Abstract
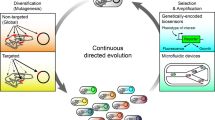
Evolutionary engineering of microbes provides a powerful tool for untargeted optimization of (engineered) cell factories and identification of genetic targets for further research. Directed evolution is an intrinsically time-intensive effort, and automated methods can significantly reduce manual labor. Here, design considerations for various evolutionary engineering methods are described, and generic workflows for batch-, chemostat-, and accelerostat-based evolution in automated bioreactors are provided. These methods can be used to evolve yeast cultures for >1000 generations and are designed to require minimal manual intervention.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Mans R, Daran J-MG, Pronk JT (2018) Under pressure: evolutionary engineering of yeast strains for improved performance in fuels and chemicals production. Curr Opin Biotechnol 50:47–56
Ho P, Swinnen S, Duitama J et al (2017) The sole introduction of two single-point mutations establishes glycerol utilization in Saccharomyces cerevisiae CEN.PK derivatives. Biotechnol Biofuels 10(1):10
Kildegaard KR, Hallström BM, Blicher TH et al (2014) Evolution reveals a glutathione-dependent mechanism of 3-hydroxypropionic acid tolerance. Metab Eng 26:57–66
Caspeta L, Chen Y, Ghiaci P et al (2014) Altered sterol composition renders yeast thermotolerant. Science 346(6205):75–78
Gonzalez-Ramos D, De Vries ARG, Grijseels SS et al (2016) A new laboratory evolution approach to select for constitutive acetic acid tolerance in Saccharomyces cerevisiae and identification of causal mutations. Biotechnol Biofuels 9(1):173
Juergens H, Niemeijer M, Jennings-Antipov LD et al (2018) Evaluation of a novel cloud-based software platform for structured experiment design and linked data analytics. Sci Data 5:180195
Zhou H, Cheng JS, Wang BL et al (2012) Xylose isomerase overexpression along with engineering of the pentose phosphate pathway and evolutionary engineering enable rapid xylose utilization and ethanol production by Saccharomyces cerevisiae. Metab Eng 14(6):611–622
Basso TO, de Kok S, Dario M et al (2011) Engineering topology and kinetics of sucrose metabolism in Saccharomyces cerevisiae for improved ethanol yield. Metab Eng 13(6):694–703
Kuyper M, Toirkens MJ, Diderich JA et al (2005) Evolutionary engineering of mixed-sugar utilization by a xylose-fermenting Saccharomyces cerevisiae strain. FEMS Yeast Res 5(10):925–934
Bracher JM, de Hulster E, Koster CC et al (2017) Laboratory evolution of a biotin-requiring Saccharomyces cerevisiae strain for full biotin prototrophy and identification of causal mutations. Appl Environ Microbiol 83(16):e00892–e00817
Marques WL, van der Woude LN, Luttik MA et al (2018) Laboratory evolution and physiological analysis of Saccharomyces cerevisiae strains dependent on sucrose uptake via the Phaseolus vulgaris Suf1 transporter. Yeast 35(12):639–652
Acknowledgments
The contribution of CM was funded by an Advanced Grant of the European Research Council (grant # 694633 to Jack T. Pronk).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic Supplementary Material
Rights and permissions
Copyright information
© 2022 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
de Hulster, E., Mooiman, C., Timmermans, R., Mans, R. (2022). Automated Evolutionary Engineering of Yeasts. In: Mapelli, V., Bettiga, M. (eds) Yeast Metabolic Engineering. Methods in Molecular Biology, vol 2513. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2399-2_15
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2399-2_15
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2398-5
Online ISBN: 978-1-0716-2399-2
eBook Packages: Springer Protocols