) The mechanism of nucleosome traversal by RNA polymerase II: roles for template uncoiling and transcript elongation factors. RNA Biol 8:581–585
Luse DS, Spangler LC, Újvári A (2011) Efficient and rapid nucleosome traversal by RNA polymerase II depends on a combination of transcript elongation factors. J Biol Chem 286:6040–6048
Crickard JB, Lee J, Lee T-H, Reese JC (2017) The elongation factor Spt4/5 regulates RNA polymerase II transcription through the nucleosome. Nucleic Acids Res 45:6362–6374
Ehara H, Kujirai T, Fujino Y, Shirouzu M, Kurumizaka H, Sekine S-I (2019) Structural insight into nucleosome transcription by RNA polymerase II with elongation factors. Science 363:744–747
Belotserkovskaya R, Oh S, Bondarenko VA, Orphanides G, Studitsky VM, Reinberg D (2003) FACT facilitates transcription-dependent nucleosome alteration. Science 301:1090–1093
Hsieh F-K, Kulaeva OI, Patel SS, Dyer PN, Luger K, Reinberg D, Studitsky VM (2013) Histone chaperone FACT action during transcription through chromatin by RNA polymerase II. Proc Natl Acad Sci U S A 110:7654–7659
Kim J, Guermah M, Roeder RG (2010) The human PAF1 complex acts in chromatin transcription elongation both independently and cooperatively with SII/TFIIS. Cell 140:491–503
Birch JL, Tan BC-M, Panov KI, Panova TB, Andersen JS, Owen-Hughes TA, Russell J, Lee S-C, Zomerdijk JCBM (2009) FACT facilitates chromatin transcription by RNA polymerases I and III. EMBO J 28:854–865
Schneider DA, French SL, Osheim YN, Bailey AO, Vu L, Dodd J, Yates JR, Beyer AL, Nomura M (2006) RNA polymerase II elongation factors Spt4p and Spt5p play roles in transcription elongation by RNA polymerase I and rRNA processing. Proc Natl Acad Sci U S A 103:12707–12712
Zhang Y, Sikes ML, Beyer AL, Schneider DA (2009) The Paf1 complex is required for efficient transcription elongation by RNA polymerase I. Proc Natl Acad Sci U S A 106:2153–2158
Zhang Y, Smith AD, Renfrow MB, Schneider DA (2010) The RNA polymerase-associated factor 1 complex (Paf1C) directly increases the elongation rate of RNA polymerase I and is required for efficient regulation of rRNA synthesis. J Biol Chem 285:14152–14159
Acknowledgments
We thank all the members of the department of Biochemistry III for constant support and discussion. This work was funded by the Deutsche Forschungsgemeinschaft (DFG) in the context of the SFB960. P. E. M. was partly supported by a fellowship of the German National Academic Foundation.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2022 The Author(s)
About this protocol
Cite this protocol
Merkl, P.E. et al. (2022). Specialization of RNA Polymerase I in Comparison to Other Nuclear RNA Polymerases of Saccharomyces cerevisiae . In: Entian, KD. (eds) Ribosome Biogenesis. Methods in Molecular Biology, vol 2533. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2501-9_4
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2501-9_4
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2500-2
Online ISBN: 978-1-0716-2501-9
eBook Packages: Springer Protocols