Abstract
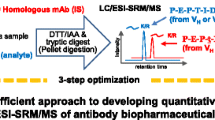
Monoclonal antibodies bind to Protein A/G resin with 100 nm-diameter pores, which orients the Fab toward the reaction solution. Then, they can be proteolyzed using trypsin immobilized on the surface of 200 nm-diameter nanoparticles. The difference between the two particle diameters allows Fab-selective proteolysis by limiting trypsin access to the antibody substrate. The specific signature peptide of monoclonal antibody is collected, which comprises the complementarity-determining regions (CDRs). Excess trypsin protease and peptide fragments from common sequences in Fc that inhibit the analysis can then be separated and removed. The resulting peptide samples are separated through high performance liquid chromatography on a 20 nm-diameter pore-size reversed-phase C18 column. These are then sequentially ionized with an electrospray interface and subjected to mass spectrometry (MS). In MS, peptide ions are trapped and fragment ions are generated by the collision-induced dissociation with argon gas. These are detected with multiple reaction monitoring measurements to perform a highly sensitive and accurate quantitative analysis.
By focusing on various physicochemical features at each analytical scene, such as characteristic structure and orientation of antibody, control of trypsin reaction field, carry-over on HPLC column, ionization suppression effect from endogenous proteins, and detection of amino acid sequence specificity of antibody, we optimized the overall conditions from the sample processing up to MS detection and developed analytical validation and clinical application of many therapeutic antibodies using our Fab-selective proteolysis technology that is based on the structure-indicated approach.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Duggan JX, Vazvaei F, Jenkins R (2015) Bioanalytical method validation considerations for LC-MS/MS assays of therapeutic proteins. Bioanalysis 7(11):1389–1395
Vande Casteele N, Herfarth H, Katz J, Falck-Ytter Y, Singh S (2017) American Gastroenterological Association Institute technical review on the role of therapeutic drug monitoring in the management of inflammatory bowel diseases. Gastroenterology 153(3):835–857. e836
Iwamoto N, Shimada T (2019) Regulated LC-MS/MS bioanalysis technology for therapeutic antibodies and fc-fusion proteins using structure-indicated approach. Drug Metab Pharmacokinet 34(1):19–24
Iwamoto N, Shimada T (2018) Recent advances in mass spectrometry-based approaches for proteomics and biologics: great contribution for developing therapeutic antibodies. Pharmacol Ther 185:147–154
Li H, Ortiz R, Tran L et al (2012) General LC-MS/MS method approach to quantify therapeutic monoclonal antibodies using a common whole antibody internal standard with application to preclinical studies. Anal Chem 84(3):1267–1273
Roman J, Qiu J, Dornadula G et al (2011) Application of miniaturized immunoassays to discovery pharmacokinetic bioanalysis. J Pharmacol Toxicol Methods 63(3):227–235
Kirresh T, Tuteja S, Russo D et al (2017) Development and validation of a liquid chromatography-mass spectrometric assay for simultaneous determination of tacrolimus and 13-O-desmethyl tacrolimus in rat kidney tissue. J Pharm Biomed Anal 136:32–37
Shibata K, Naito T, Okamura J et al (2017) Simple and rapid LC-MS/MS method for the absolute determination of cetuximab in human serum using an immobilized trypsin. J Pharm Biomed Anal 146:266–272
Jenkins R, Duggan JX, Aubry AF et al (2015) Recommendations for validation of LC-MS/MS bioanalytical methods for protein biotherapeutics. AAPS J 17(1):1–16
He J, Su D, Ng C et al (2017) High-resolution accurate-mass mass spectrometry enabling in-depth characterization of in vivo biotransformations for intact antibody-drug conjugates. Anal Chem 89(10):5476–5483
Carter P (2001) Improving the efficacy of antibody-based cancer therapies. Nat Rev Cancer 1(2):118–129
Scott AM, Wolchok JD, Old LJ (2012) Antibody therapy of cancer. Nat Rev Cancer 12(4):278–287
Lee SS, Bindokas VP, Kron SJ (2019) Multiplex three-dimensional mapping of macromolecular drug distribution in the tumor microenvironment. Mol Cancer Ther 18(1):213–226
Meroni PL, Valentini G, Ayala F et al (2015) New strategies to address the pharmacodynamics and pharmacokinetics of tumor necrosis factor (TNF) inhibitors: a systematic analysis. Autoimmun Rev 14(9):812–829
Iwamoto N, Hamada A, Shimada T (2018) Antibody drug quantitation in coexistence with anti-drug antibodies on nSMOL bioanalysis. Anal Biochem 540-541:30–37
Riaz N, Havel JJ, Makarov V et al (2017) Tumor and microenvironment evolution during immunotherapy with Nivolumab. Cell 171(4):934–949. e916
Ray A, Williams MA, Meek SM et al (2016) A phase I study of intratumoral ipilimumab and interleukin-2 in patients with advanced melanoma. Oncotarget 7(39):64390–64399
Mulleman D, Meric JC, Paintaud G et al (2009) Infliximab concentration monitoring improves the control of disease activity in rheumatoid arthritis. Arthritis Res Ther 11(6):R178
Force J, Leal JHS, McArthur HL (2019) Checkpoint blockade strategies in the treatment of breast cancer: where we are and where we are heading. Curr Treat Options in Oncol 20(4):35
Hassel JC, Heinzerling L, Aberle J et al (2017) Combined immune checkpoint blockade (anti-PD-1/anti-CTLA-4): evaluation and management of adverse drug reactions. Cancer Treat Rev 57:36–49
Wu TT, Kabat EA (1970) An analysis of the sequences of the variable regions of Bence Jones proteins and myeloma light chains and their implications for antibody complementarity. J Exp Med 132(2):211–250
Takai T (2002) Roles of Fc receptors in autoimmunity. Nat Rev Immunol 2(8):580–592
Iwamoto N, Shimada T, Umino Y et al (2014) Selective detection of complementarity-determining regions of monoclonal antibody by limiting protease access to the substrate: nano-surface and molecular-orientation limited proteolysis. Analyst 139(3):576–580
Iwamoto N, Umino Y, Aoki C et al (2016) Fully validated LCMS bioanalysis of bevacizumab in human plasma using nano-surface and molecular-orientation limited (nSMOL) proteolysis. Drug Metab Pharmacokinet 31(1):46–50
Iwamoto N, Shimada T, Terakado H, Hamada A (2016) Validated LC-MS/MS analysis of immune checkpoint inhibitor Nivolumab in human plasma using a Fab peptide-selective quantitation method: nano-surface and molecular-orientation limited (nSMOL) proteolysis. J Chromatogr B Analyt Technol Biomed Life Sci 1023–1024:9–16
Iwamoto N, Yokoyama K, Takanashi M et al (2018) Verification between original and biosimilar therapeutic antibody infliximab using nSMOL coupled LC-MS bioanalysis in human serum. Curr Pharm Biotechnol 19(6):495–505
Iwamoto N, Yokoyama K, Takanashi M et al (2018) Application of nSMOL coupled with LC-MS bioanalysis for monitoring the Fc-fusion biopharmaceuticals Etanercept and Abatacept in human serum. Pharmacol Res Perspect 6(4):e00422
Iwamoto N, Takanashi M, Yokoyama K et al (2019) Multiplexed monitoring of therapeutic antibodies for inflammatory diseases using Fab-selective proteolysis nSMOL coupled with LC-MS. J Immunol Methods 472:44–54
Iwamoto N, Yonezawa A, Matsubara K, Shimada T (2019) Acceleration of nano-surface and molecular-orientation limited (nSMOL) proteolysis with acidified reduction pretreatment for quantification of tocilizumab. J Pharm Biomed Anal 164:467–474
Acknowledgments
nSMOL development and clinical application are part of the collaboration results with Akinobu Hamada, Ph.D. and Hitoshi Nakagama, M.D., Ph.D. of National Cancer Center (Tokyo, Japan), Atsushi Yonezawa, Ph.D. and Kazuo Matsubara, Ph.D. of Kyoto University Hospital (Kyoto, Japan), Bernard Fox, Ph.D., William Redmond, Ph.D., Eric Tran, Ph.D., Brian Piening, Ph.D., Yoshinobu Koguchi, M.D., Ph.D., and Walter Urba, M.D., Ph.D. of Providence Cancer Institute (Portland, Oregon), Naoe Yamane, Ph.D. of CMIC (Tokyo, Japan), and Ms. Misato Hara and Mr. Naohiro Hanyu of Tamagawa Seiki (Nagano, Japan). We would sincerely express our deeply appreciation to this collaboration and their colleagues for significant support and discussion.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Iwamoto, N., Shimada, T. (2022). Structure-Indicated LC-MS/MS Bioanalysis of Therapeutic Antibodies. In: Houen, G. (eds) Therapeutic Antibodies. Methods in Molecular Biology, vol 2313. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1450-1_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1450-1_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1449-5
Online ISBN: 978-1-0716-1450-1
eBook Packages: Springer Protocols