Abstract
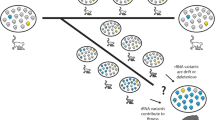
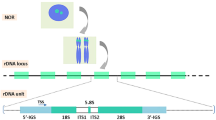
Although commonly regarded as nearly identical in sequence, 45S ribosomal RNA genes represent a massive source of genetic variation at different levels. Unfortunately, due to their repetitive nature and the difficulty to assemble their multiple copies in tandem, these important genomic elements remain largely unexplored in sequencing projects. Here, I describe how to exploit next generation sequencing data to estimate their copy number in an organism and detect true polymorphic sites within and among individuals. Furthermore, for species that carry multiple 45S ribosomal RNA gene clusters, I show how to make use of experimental populations to assign some of these variants to their cluster of origin.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Moss T, Langlois F, Gagnon-Kugler T et al (2007) A housekeeper with power of attorney: the rRNA genes in ribosome biogenesis. Cell Mol Life Sci 64:29–49
Layat E, Sáez-Vásquez J, Tourmente S (2012) Regulation of pol I-transcribed 45S rDNA and pol III-transcribed 5S rDNA in Arabidopsis. Plant Cell Physiol 53:267–276
Ritossa FM, Spiegelman S (1965) Localization of DNA complementary to ribosomal RNA in the nucleolus organizer region of drosophila melanogaster. Proc Natl Acad Sci U S A 53:737–745
Wallace H, Birnstiel ML (1966) Ribosomal cistrons and the nucleolar organizer. Biochim Biophys Acta 114:296–310
Michael TP, Jupe F, Bemm F et al (2018) High contiguity Arabidopsis thaliana genome assembly with a single nanopore flow cell. Nat Commun 9:541
Sims J, Sestini G, Elgert C et al (2021) Sequencing of the Arabidopsis NOR2 reveals its distinct organization and tissue-specific rRNA ribosomal variants. Nat Commun 12:387
Guo Y, Li J, Li C-I et al (2012) The effect of strand bias in Illumina short-read sequencing data. BMC Genomics 13:666
Guo Y, Cai Q, Samuels DC et al (2012) The use of next generation sequencing technology to study the effect of radiation therapy on mitochondrial DNA mutation. Mutat Res 744:154–160
Rabanal FA, Mandáková T, Soto-Jiménez LM et al (2017) Epistatic and allelic interactions control expression of ribosomal RNA gene clusters in Arabidopsis thaliana. Genome Biol 18:75
Copenhaver GP, Pikaard CS (1996) RFLP and physical mapping with an rDNA-specific endonuclease reveals that nucleolus organizer regions of Arabidopsis thaliana adjoin the telomeres on chromosomes 2 and 4. Plant J 9:259–272
Copenhaver GP, Browne WE, Preuss D (1998) Assaying genome-wide recombination and centromere functions with Arabidopsis tetrads. Proc Natl Acad Sci 95:247–252
Anaconda Software Distribution (2020) Anaconda Documentation. Anaconda Inc. https://docs.anaconda.com/
R Core Team (2020) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Grüning B, Dale R, Sjödin A et al (2018) Bioconda: sustainable and comprehensive software distribution for the life sciences. Nat Methods 15:475–476
Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796–815
Lamesch P, Berardini TZ, Li D et al (2012) The Arabidopsis information resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res 40:D1202–D1210
Rabanal FA, Nizhynska V, Mandáková T et al (2017) Unstable Inheritance of 45S rRNA Genes in Arabidopsis thaliana. G3 (Bethesda) 7:1201–1209
Alonso-Blanco C, El-Assal SE, Coupland G et al (1998) Analysis of natural allelic variation at flowering time loci in the Landsberg erecta and Cape Verde Islands ecotypes of Arabidopsis thaliana. Genetics 149:749–764
Pontvianne F, Abou-Ellail M, Douet J et al (2010) Nucleolin is required for DNA methylation state and the expression of rRNA gene variants in Arabidopsis thaliana. PLoS Genet 6:e1001225
Abou-Ellail M, Cooke R, Sáez-Vásquez J (2011) Variations in a team: major and minor variants of Arabidopsis thaliana rDNA genes. Nucleus 2:294–299
Havlová K, Dvořáčková M, Peiro R et al (2016) Variation of 45S rDNA intergenic spacers in Arabidopsis thaliana. Plant Mol Biol 92:457–471
Chandrasekhara C, Mohannath G, Blevins T et al (2016) Chromosome-specific NOR inactivation explains selective rRNA gene silencing and dosage control in Arabidopsis. Genes Dev 30:177–190
Van Rossum G, Drake FL (2009) Python 3 reference manual. CreateSpace
SRA Toolkit Development Team. http://ncbi.github.io/sra-tools/
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.J 17:10–12
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760
Li H, Handsaker B, Wysoker A et al (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079
Acknowledgments
This work has been supported by a Human Frontiers Science Program (HFSP) Long-Term Fellowship (LT000819/2018-L) to FAR and by the Max Planck Society. I thank Sonja Kersten and Ilja Bezrukov for proofreading the manuscript and testing the code.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Rabanal, F.A. (2022). Characterization of the Different Levels of Variation in 45S rRNA Genes. In: Lambing, C. (eds) Plant Gametogenesis. Methods in Molecular Biology, vol 2484. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2253-7_24
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2253-7_24
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2252-0
Online ISBN: 978-1-0716-2253-7
eBook Packages: Springer Protocols