Abstract
Cellular RNAs, both coding and noncoding, contain several chemical modifications. Both ribose sugars and nitrogenous bases are targeted for these chemical additions. These modifications are believed to expand the topological potential of RNA molecules by bringing chemical diversity to otherwise limited repertoire. Here, using ribosomal RNA of yeast as an example, a detailed protocol for systematically mapping various chemical modifications to a single nucleotide resolution by a combination of Mung bean nuclease protection assay and RP-HPLC is provided. Molar levels are also calculated for each modification using their UV (254 nm) molar response factors that can be used for determining the amount of modifications at different residues in other RNA molecules. The chemical nature, their precise location and quantification of modifications will facilitate understanding the precise role of these chemical modifications in cellular physiology.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
1 Introduction
Chemical modification of biomolecules is an impressive way of nature to expand their functional diversity. Like DNA and proteins, RNA molecules also undergo a variety of chemical modifications that allow them to enrich their topological potential and perform a variety of functions that go beyond their capacity to encode proteins [1, 2].
Around 163 different chemical modifications of RNA has been cataloged [3]. Both ribose sugar and the heterocyclic ring of the nucleobases are targeted for these chemical additions. Although the presence of these modifications was first acknowledged by some elegant studies in the middle of last century, it is only very recently that we have started compiling modification profiles for the cellular RNAs of model organisms, and explore the functions of few of these modifications [4]. This time lag has been primarily due to unavailability of advanced technology in studying these modifications, which has been revolutionized in last two decades by introduction of both high-performance liquid chromatography (HPLC) , mass spectrometry (MS) , and RNA sequencing (RNA Seq ) [5,6,7,8].
Ribosomes are highly conserved ribonucleoprotein complexes that synthesize cellular proteins. Eukaryotic ribosomes comprise of 4 ribosomal RNAs (rRNAs) and around 80 ribosomal proteins (r-proteins ). Both enzymatic reactions of ribosomes, namely, peptidyl transfer and peptidyl hydrolysis are performed by rRNA [9]. In eukaryotes including Saccharomyces cerevisiae , a functional ribosome contains two asymmetric subunits, a small 40S and a large 60S . A small subunit (SSU) of the ribosome (the 40S ) in yeast contains a single 18S rRNA of 1.8 kb together with 33 ribosomal proteins [10]. The 60S or the large subunit (LSU) of the yeast ribosome contains three rRNAs ; 25S, 5.8S and 5S, and 46 ribosomal proteins . The 40S decodes the genetic information carried by mRNA , whereas the 60S catalyzes the joining of amino acids [11, 12]. Ribosomes have always been seen as homogeneous, constitutive protein-synthesizing machine, lacking any significant contribution in regulating gene expression . Typically, the efficiency of translation is suggested to be determined either by features intrinsic to the mRNAs or assisted by protein or RNA adaptors (translational factors). However, in contrast to this view, several recent studies have underscored the undeniable roles of ribosomes in gene regulation [13,14,15,16,17]. Emerging data have buttressed the notion that the ribosome population in cells is heterogeneous by virtue of its different components, including ribosomal proteins , rRNAs and their chemical modifications . It becomes more and more evident that this ribosomal heterogeneity provides a further regulatory level to the translation .
Ribosomal RNA contains three types of chemical modifications , 2′-O methylation of ribose sugars (Nm), base isomerization (pseudouridylation (Ψ)), and base modifications (methylation (mN) and acetylation (acN) , aminocarboxypropylation (acpN)) [1, 2]. Methylated ribose sugars and pseudouridines represent the majority of rRNA modifications [18]. A methylated sugar is generated by the addition of one methyl group at the 2′-OH position of the ribose on the nucleoside and is independent of the nature of the base. Pseudouridylation results from uridine isomerization, involving a 180° rotation around the N3–C6 axis [19].
Box C/D snoRNPs (small nucleolar ribonucleoproteins) catalyzes site-directed methylation at the 2′-OH position on the sugar of the targeted nucleotide, whereas the box H/ACA snoRNPs isomerize targeted uridine to pseudouridine [20]. Remarkably, the substrate specificity for both ribose methylation and pseudouridylation is dictated by the RNA component of these snoRNP complexes [21, 22]. Here the complementary sequences in the guide RNA base pair with target RNAs to decide the nucleotide for modification [21, 22]. The catalytic activities are provided by the methyltransferase (Nop1 or Fibrillarin ) or the pseudouridine synthase (Cbf5), respectively [23, 24]. In contrast, majority of base modifications are catalyzed by “protein only” enzymes with only exception being rRNA cytosine acetylation (ac4C) [25].
To analyze and explore the significance of these modifications in ribosome biogenesis and ribosome function, a comprehensive analysis of their chemical nature and precise location on the ribosome is central. In this chapter, a detailed protocol for mapping these modifications on to the ribosomal RNA by mung bean nuclease (MBN) assay and RP-HPLC (reversed phase high-performance liquid chromatography) is provided [26].
MBN is a single strand specific nuclease , and can be used to isolate specific fragments of RNA by utilizing synthetic complementary DNA. These RNA fragments can then be subjected to RP-HPLC or LC-MS /MS analysis to identify the modification or modifications they harbor (Fig. 1) [26, 27]. This method also allows to achieve a single nucleotide resolution for mapping these modifications by isolating overlapping fragments [28, 29].
Reversed phase high-performance liquid chromatography (RP-HPLC ) analysis. (a) Sucrose gradient sedimentation profile of yeast total RNA separated on 5% to 25% sucrose gradient. Fractions corresponding to tRNA (predominantly)), 18S rRNA , and 25S rRNA were collected and the rRNA was isolated using 95% ethanol precipitation at −80 °C. (b) A representative 1.5% Agarose gel showing the rRNA recovered after ethanol precipitation. (c) RP-HPLC chromatogram of different commercially available nucleosides, showing peak corresponding to major nucleosides (adapted from [26])
Although MBN assay and LC-MS /MS are work intensive, these are so far the only methods for a direct analysis of these chemical modifications . All other RNA sequencing based methods relies either on antibodies, which most of the time have issues with cross-reactivities or on the chemical reactivities of the modified residues, which are not very specific and are similar among many different modifications [30, 31]. The major limitation of the LC-MS /MS , on the other hand, is that it cannot be applied in a high throughput manner to reveal both the modification and the sequence context as in case of RNA Seq based methods. Oxford nanopore sequencing appears to be an ideal method of choice for the direct transcriptome-wide sequencing of these modifications [32].
2 Materials
2.1 Isolation of Intact 18S and 25S rRNA
-
1.
Acid phenol–chloroform–isoamyl alcohol, pH 4.5 (25:24:1) (InvitrogenTM, Catalog number: AM9720).
-
2.
TEN buffer.
-
100 mM Tris–HCl pH 7.8.
-
1 mM EDTA.
-
100 mM NaCl in nuclease-free water.
-
-
3.
5% and 25% sucrose solutions (w/v) in TEN buffer.
2.2 Mung Bean Nuclease Protection Assay
-
1.
Hybridization buffer.
-
250 mM HEPES pH 7.0.
-
500 mM KCl in nuclease-free water.
-
-
2.
Mung bean nuclease (New England BioLabs (NEB), Catalog number: M0250L).
-
3.
RNAse A DNase and protease free (Thermo ScientificTM, Catalog number: EN0531).
-
4.
8M LiCl.
-
5.
3M sodium acetate (NaAc), pH 5.3.
-
6.
GlycoBlueTM (InvitrogenTM, Catalog number: AM9515).
-
7.
UltraPureTM Formamide (InvitrogenTM, Catalog number: 15515026).
-
8.
2× RNA loading buffer (Thermo ScientificTM, Catalog number: R0641).
-
9.
Novex™ TBE-Urea Gels, 10% (InvitrogenTM, Catalog number: EC68755BOX).
2.3 Quantitative Reversed Phase High-Performance Liquid Chromatography (qRP-HPLC)
-
1.
Millipore Water (18 MΩ·cm resistivity at 25°C).
-
2.
Nuclease P1 from Penicillium citrinum (Sigma-Aldrich, Catalog number: N8630) (200 units per mL in 30 mM sodium acetate, pH 5.4).
-
3.
Phosphatase, Alkaline from Escherichia coli (Sigma-Aldrich, Catalog number: P4252).
-
4.
10mM ZnSO4.
-
5.
0.5M Tris–HCl, pH 8.3.
-
6.
HPLC grade Methanol (AppliChem, Catalog number: 361091).
-
7.
HPLC Grade Acetonitrile (AppliChem, Catalog number: 221881).
-
8.
Buffer A (10 mM of NH4H2PO4, 2.5% of methanol, pH 5.3).
-
9.
Buffer B (10 mM of NH4H2PO4, 20% of methanol, pH 5.1).
-
10.
Buffer C (66% acetonitrile).
-
11.
All canonical and modified nucleosides standard—Carbosynth.
3 Methods
3.1 Isolation of Intact 18S and 25S rRNA
-
1.
Harvest yeast cells at OD 600 0.8 (see Note 1 ).
-
2.
Extract the total RNA with hot acidic phenol exactly as described before [33].
-
3.
To isolate intact 18S and 25S rRNA , layer 500 μg of total RNA onto a freshly prepared 5–25% sucrose gradient in TEN buffer (see Note 2 ). Prepare the sucrose gradients using Gradient Master 107 (Biocomp).
-
4.
Centrifuge the samples at 67,000 × g for 25 h at 4 °C in an SW40 rotor in an L-70 Beckman ultracentrifuge.
-
5.
Fractionate the gradients by using a Teledyne Isco density gradient fraction collector. Collect 18S and 25S rRNA fractions in separate tubes and to each add 2.5 × volume of chilled ethanol. Incubate them overnight in a −20 freezer.
-
6.
Next day, centrifuge the rRNA samples at 16,000 × g for 30 min at 4 °C, air-dry the pellet, and resuspend it in nuclease-free water.
-
7.
For the quantification of modified residues, it is very important to have intact rRNA . Since rRNA are abundant species, the quality (integrity) of purified 18S and 25S rRNA could be easily assesed by using 1.5% agarose gel stained with ethidium bromide.
3.2 Mung Bean Nuclease Protection assay (MBN Assay )
3.2.1 Hybridization
-
1.
Mix the following components in a 1.5 mL Eppendorf tubes. DNA oligos listed in Table 4 have been used for mapping yeast rRNA modifications [26].
Components
Amount
18S or 25S rRNA (see Note 3)
100 pmoles
DNA oligo (Table 4)
1000 pmoles
-DMSO
3 μL
Nuclease-free water
Up to 100 μL
Hybridization buffer
30 μL
-
2.
Heat the mixture to 90 °C for 5 min, and let it cool down to 45 °C on heating block (approximately 1.5 h). When the temperature drops down to 45 °C, move to the next step.
3.2.2 Mung Bean Nuclease Digestion
-
1.
Mung bean nuclease digestion are carried out by mixing following components.
Components
Amount (μL)
RNA oligo mix
130
10× MBN buffer
20
RNAse A (0.05 μg/μL)
14
Mung bean nuclease
10
Nuclease-free water
26
-
2.
Incubate the reaction at 35 °C for 1 h.
3.2.3 Purification of the rRNA Fragments
-
1.
After the MBN digestion, purify the rRNA fragment using phenol–chloroform extraction. Add 250 μL of acid phenol–chloroform–isoamyl alcohol, pH 4.5 (25:24:1) mixture to the 200 μL reaction mixture and vortex it for 1 min.
-
2.
Centrifuge the Eppendorf tubes at 4000 × g for 5 min at 4 °C.
-
3.
Precipitate the rRNA fragments overnight at −20 °C by adding 350 μL of 8 M LiCl and 1 mL ethanol in 2 mL Eppendorf tubes.
-
4.
Next day, centrifuge the tube at 16,000 × g for 30 min at 4 °C, air-dry the pellet and resuspended the rRNA fragments in 30 μL nuclease-free water–formamide (1:2).
-
5.
Add 30 μL of 2× RNA loading buffer to the tubes and heat the samples to 70 °C for 10 min.
-
6.
Load the samples on a Novex™ TBE-Urea Gels, 10% and run the gels as per manufacturer’s instructions. Use DNA oligo and DNA oligo + total RNA as marker (see Note 5 ).
-
7.
Slice out the gel containing the protected rRNA fragment using a fresh blade (see Note 5 ).
-
8.
Wash gel slices with Millipore water 2–3 times with a pipette, do not crush the gel.
-
9.
Add 450 μL nuclease-free water and 50 μL of 3 M NaAc pH 5.3 (final conc. = 0.3 M NaAc).
-
10.
Elute the rRNA fragment from the gel slices by rotating on the rotor overnight at 4 °C (see Note 6 ).
-
11.
Transfer the eluate into a 2 mL Eppendorf tube (Keep the gel slices in the old Eppendorf tube).
-
12.
Add 1.25 mL (2.5× elution), 100% EtOH, and 0.5 μL GlycoBlue™ to the eluate, and incubate the mix at −80 °C for at least 12 h for efficient precipitation.
-
13.
Next day, centrifuge the tube at 16,000 × g for 30 min at 4 °C, air-dry the pellet and resuspend in 45 μL of nuclease-free water. Store the samples at −20 °C.
-
14.
Repeat the elution step. We get a better yield with two elution.
-
15.
Quantify the rRNA fragment using a nanodrop.
3.2.4 Mapping of Chemical Modifications to a Single Nucleotide Resolution
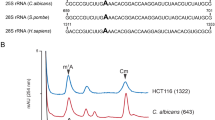
As illustrated in Fig. 2, MBN assay could be easily used to achieve single nucleotide resolution by designing overlapping fragment around the modifications (see Note 4).
Mung bean nuclease protection assay . (a) Schematic illustration of mung bean nuclease protection assay (MBN ). (b) Representative Urea PAGE gel for the MBN assay showing an intact protected fragment retrieved after the mung bean nuclease digestion. The synthetic antisense oligonucleotide used for the protection is used as a marker (see Note 5 ) (adapted from [26])
3.3 Quantitative Reversed Phase High-Performance Liquid Chromatography (qRP-HPLC)
3.3.1 Preparation of Nucleosides
Nucleosides for RP-HPLC are prepared as described by Gehrke and Kuo, and adapted for rRNA as described previously [7, 26].
-
1.
Heat the rRNA fragments isolated after MBN assay as described above to 90 °C for 2 min on a heating block followed by rapid cooling on ice.
-
2.
Add 5 μL of 10 mM ZnSO4 and 10 μL of P1 nuclease. Incubate the samples for 16 h at 37 °C.
-
3.
After 16 h, add 10 μL of 0.5 M Tris–HCl buffer, pH 8.3, and 10 μL of bacterial alkaline phosphatase and incubate the tubes at 37 °C for 2 h.
-
4.
Next, before loading on the HPLC column clarify the nucleosides hydrolysates by centrifugation at 16, 000 × g.
3.3.2 RP-HPLC
-
1.
Load the nucleosides hydrolysates on Supelcosil LC-18-S HPLC column (25 cm × 4.6 mm, 5 mm) equipped with a precolumn (4.6 × 20 mm) at 30 °C on an Agilent 1200 HPLC system.
-
2.
Use the following elution protocol (Table 1) described by Gehrke and Kuo for resolving majority of canonical and modified nucleosides in the rRNA (Fig. 1) [7]. For some modifications like m3U protocol has to be adjusted for better separation. In contrast to gradient elution for rest of the modified nucleosides, the elution conditions for m3U are changed to an isocratic mode using 50% buffer A and 50% buffer B.
3.3.3 Quantification of the Modified Nucleosides
Approximate modification levels for each ribose and base modifications can be calculated from HPLC peak areas (UV254nm absorption). For the quantification, the peak areas are divided by the standard molar response factors. Table 2 contains the UV254nm molar response factors calculated in our previous study. Similarly, mole % (Seq) for yeast (S. cerevisiae ) are calculated assuming that 18S rRNA (1800 nts) contains 473 unmodified adenosines (A), 494 unmodified uridines (U), 453 unmodified guanosines (G), and 343 unmodified cytidines (C), and similarly for 25S rRNA (3396 nts) assuming that it contains 885 unmodified adenosines (A), 828 unmodified uridines (U), 956 unmodified guanosines (G), and 653 unmodified cytidines (C) Table 3. Likewise, for MBN protected fragments residues per moles or extent of modifications for each residue can be established using the sequence of the isolated fragment and its comparison to the residues/moles calculated from the 18S rRNA or 25S rRNA (see Note 7 ).
4 Notes
-
1.
Make sure that the yeast cells are grown for at least 6–7 generations.
-
2.
Always prepare fresh TEN buffer and sucrose solutions. For isolation of rRNAs from higher eukaryotes including humans, the sucrose gradient should be adjusted—the one used here is optimized for both budding and fission yeast .
-
3.
Instead of purified 18S and 25S rRNA , MBN assay can also be performed using total RNA , although the yield and efficiency was significantly higher with purified 18S and 25S rRNA .
-
4.
Since several modifications impact the hydrogen bonding, isolation of fragment using MBN assay from highly modified regions of the rRNA is sometime challenging. The best oligo set may have to be empirically determined by altering the Tm, either by increasing the length or sequence.
-
5.
After MBN assay , fragments must be PAGE purified using appropriate marker—(DNA oligo used for the reaction can also be used). Often we see the protected fragment running a little higher than the expected size.
-
6.
Instead of passive elution of the rRNA fragments, one can also use electro elution using Model 422 Electro-Eluter (BioRad).
-
7.
MBN assay in combination with RP-HPLC is a valuable tool for both qualitative and quantitative of relatively abundant RNA species. For low abundant RNAs it is advised to use more sensitive methods like LC-MS /MS or upcoming Oxford nanopore base sequencing.
-
8.
Molar response factors given here (Table 2) are calculated for NH4H2PO4 buffer system. NH4H2PO4 has a low UV absorption and higher solubility as compared to the potassium or sodium salts [7]. If other buffer system is used, the molar response factors should be recalculated.
References
Sharma S, Lafontaine DLJ (2015) ‘View from a bridge’: a new perspective on eukaryotic rRNA base modification. Trends Biochem Sci 40:560–575
Sloan KE, Warda AS, Sharma S, Entian K-D, Lafontaine DLJ, Bohnsack MT (2016) Tuning the ribosome: the influence of rRNA modification on eukaryotic ribosome biogenesis and function. RNA Biol 14(9):1138–1152. https://doi.org/10.1080/15476286.2016.1259781
Boccaletto P, Machnicka MA, Purta E, Piatkowski P, Baginski B, Wirecki TK, de Crécy-Lagard V, Ross R, Limbach PA, Kotter A et al (2018) MODOMICS: a database of RNA modification pathways. 2017 update. NAR 46:D303–D307
Klootwijk J, Planta RJ (1973) Analysis of the methylation sites in yeast ribosomal RNA. Eur J Biochem 39(2):325–333. https://doi.org/10.1111/j.1432-1033.1973.tb03130.x
Banoub JH, Limbach PA (2009) Mass spectrometry of nucleosides and nucleic acids. CRC Press
Gehrke CW, Kuo KCT (eds) (1989) Analytical methods for major and modified nucleosides - HPLC, GC, MS, NMR, UV and FT-IR. Elsevier
Gehrke CW, Kuo KC (1990) Chapter 1 Ribonucleoside analysis by reversed-phase high performance liquid chromatography. In: Chromatography and modification of nucleosides: analytical methods for major and modified nucleosides, HPLC, GC, MS, NMR, UV, and FT-IR, Journal of chromatography library, vol 45. Elsevier, pp A3–A71
Motorin Y, Helm M (2019) Methods for RNA modification mapping using deep sequencing: established and new emerging technologies. Genes 10:35
Woolford JL, Baserga SJ (2013) Ribosome biogenesis in the yeast Saccharomyces cerevisiae. Genetics 195:643–681
Henras AK, Plisson-Chastang C, O'Donohue M-F, Chakraborty A, Gleizes P-E (2015) An overview of pre-ribosomal RNA processing in eukaryotes. WIREs RNA 6:225–242
Ramakrishnan V (2002) Ribosome structure and the mechanism of translation. Cell 108:557–572
Lilley DM (2001) The ribosome functions as a ribozyme. Chembiochem 2:31–35
Shi Z, Fujii K, Kovary KM, Genuth NR, Röst HL, Teruel MN, Barna M (2017) Heterogeneous ribosomes preferentially translate distinct subpools of mRNAs genome-wide. Mol Cell 67:71–83.e7
Simsek D, Tiu GC, Flynn RA, Byeon GW, Leppek K, Xu AF, Chang HY, Barna M (2017) The mammalian Ribo-interactome reveals ribosome functional diversity and heterogeneity. Cell 169:1051–1057.e18
Xue S, Barna M (2012) Specialized ribosomes: a new frontier in gene regulation and organismal biology. Nat Rev Mol Cell Biol 13:355–369
Schosserer M, Minois N, Angerer TB, Amring M, Dellago H, Harreither E, Calle-Perez A, Pircher A, Gerstl MP, Pfeifenberger S et al (2015) Methylation of ribosomal RNA by NSUN5 is a conserved mechanism modulating organismal lifespan. Nat Commun 6:6158
Sharma S, Hartmann JD, Watzinger P, Klepper A, Peifer C, Kötter P, Lafontaine DLJ, Entian K-D (2018) A single N1-methyladenosine on the large ribosomal subunit rRNA impacts locally its structure and the translation of key metabolic enzymes. Sci Rep 8:11904
Piekna-Przybylska D, Decatur WA, Fournier MJ (2008) The 3D rRNA modification maps database: with interactive tools for ribosome analysis. NAR 36:D178–D183
Gray MCMW (2000) Pseudouridine in RNA: what, where, how, and why. IUBMB Life 49:341–351
Kiss T (2002) Small nucleolar RNAs: an abundant group of noncoding RNAs with diverse cellular functions. Cell 109:145–148
Kiss-László Z, Henry Y, Bachellerie JP, Caizergues-Ferrer M, KISS, T. (1996) Site-specific ribose methylation of preribosomal RNA: a novel function for small nucleolar RNAs. Cell 85:1077–1088
Ganot P, Bortolin ML, KISS, T. (1997) Site-specific pseudouridine formation in preribosomal RNA is guided by small nucleolar RNAs. Cell 89:799–809
Zebarjadian Y, King T, Fournier MJ, Clarke L, Carbon J (1999) Point mutations in yeast CBF5 can abolish in vivo pseudouridylation of rRNA. Mol Cell Biol 19:7461–7472
Tollervey D, Lehtonen H, Jansen R, Kern H, Hurt EC (1993) Temperature-sensitive mutations demonstrate roles for yeast fibrillarin in pre-rRNA processing, pre-rRNA methylation, and ribosome assembly. Cell 72:443–457
Sharma S, Yang J, van Nues R, Watzinger P, Kötter P, Lafontaine DLJ, Granneman S, Entian K-D (2017) Specialized box C/D snoRNPs act as antisense guides to target RNA base acetylation. PLoS Genet 13:e1006804
Yang J, Sharma S, Watzinger P, Hartmann JD, Kötter P, Entian K-D (2016) Mapping of complete set of ribose and base modifications of yeast rRNA by RP-HPLC and mung bean nuclease assay. PLoS One 11:e0168873
Buchhaupt M, Sharma S, Kellner S, Oswald S (2014) Partial methylation at Am100 in 18S rRNA of baker’s yeast reveals ribosome heterogeneity on the level of eukaryotic rRNA modification. PLoS One 9(2):e89640. https://doi.org/10.1016/j.ab.2006.11.001
Sharma S, Yang J, Watzinger P, Kötter P, Entian K-D (2013) Yeast Nop2 and Rcm1 methylate C2870 and C2278 of the 25S rRNA, respectively. Nucleic Acids Res 41:9062–9076
Sharma S, Langhendries J-L, Watzinger P, Kötter P, Entian K-D, Lafontaine DLJ (2015) Yeast Kre33 and human NAT10 are conserved 18S rRNA cytosine acetyltransferases that modify tRNAs assisted by the adaptor Tan1/THUMPD1. Nucleic Acids Res 43:2242–2258
Helm M, Lyko F, Motorin Y (2019) Limited antibody specificity compromises epitranscriptomic analyses. Nat Commun 10:5669
Helm M, Motorin Y (2017) Detecting RNA modifications in the epitranscriptome: predict and validate. Nat Publ Group 18:275–291
Smith AM, Jain M, Mulroney L, Garalde DR, Akeson M (2019) Reading canonical and modified nucleobases in 16S ribosomal RNA using nanopore native RNA sequencing. PLoS One 14:e0216709
Collart MA, Oliviero S (2001) Preparation of yeast RNA. Curr Protoc Mol Biol Chapter 13:Unit13.12
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2022 The Author(s)
About this protocol
Cite this protocol
Yang, J., Watzinger, P., Sharma, S. (2022). Mapping of the Chemical Modifications of rRNAs. In: Entian, KD. (eds) Ribosome Biogenesis. Methods in Molecular Biology, vol 2533. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2501-9_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2501-9_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2500-2
Online ISBN: 978-1-0716-2501-9
eBook Packages: Springer Protocols