Abstract
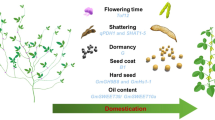
Over time, wild crops have been domesticated by humans, and the knowledge gained from parallel selection and convergent domestication-related studies in cereals has contributed to current techniques used in molecular plant breeding. Sorghum (Sorghum bicolor (L.) Moench) is the world’s fifth-most popular cereal crop and was one of the first crops cultivated by ancient farmers. In recent years, genetic and genomic studies have provided a better understanding of sorghum domestication and improvements. Here, we discuss the origin, diversification, and domestication processes of sorghum based on archeological discoveries and genomic analyses. This review also comprehensively summarized the genetic basis of key genes related to sorghum domestication and outlined their molecular mechanisms. It highlights that the absence of a domestication bottleneck in sorghum is the result of both evolution and human selection. Additionally, understanding beneficial alleles and their molecular interactions will allow us to quickly design new varieties by further de novo domestication.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Crop domestication has contributed to the rise of agriculture and the transition away from hunting and gathering, laying the groundwork for modern human civilization (Purugganan and Fuller 2009). Evidence collected from archeological discoveries and genetic and genomic studies indicates that crop domestication is a protracted process rather than a short, discrete event (Purugganan 2019). In general, the domestication process can be divided into four stages: in stage 1, humans harvest and consume wild plants; in stage 2, humans deliberately cultivate crops, resulting in genetic bottlenecks and a decline in wild alleles (Doebley et al. 2006); in stage 3, domesticated plants expand to new geographical regions and adapt to the local environment, resulting in a diversification of domesticated alleles and the formation of landraces; in stage 4, humans deliberately breed improved cultivars to meet the demands of modern diets (Gaut et al. 2018; Lenser and Theissen 2013). Compared with their wild ancestors, domesticated crops share some common morphological and physiological characteristics, which is known as domestication syndrome, including loss of seed shattering (non-shattering), loss of seed dormancy, seed enlargement, synchronous germination, and changes in tiller number and stature (Gepts 2014; Stetter et al. 2017). These traits typically occur under parallel/convergent domestication among different crops (Lenser and Theissen 2013; Purugganan 2019). The plant domestication center refers to the original geographic region where a specific species was domesticated. Three main cereal crops (rice, wheat, and maize) supply more than 50% of human calories, out of ~ 5500 food crops cultivated worldwide (Ross-Ibarra et al. 2007; Zhao et al. 2021), and were originally domesticated in East Asia, the Middle East, and Central America, respectively (Gepts 2014; Larson et al. 2014).
Sorghum (Sorghum bicolor (L.) Moench) is the fifth-most popular cereal crop in the world and is a staple food for more than 500 million people in Africa and Asia (Xin et al. 2021; FAOSTAT, https://www.fao.org/faostat/en/#data). It is estimated that sorghum was first cultivated as a food source in the Sahelian belt of Africa and was originally domesticated in central eastern Sudan approximately 6000 to 4000 years ago (Winchell et al. 2017). In addition to being a valuable source of calories, sorghum can also be used as fiber, forage, and fuel (Hao et al. 2021; Silva et al. 2021; Xie and Xu 2019). Since sorghum was domesticated and evolved in arid and semiarid ecosystems, it exhibits strong resistance to many abiotic stresses, such as drought, high light, barrenness, salt, and alkalinity, making it an ideal resistant plant resource to meet the modern demands of crop breeding to ensure food security under climate change (Ma et al. 2020; Prasad et al. 2021; Varoquaux et al. 2019; Xie and Xu 2019; Yang et al. 2020). Apart from the archeological and fossil discoveries, recent evidence from genetic and genomic studies of sorghum provides clues about its domestication at the molecular level (Baye et al. 2022; Wu et al. 2022, 2019; Xie et al. 2022; Zhang et al. 2018; Zhou et al. 2021). With the help of modern advanced biotechnologies, such as synthetic biology and gene editing techniques, we can rapidly improve current cultivars or create new crops by de novo domestication (such as in tomato and rice) to address the crises caused by food deficiency once we understand the underlying genetic basis of domestication-related genes (Li et al. 2018; Yu et al. 2021).
In this review, we discuss widely accepted opinions about sorghum domestication and characteristics based on archeological records and genomic studies. We comprehensively and systematically summarized the genetic basis and molecular mechanisms of major sorghum domestication-related genes. The conclusions regarding the artificial selection of sorghum will accelerate sorghum breeding processes when combined with advanced biotechnologies.
Sorghum origin, distribution, and classification
Although some issues are still under debate, it is widely accepted that sorghum originated from Africa in approximately 7500 BC, based on evidence gathered from archeological discoveries (Ananda et al. 2020). Some studies have reported the potential evolutionary history and dispersal route of domesticated sorghum (Burgarella et al. 2021; Fuller and Stevens 2018; Venkateswaran et al. 2019; Winchell et al. 2018). It has been proposed that sorghum was first domesticated in the eastern Sahelian zone in approximately 4000 BC and propagated to South Asia approximately 1000 years later (Winchell et al. 2017, 2018). Then, sorghum was introduced to China and domesticated into Chinese kaoliang (Zhang and Ping 2022).
The genus Sorghum consists of 23 or 24 species, though the taxonomy of this genus is still being debated (Ananda et al. 2020; Ohadi et al. 2018). Sorghum bicolor (L.) Moench subsp. bicolor contains all cultivated sorghum varieties, which were derived from the wild progenitor S. bicolor subsp. verticilliflorum (formerly known as subsp. arundinaceum), which is widely distributed in Africa (Berenji et al. 2011; Wet and Harlan 1971; Wet and Huckabay 1967). Instead of the complex early classification that divided cultivated sorghum into 158 varieties, Harlan and de Wet proposed a revised version that has been accepted by the following researchers (Harlan and de Wet 1972). According to the simplified morphological characteristics of the spikelet and head type, cultivated sorghum can be classified into five basic races: bicolor, guinea, caudatum, kafir, and durra. Race bicolor is considered the oldest race, and its seeds are more tightly covered by glumes than those of the other four races (de Wet and Shechter 1977). Race guinea likely evolved in West Africa and adapted to humid habitats with a loose head type and glumes opening at a large angle (de Wet et al. 1972). Race durra could be derived from hybridization between the race bicolor and local wild species in India (Harlan and Stemler 1976). It has a compact panicle and lower glumes than bicolor. Glumes typically have different textures between the tip and the base (de Wet and Shechter 1977). Race kafir shows glumes varying in length, derived from early bicolor or independently evolved from the local wild race in South Africa. The origin of race caudatum has not been fully determined. It could be directly selected from early bicolor with lower glumes and has become a major sorghum donor widely distributed in eastern Nigeria, Sudan, and Uganda (Shechter and de Wet 1975). The proposed original domestication center, main distribution, and spikelet morphology of five cultivated sorghum races in Africa are depicted in Fig. 1.
Domestication centers, original distribution, and spikelet morphology of five domesticated sorghum races (bicolor, guinea, caudatum, kafir, and durra). The early bicolor race was distributed in the eastern Sahelian zone and spread to other African regions. Its seeds are more tightly covered than those of the other four races. Race guinea probably evolved in West Africa and adapted to humid habitats with glumes opening at a large angle. Race kafir is widely grown in South Africa, with glumes varying in length. Race caudatum is dominant in eastern Nigeria, Sudan, and Uganda, with a relatively lower glume coverage. Race durra could be derived from the hybridization between race bicolor and the local wild species in India, and then subsequently re-introduced to Africa. Arrows indicate the possible spreading routes of race durra. The gray background represents the Sahelian zone
During agricultural cultivation, sorghum can be classified into different subgroups based on its end use. In the literature, domesticated sorghum has been classified into grain sorghum, sweet sorghum, forage sorghum, and biomass sorghum (Silva et al. 2021). However, the boundary between forage and biomass sorghum is unclear, and another classification added broom sorghum instead of biomass sorghum (Li and Yu 2022). Grain sorghum typically has larger seeds and a short lifespan, while sweet sorghum accumulates abundant soluble sugars in its stem. Forage sorghum is bred to feed animals with higher biomass and better palatability. Broom sorghum with long panicles is used to produce traditional homemade brooms (Berenji et al. 2011). The various morphological characteristics of sorghum are due to evolution and artificial selection, indicating that there are abundant genetic resources related to these traits.
Genomic footprints of sorghum domestication
Morphological variation during crop evolution is mainly determined by genetic modification in the nucleotide sequence of the genome. Sorghum is a diploid crop (2n = 2x = 20). The whole-genome assembly of the sorghum cultivar BTx623 was first achieved using de novo sequencing in 2009, and the reference genome size is approximately 730 Mb (Paterson et al. 2009). A subsequently improved genome assembly version annotated 34,211 genes (McCormick et al. 2018). Additionally, the reference genomes of the sorghum cultivar Tx430 and a sweet sorghum line Rio were also released in recent years (Cooper et al. 2019; Deschamps et al. 2018). The genome sequences of the other three sorghum accessions, BTx642, RTx430, and SC187, are also available on the Phytozome website (https://phytozome-next.jgi.doe.gov/). A comparison of genomic and transcriptomic data between BTx623 and Rio revealed structural variations and differentially expressed genes involved in sugar metabolism (Cooper et al. 2019). The pangenome of cultivated and wild sorghum revealed new annotated genes and presence and absence variations (PAVs) that could play an important role in sorghum domestication and diversification (Ruperao et al. 2021; Tao et al. 2021).
Advancements in sequencing technology and bioinformatics make it possible to analyze genetic information at the population level, increasing our knowledge of the evolutionary trajectory on many species, such as rice, soybean, tomato, lettuce, and apricots (Groppi et al. 2021; Lin et al. 2014; Lu et al. 2020; Qin et al. 2021; Wei et al. 2021). In sorghum, resequencing analysis of nine archeological accessions discovered at Qasr Ibrim from approximately 800 BC, as well as wild and cultivated sorghum genomes, revealed that there was no obvious domestication bottleneck in sorghum, but that there was a gradual decline in genetic diversity. It also induced an increase in deleterious mutations in cultivated sorghum lines, known as the “cost of domestication” (Gaut et al. 2018; Smith et al. 2019). Evidence from the genome information demonstrated the presence of occasional hybridization between the Asian durra type and African bicolor type, which led to genetic rescue and diversification in sorghum domestication. Hybridization or genetic introgression was also discovered between cultivated sorghum and their weedy or wild relatives, indicating the occurrence of high-frequency active gene flow in sorghum. This differs from other strict self-pollination crops (Ohadi et al. 2018).
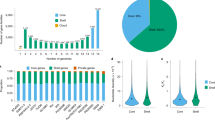
A recent population genetic study deepened our understanding of sorghum domestication and diversification at the whole-genome level. A total of 445 sorghum accessions were classified into seven subpopulations, including wild, sudangrass, landrace broom (LB), landrace grain (LG), improved grain (IG), improved sweet (IS), and ambiguous lines (AL), based on distinct phenotypic differentiation. Analysis of whole-genome resequencing data also revealed the existence of frequent genetic exchanges between LG and the wild type. LB lines, mostly collected from Asia, received gene flow from the common ancestor with LG, while IS received gene flows from both LB and sudangrass (Wu et al. 2022). In addition, genomic regions with selection signals were identified in different groups, and eight models based on the haplotype changes of domesticated genes were proposed. Among them, two important models represented soft selection with multiple domestication origins and hard selection with only one domestication event. Deep whole-genome resequencing and analysis of a sorghum association panel (SAP) containing 400 sorghum accessions identified 18 genomic regions with significant Fst peaks. These peaks varied in the five typical domesticated races, which reflects evolutionary differences and relationships during sorghum domestication. More importantly, these regions overlap with some previously reported quantitative trait loci (QTLs), indicating that the potential genetic mechanism underlying domestication-related traits could be uncovered in the future (Boatwright et al. 2022).
Genetic dissection of domestication-related traits in sorghum
Studies analyzing the genetics and molecular biology of sorghum have identified many genes related to sorghum domestication, and the underlying genetic basis has been well elucidated. In the following text, we summarize the major domesticated genes in sorghum and introduce their molecular mechanisms. These genes are mostly involved in panicle-related traits, such as seed shattering, tannin content, awn and glume coverage, while others are related to stem juiciness, flowering, and plant architecture, including plant height and tiller number (Fig. 2 and Table 1).
Plant architecture of wild and weedy sorghum and domesticated sorghum cultivar. The major genes controlling sorghum domestication-related traits are SbTB1 (reduced tillers), Dw1, Dw2, and Dw3 (dwarfism), Dry (stem juiciness), SbPRR37, Ma2, PhyB, Ma4, PhyC, and SbGhd7 (early-flowering time in LD), GC1 (naked grains), awn1 (awnless), Sh1 and SpWRKY (non-shattering), and Tan1 and Tan2 (non-tannin). LD long day, Sb, Sorghum bicolor
Plant height and tillers
Plant height is one of the most prominent target characteristics in crop breeding. Dwarfism in crops can increase lodging resistance and decrease grain yield loss. In rice and wheat, semidwarf varieties with deficiencies in gibberellin biosynthesis and signaling pathways were bred by breeders and subsequently initiated the “green revolution” (Wang et al. 2021). However, dwarfism is not a universal breeding goal for sweet or forage sorghum due to the demand for varieties with tall and robust statures and high biomass. Additionally, decreased plant height is widely selected in grain sorghum or other races with high grain yield.
It has been reported that sorghum plant height is controlled by four independent loci, named Dw1, Dw2, Dw3, and Dw4 (Quinby and Martin 1954). Recessive mutations of each locus could lead to decreases in plant height. The underlying genes of the Dw1, Dw2, and Dw3 loci have been cloned. Dw3 encodes a protein containing transmembrane domains and ATP-binding domains. It functions in transporting auxin from the middle to lower stem tissues in a light-dependent manner. A nonfunctional allele with an 882 bp duplication in the fifth exon of Dw3 was found in the dw3 mutant. This mutation caused the dwarf phenotype with a reduction in internode length. However, the unequal crossing over of this duplication led to an unstable dwarf phenotype as a few accidental offspring individuals with tall plant height (Multani et al. 2003). Dw1 was finely mapped to a 33 kb region on chromosome 6, and Sobic.009G229800 was considered the candidate gene (Hilley et al. 2016). Dw1 encodes a putative membrane protein with a higher expression level in elongating middle stems. The dwarf parent 80M carries the dw1 allele with a truncated protein caused by an A to T mutation in the second exon (Yamaguchi et al. 2016). However, the molecular mechanism of Dw1 remains unknown. The Dw2 locus has been detected in many studies. Through fine-mapping from two recombinant inbred lines (RIL) populations, Sobic.006G067700 was finally confirmed as the candidate gene instead of the previously reported gene Sobic.006G067600 (Hilley et al. 2017). Dw2 encodes a protein kinase belonging to the AGCVIII subfamily. The recessive dw2 allele carries an Indel in the first exon, which causes a frameshift and a truncated protein. A subsequent study revealed that cell proliferation was repressed in NIL-DDYM (dw2 allele), resulting in a shortened internode. In addition, mutation of Dw2 caused irregular cell shapes and altered the morphology of vascular bundles. Cell wall polysaccharide components were also changed in dw2. Phosphoproteomic analysis demonstrated that Dw2 was also involved in regulating lipid signaling and endomembrane trafficking (Oliver et al. 2021). Genome-wide selection signal detection found that the Dw2 region has a much higher Fst value in the improved accessions than in the five basic races, indicating that Dw2 was under positive selection in crop improvement practice after domestication (Boatwright et al. 2022). Other plant height-related regions were also detected in a genome-wide association study (GWAS), but it is unclear whether the genes are related to domestication.
Another agronomic trait related to shoot architecture is tiller number. Compared with their wild ancestors, domesticated and improved crop cultivars exhibit a great reduction in tiller number due to the benefits of close planting. TB1, a basic helix-loop-helix transcription factor negatively regulating tiller number, was first cloned in maize, and its orthologous gene in rice or sorghum plays similar roles (Doebley et al. 1997). A higher expression level of SbTB1 observed in the sorghum phyB-1 mutant revealed the relationship between tiller development and light signals (Kebrom et al. 2006). A population genetic study proved that the SbTB1 region was under strong selection during sorghum domestication. All sorghum cultivars carry the same haplotype, while variations in the promoter region of SbTB1 could explain the changes in tiller number between wild and domesticated species (Wu et al. 2022).
Stem juiciness
The crop stem parenchyma cells play a role in storing water and nutrients, but become dry or form cavities surrounded by epidermal cells at the mature stage. Grain sorghum and sweet sorghum exhibit distinct stem morphology: grain sorghum stems become dry and pithy, while sweet sorghum stems maintain much higher water content and are juicy. This absorbing phenotype in sorghum has been investigated since last century (Swanson and Parker 1931). Early studies suggested that pithy or juicy stems in sorghum were determined by a single D locus, and that pithy was dominant to juicy. It has also been observed that pithy and juicy stems were closely associated with white and green midribs, respectively. In 2001, Hart et al. reported that the D locus was cosegregated with the Xtxp97 marker in a sorghum genetic map (Hart et al. 2001). Both later GWAS and bulked segregant analysis (BSA) have mapped the D locus, and another study narrowed the D locus region with only six genes inside by map-based cloning (Han et al. 2015; Upadhyaya et al. 2022; Zhai et al. 2014). However, the underlying gene and its role in the origin of sweet sorghum remained obscure until two 2018 studies.
Using an F2 population derived from a cross between a dry-stem sorghum SKS and a juicy-stem sorghum MS3B, Fujimoto et al. (2018) finely mapped the D locus to an 18.99-kb interval on chromosome 6, and one gene, Sobic.006G147400 (referred to as the D gene), was confirmed as the candidate gene controlling the juicy content in sorghum. The D gene is a NAC domain transcription factor that can trigger the programmed cell death (PCD) of stem parenchyma cells, which leads to dry and pithy stems in sorghum. All the investigated wild sorghum and dry-stem sorghum cultivars possess a functional D gene, while six nonfunctional alleles of the D gene were discovered in juicy-stem sorghum accessions. Haplotype distribution of the D gene was found in both African and Asian germplasms. Two nonfunctional alleles were only distributed in African germplasms, which indicated that a nonfunctional D gene could have been selected at an early stage in Africa.
In another study, the same underlying gene (referred to as Dry) in the D locus was also identified using GWAS and map-based cloning (Zhang et al. 2018). The authors discussed more about the relationship between Dry selection and the origin of domesticated sweet sorghum. Principal component analysis (PCA) based on the whole-genome resequencing data of 241 sorghum accessions collected worldwide revealed that the wild, pithy, and juicy accessions were clustered into three different groups, although there were some exceptions in both the pithy and juicy groups. Dry acts as a master transcription factor that can regulate a series of genes involved in cell wall biosynthesis and loss of function of the Dry gene in juicy sorghum causes irregular parenchyma cells and thinner secondary cell walls. A positive selection signal was detected in the juicy sorghum subgroup, which had a significantly lower π value than the pithy subgroup. Twenty-three haplotypes of the Dry gene were found in 42 wild sorghum lines with pithy stems. Two of them were discovered in 86 landraces exhibiting dry pithy stems, and 112 improved cultivars carried four nonfunctional haplotypes. This variation in Dry gene haplotype diversity indicates that a bottleneck effect exists in the Dry gene during sweet sorghum domestication. In addition, the Dry locus also exists in collinear genomic regions in other cereal crops, such as rice, wheat, millet, and maize, indicating that the Dry locus is an important target in designing crops with both high grain yield and high stem biomass.
Flowering
Sorghum is a short-day C4 grass with photoperiod-sensitive characteristics that evolved in a tropical, equatorial region. It adapted to long-day environments after dispersing to high latitudes and temperate regions. Early flowering was generally selected for grain sorghum to ensure reproduction by avoiding drought or low temperature, while other sorghum types, such as sweet sorghum, forage sorghum, and energy sorghum, were selected to have a longer duration of vegetative growth to acquire a higher biomass yield. The diversification of sorghum flowering time under long-day conditions indicates that multiple genes could be selected to produce photoperiod-insensitive varieties. Breeders have paid close attention to identifying genes controlling sorghum maturity, and a series of maturity loci (named Ma1 to Ma6) have been reported over the last century (Quinby and Karper 1945; Quinby 1966, 1967; Rooney and Aydin 1999). All six loci are dominant in suppressing flowering under long-day conditions, and the Ma1, Ma2, Ma3, Ma5, and Ma6 loci were identified in these years, which allowed us to obtain a better understanding of their related genetic basis.
Using a BC1F1 population and another F2 population, Ma1 was fine-mapped to an 86-kb interval on chromosome 6, and Sb06g014570 (SbPRR37) was confirmed as the candidate gene (Murphy et al. 2011). SbPRR37 encodes a pseudoresponse regulator, and its expression has two peaks in the morning and evening on long days instead of only one morning peak on short days. The higher expression level of SbPRR37 on long days activates the expression of the floral inhibitor gene CONSTANS (CO) and represses SbEhd1 (Early Heading Date 1), which ultimately downregulates FT genes (SbCN8 and SbCN12) expression. SbCN8 and SbCN12 are florigen genes that play substantial roles in inducing flowering time (Turck et al. 2008; Yang et al. 2014b). Three nonfunctional alleles were found in early-flowering accessions cultivated in long-day conditions, which indicates the possible multiple origins of photoperiod-insensitive sorghum in diversification progress when sorghum spreads to temperate regions. Another association analysis study of sorghum maturity also identified the SbPRR37 gene by using GWAS in a sorghum mini-core collection (Upadhyaya et al. 2013). Multiple variant alleles of the SbPRR37 genomic sequence from 253 landraces and historic sorghum cultivars were found. Some alleles dominantly distributed in a specific sorghum race or geographic region reveals the selection history and gene flow of PRR37 when sorghum was introduced to high latitudes in a new continent by human activity. For example, prr37Kafir−1 and prr37Kafir−2 alleles were mainly distributed in the Kafir race, and prr37Durra was dominant in Chinese kaoliang (Klein et al. 2015).
SbGhd7, encoding a protein containing a CCT domain (CONSTANS, CO-like, and TOC1), was identified as the candidate gene in the Ma6 loci (Murphy et al. 2014). Its orthologous gene in rice, GHD7, inhibits the expression of EHD1, which regulates the expression of Hd3a (FT gene in rice) in response to day length. SbGhd7 has a similar expression pattern to SbPRR37 and inhibits the expression of SbEhd. Two recessive ghd7 alleles were found in photoperiod-insensitive cultivars. The dominant alleles of SbGhd7 and SbPRR37 act in an additive fashion to delay flowering under long-day conditions. Other studies confirmed that phytochrome B (PhyB) is the causal gene in Ma3 loci, and phytochrome C (PhyC) was proposed as the underlying gene for Ma5 (Childs et al. 1997; Yang et al. 2014a). Both PhyB and PhyC have been reported to be related to flowering regulation in a light-dependent manner in Arabidopsis and rice. Genetic evidence revealed that PhyB is epistatic to Ma1 (SbPRR37) and Ma6 (SbEhd). PHYB inhibits the expression of SbEhd1, which activates the expression of SbCN8 and SbCN12 under long-day conditions. Another sorghum FT gene, SbCN15, is also repressed by PhyB regardless of photoperiod. The underlying gene of Ma2 was finely mapped as Sobic.002G302700, which encodes a lysine methyltransferase with a SET and MYND (SYMD) domain (Casto et al. 2019). It enhances the expression of SbPRR37 and SbCO. A genetic interaction between Ma2 and Ma4 was observed. Two recessive ma2 alleles were discovered in early-flowering sorghum lines under long-day conditions. These studies shed light on the scenario of sorghum diversification and improvement under human selection for adaptation in temperate regions.
Awn
The awn is a needle-like structure that extends from the lemma and is very common in gramineous crops, such as wheat, barley, rice, oats, and sorghum (Gu et al. 2015). There are some advantages to having awns for wild species. For example, the awn can prevent insects and birds from predating the seeds, while a barbed awn can help the seeds efficiently spread by sticking to animal furs (Hua et al. 2015; Jagathesan et al. 1961). It has also been reported that awns can contribute to yield by producing more photosynthate in wheat and barley (Du et al. 2021). In wild wheat, awns can help the seeds germinate by pushing them into the soil. Awns can bend or twist when the surrounding humidity changes and therefore produce the mechanical force needed to orient the spikelet (Elbaum et al. 2007). However, long cultivar awns cause difficulties in harvest, processing, and storage. As a result, breeding short-awn or awnless varieties occurs during crop domestication and improvement.
In addition to the normal seed-bearing spikelets, sorghum inflorescence has sterile pedicellate spikelets. It has been proven that the sterile spikelets in sorghum, not the awn, can have the capacity for photosynthesis (AuBuchon-Elder et al. 2020). Therefore, sorghum awns are likely not a carbon source, although all wild sorghum species have long awns. Girma et al. (2019) conducted a GWAS of the presence or absence of awns using 1425 Ethiopian landrace accessions. The GWAS results revealed a leading peak at 72.6 Mb on chromosome 3 and identified eight significant SNPs (Girma et al. 2019). However, no genetic confirmation was performed in this study, and the underlying mechanism remains unclear.
A recent study identified a major gene, Awn1, which is responsible for awn loss in cultivated sorghum (Zhou et al. 2021). In this study, Zhou et al. finely mapped the awn1 gene through a RIL population constructed from a cross between the wild sorghum progenitor Sorghum virgatum (SV) and the improved sorghum cultivar Tx623. After narrowing down the interval into a 9.5-kb region on chromosome 3, sequence comparison showed a large 5.4-kb insertion in the domesticated sorghum cultivar Tx623. Only one gene, Sobic.003G42130, was annotated in this insertion, which was the same candidate gene in the abovementioned GWAS. Sobic.003G42130 was named awn1 and was proven to be derived from an ancestral homologous gene on chromosome 10, Sobic.010G225100, which was referred to as awn1-10. Awn1 encodes the identical protein with the ALOG domain but recruits a new promoter and has a higher expression level than awn1-10. Transcriptional activity assays and yeast two-hybrid assays proved that Awn1 is a transcriptional repressor. RNA-seq and DAP-seq analysis revealed that it can downregulate some MADS-box genes involved in flower development and the orthologous genes of DL and LKS2 of rice, leading to a reduction in awn elongation in sorghum. Genomic sequence comparison found that this 5.4-kb fragment on chromosome 10 between SV and Tx623 had more SNPs than between awn1 and awn1-10 in Tx623, indicating that duplication on chromosome 3 could have occurred after domestication. Tajima’s D test revealed a significant selection signal in the neighboring regions of Awn1, and the awnless sorghums had the lowest genomic diversity around this region. Awn1 is a largely effective gene for more than 30% of phenotypic explanations of awn presence or absence in a natural sorghum population, which could be used in further awnless sorghum breeding. Homologs of Awn1 could be further exploited in other cereals, and it is unclear whether it also experienced similar parallel selection.
Glume coverage
Wild sorghum has a pair of tenacious glumes that cover the seed, which can protect the seed from being infected by fungi, birds, and insects in the natural environment. However, it caused a huge obstacle for threshing during sorghum domestication. In agricultural practice, seeds tightly covered by glumes pose difficulties for modern automated planting, threshing, and processing (Adeyanju et al. 2015). As a result, farmers favor sorghum grains with low glume coverage, which induces enriching variations in glume coverage in modern sorghum accessions. The spikelet morphology of sorghum differs from that of rice and maize. In rice, the glumes degenerated. The revolved hard lemma and palea act as glumes to cover the rice seed. In maize cultivars, the seeds are naked, and the whole ear is covered by bracts (Wu et al. 2019). The different spikelet structures indicate that sorghum could acquire a distinct regulatory network to control glume coverage during domestication.
Recently, Xie et al. (2022) identified a major gene located on chromosome 1, GC1, which controls sorghum glume coverage using GWAS and positional cloning. Five main haplotypes were identified from 482 sorghum accessions, named WT GC1, and mutated gc1-a, gc1-b, gc1-c, and gc1-d. Among them, gc1-b, gc1-c, and gc1-d were rare, while the GC1 (71%) and gc1-a (24%) haplotypes were dominant in the evaluated accessions. The association test demonstrated that GC1 variation was highly associated with glume coverage instead of yield-related traits (seed length, seed width, and thousand seed weight). GC1 encodes a protein of 198 amino acids with a Gγ-like domain (referred to as GC1-G) and a predicted transmembrane domain (GC1-T), while gc1-an obtains a stop codon at amino acid position 137 but reserves the entire GC1-G and GC1-T domains. The truncated gc1-a exhibited much lower glume coverage than WT GC1. Overexpression of GC1 reduced glume coverage in independent transgenic lines. Interestingly, knocking out GC1 increased glume coverage, which was different from what was observed in gc1-a, indicating that the truncated protein could still function during the regulation of glume coverage. gc1-a-overexpressing plants confirmed this hypothesis with significantly reduced glume coverage compared with GC1. These results indicated that both GC1 and the truncated gc1-a negatively control glume coverage in sorghum. Overexpressing or knocking out the orthologous gene of GC1 in millet also supported this conclusion.
Further study confirmed that the truncated C-terminus of GC1 caused a higher protein accumulation in vivo, which inhibited glume cell proliferation by downregulating cyclin-CDK-related genes. An interacting phospholipase protein, SbpPLAII-1, identified by immunoprecipitation-mass spectrometry (IP-MS), promoted the expression of cyclin-CDK-related genes and longer glumes. SbpPLAII-1 could be degraded when GC1 or gc1-a was accumulated. The more stable gc1-a accelerated this degradation process compared with GC1. These results demonstrated that naturally truncated variations of GC1 contributed to lower glume coverage in sorghum by degrading SbpPLAII-1 and downregulating the expression of cell division-related genes. Tajima’s D test showed a significant selection signal in landraces and improved cultivars with low glume coverage. In addition, the nucleotide diversity in the exon 5 and 3’UTR of GC1 was also significantly reduced in the naked landraces and improved lines compared with wild sorghum. The geographic distributions of five different haplotypes in this study also indicated that the Sahelian zone is a domestication center of sorghum.
Tannin content
Proanthocyanidins (PAs) are condensed tannins and are products of the flavonoid biosynthesis pathway. They have astringent properties and are present in sorghum grains but absent in other main cereal crops, such as corn, wheat, and rice (Wu et al. 2012). Tannin evaluation of 11,577 cultivated sorghum accessions showed that tannin and non-tannin types were present in approximately equal proportions (45% and 55%, respectively) (Wu et al. 2019). However, it is unclear why domesticated sorghum still preserved these bitter chemicals under human selection. An early study reported that the presence of tannin in sorghum grain was regulated by two genes (B1 and B2) (Smith and Frederiksen 2000). Wu et al. (2012) identified the Tannin1 (Tan1) gene involved in condensed tannins biosynthesis in sorghum. In 2019, two studies identified the same locus related to the presence of tannin in sorghum and shed light on the underlying regulatory mechanism between tannin content and bird feeding behavior.
Xie et al. (2019) first collected phenotypic data by evaluating bird-preference and bird-avoidance characteristics from the field and identified a significant SNP within the Tan1 gene involved in flavonoid and PA biosynthesis by GWAS. This locus was stably detected regardless of the phenotype of tannin content or by bird damage levels under different populations and field conditions, indicating that there was a relationship between tannin content and the level of bird damage. This was proven because bird-preference sorghum accessions had significantly reduced levels of metabolites involved in anthocyanin and PA biosynthesis. Two mutated alleles, tan1-a and tan1-b, with no detected tannins were associated with a much more severe bird damage phenotype compared with the wild-type Tannin1 (Tan1). Sparrow feeding experiments demonstrated that sparrows fed on fewer seeds coated with malvidin or PA than untreated seeds. On the other hand, bird-preference seeds with tan1-a/b alleles produced more fragrant volatile organic compounds, such as 1-octen-3-ol and hexanal, which caused a longer residence time for sparrow feeding. It is known that the WD40 protein can form a ternary complex by interacting with MYB and basic helix-loop-helix (bHLH) proteins to control multiple biological processes (Ramsay and Glover 2005). Further studies revealed that tan1-a/b likely promoted more fatty acid-derived volatiles than Tan1, possibly by repressing the expression of SbGL2, a key negative regulator of fatty acid biosynthesis.
Wu et al. (2019) reported that Tan1 and another gene, Tannin2 (Tan2), were both involved in sorghum tannin content regulation. It was identified by QTL mapping and a combined GWAS. Using the different bird damage levels as phenotype data, five significant loci were identified from QTL mapping of a RIL population derived from a cross between P898012 and Tx430. Three of them were related to plant height, but only two significant loci located on chromosomes 4 and 2 were left when using the tannin presence as the phenotype data. These two loci were also identified in the GWAS using tannin content as the input data. The mapping region on chromosome 4 contained Tannin1, which has been proven to regulate tannin presence in sorghum grains in a previous study (Wu et al. 2012), and the allele carrying a 1-base pair insertion (referred to as tan1-a) in the coding sequence caused the non-tannin grains in Tx430. After sequence variation analysis, Sobic.002G076600 was considered the candidate gene of Tan2. This was also confirmed by complementing the Arabidopsis tt8 mutant with modified seed pigmentation (Nesi et al. 2000). A 5-bp insertion in Tan2 (encoding a protein with bHLH domain) caused a frameshift in Tx430 and was denoted as tan2-a.
Tannin and non-tannin plants are segregated at a ratio of 1:3 in the RIL population. These results indicated that Tan1 and Tan2 were the underlying genes of the B1 and B2 loci, respectively, and sorghum grain tannin was present only when both genes were dominant. The recessive alleles, tan1-b, tan1-c, and tan2-b, tan2-c, of each gene were discovered from 88 non-tannin accessions, with tan1-b and tan2-b at a low frequency. tan1-a was mainly distributed in East and West Africa, while tan2-a dominated in South and West Africa. In addition, tan1-c and tan2-c were mainly present in South and East Africa, and no tan2-a allele was observed in East Africa. These results indicate that non-tannin sorghum could have multiple domestication origins. Further study confirmed that tannin sorghum had a higher proportion in East and South Africa where bird damage was severe, while non-tannin sorghum dominated in West Africa where bird threats were mild. Interestingly, the geographic distribution of tannin sorghum was also connected to human TAS2R variants. It was assumed that humans carrying a TAS2R haplotype were insensitive to bitter tastes, thus contributing to the selection of tannin sorghum to address local severe bird threats in East and South Africa. In contrast, humans carrying another TAS2R haplotype could perceive a bitter taste and prefer non-tannin sorghum in West Africa. The interactions among plants, humans, and the environment demonstrate the complexity of tannin content domestication in sorghum.
Seed shattering
Loss of seed shattering is considered a hallmark of crop domestication (Li et al. 2006). Wild ancestors of modern crops disperse their seeds by forming an abscission layer between the seed and pedicel, which causes seed shattering and helps their seeds fall off into the soil in a timely manner and propagate efficiently. However, seed shattering is an unfavorable phenotype for farmers, since it results in great yield losses and causes difficulties in harvesting. As a result, non-shattering variants were selected during domestication among most crops.
Sh1 is the major gene controlling sorghum seed shattering (Lin et al. 2012). It is cloned using a large F2 population containing approximately 15,000 individuals derived from a cross of a wild sorghum Sorghum virgatum (SV) and a domesticated cultivar Tx430. A YABBY domain transcription factor was identified as the candidate gene of Sh1 loci. Sequence variation in Sh1 revealed four main haplotypes, including the wild haplotype SV-like Sh1 (Sh1SV−like) and three domesticated haplotypes, SC265-like Sh1 (Sh1SC265−like), Tx430-like Sh1 (Sh1Tx430−like), and Tx623-like Sh1 (Sh1Tx623−like). Compared to Sh1SV−like, Sh1Tx430−like has four causal changes in the promoter and the second intron, which leads to a lower expression level. Variations in Sh1Tx623−like and Sh1SC265−like cause frameshift mutations, resulting in truncated proteins lacking the zinc finger and YABBY domains. The proportion of these three non-shattering haplotypes varies in domesticated sorghum races. For example, in the investigated sorghum accessions, Sh1Tx430−like is prevalent in the caudatum race, while all durra races, most guinea races, and approximately half of bicolor races carry Sh1SC265−like. The other haplotype, Sh1Tx623−like, is mainly comprised of kafir and bicolor races and is widely distributed in South and East Africa. These results indicate that non-shattering sorghum could have been simultaneously domesticated in different regions from local relative wild ancestors. Another study found that the three wild sorghum accessions SL129, SL12, and SL32, collected from Kenya, Nigeria, and Tanzania, respectively, could be the ancestors of the non-shattering haplotypes Sh1Tx430−like, Sh1SC265−like, and Sh1Tx623−like, respectively (Wu et al. 2022). Interestingly, the syntenic region harboring orthologous genes related to the non-shattering phenotype in rice and maize was also under positive selection, which illustrates that the orthologous genomic region of Sh1 underwent parallel selection in different cereal lineages.
SpWRKY, identified from a wild sorghum species, Sorghum propinquum, is also a major gene for sorghum seed shattering (Tang et al. 2013). Compared to the non-shattering allele SbWRKY, SpWRKY has a longer translated protein since it recruits a new start codon. However, it seems that SbWRKY is not related to domestication. It is more likely that Sorghum propinquum keeps the seed shattering by obtaining a functional SpWRKY from a shared common ancestor with Sorghum bicolor. Intriguingly, the expression of SpWRKY in non-shattering RTx430 restored the shattering phenotype. Regardless, the relationships and molecular mechanisms between the two genes remain largely unknown and must be further elucidated.
Conclusions and perspectives
Crop domestication is a protracted process that entails the early selection of wild progenitors, subsequent diversification when the landrace spreads to a new environment, and modern improvement breeding. In this review, we discussed the origin and classification of five basic cultivated breeds of sorghum. Genomic information reveals that there is no obvious domestication bottleneck in sorghum, which differs from other main crops. We summarized key genes that have recently been reported to be involved in sorghum domestication and elucidated their genetic molecular mechanisms. The underlying genes of these agronomic traits were discussed in detail, and the genomic footprints of these genes give us a better understanding of how plants can achieve ideal traits between the environment and human activity during their evolutionary process.
Cereal domestication started in the Neolithic Age approximately 10,000 years ago. During the evolutionary process of plants, multiple agronomic traits were altered that benefited humans during parallel selection across major crops, which is known as domestication syndrome (Gepts 2014; Stetter et al. 2017). The genes that occur in parallel selection always occupy few but important signaling pathway positions. Loss or gain of function in these paralleled selected genes are conserved in related crop species despite different mutation forms (Lenser and Theissen 2013). Recently, an increase in our knowledge about key genes underlying domestication-related traits has been identified in rice, maize, and wheat (Fernie and Yan 2019), which could inspire research of sorghum domestication. For example, the only identified major gene, SbTB1, controlling tiller number in sorghum, is the ortholog of well-known TB1 in maize (Doebley et al. 1997). However, this does not occur in some homologs of those reported distinguished genes due to distinct selection pressure. Millet SiGC1, the ortholog to sorghum GC1, did not exhibit a parallel section in naked grains since it already had thin glumes and easy-threshing grains (Xie et al. 2022). It is noteworthy that these key domesticated genes are also located in simple regulatory pathways and have minimal pleiotropic effects, which can prevent additional side effects on other important agronomic traits, such as the flowering time pathway (Turck et al. 2008).
One cost of crop domestication is mutation load, which refers to the increase in deleterious mutations in the genome (Gaut et al. 2018). Domestication also caused a loss of rare or elite alleles that could resist biotic or abiotic stress. Interestingly, domesticated crops or animals can reacquire wild-like traits, which is called the de-domestication process (Wu et al. 2021). In sorghum, the effects of mutation load can be mitigated by occasional hybridization between different sorghum accessions (Brown 2019; Smith et al. 2019). A modern understanding of the molecular mechanisms underlying domestication-related genes and the site-directed mutations by highly-efficient genome editing techniques is ushering in the era of Breeding 4.0 (Wallace et al. 2018). Using advanced biotechnologies such as genome editing techniques, researchers can now rapidly achieve de novo domestication of a crop from its wild ancestors. This could preserve the elite genes lost during human selection (Huang et al. 2022; Lyzenga et al. 2021). Sorghum has the potential to be widely utilized not only for food and forage, but also for industrial raw materials, such as sugar and biofuel. The results of genetic studies and molecular biology provide a better understanding of sorghum domestication at the molecular level. Additionally, the beneficial alleles of the major domestication-related traits could contribute to the efficient and accurate breeding of sorghum.
Data availability
Data sharing is not applicable to this article, as no datasets were generated or analyzed during the current study.
References
Adeyanju A, Perumal R, Tesso T (2015) Genetic analysis of threshability in grain sorghum [Sorghum bicolor (L.) Moench]. Plant Breeding 134(2):148–155. https://doi.org/10.1111/pbr.12244
Ananda GKS, Myrans H, Norton SL, Gleadow R, Furtado A, Henry RJ (2020) Wild sorghum as a promising resource for crop improvement. Front Plant Sci 11:1108. https://doi.org/10.3389/fpls.2020.01108
AuBuchon-Elder T, Coneva V, Goad DM, Jenkins LM, Yu Y, Allen DK, Kellogg EA (2020) Sterile spikelets contribute to yield in sorghum and related grasses. Plant Cell 32(11):3500–3518. https://doi.org/10.1105/tpc.20.00424
Baye W, Xie Q, Xie P (2022) Genetic architecture of grain yield-related traits in sorghum and maize. Int J Mol Sci 23(5):2405. https://doi.org/10.3390/ijms23052405
Berenji J, Dahlberg J, Sikora V, Latkovi D (2011) Origin, history, morphology, production, improvement, and utilization of broomcorn [Sorghum bicolor (L.) Moench] in Serbia. Econ Bot 65(2):190–208. https://doi.org/10.1007/s12231-011-9155-2
Boatwright JL, Sapkota S, Jin H, Schnable JC, Brenton Z, Boyles R, Kresovich S (2022) Sorghum association panel whole-genome sequencing establishes cornerstone resource for dissecting genomic diversity. Plant J 111(3):888–904. https://doi.org/10.1111/tpj.15853
Brown TA (2019) Is the domestication bottleneck a myth? Nat Plants 5(4):337–338. https://doi.org/10.1038/s41477-019-0404-1
Burgarella C, Berger A, Glemin S, David J, Terrier N, Deu M, Pot D (2021) The road to sorghum domestication: evidence from nucleotide diversity and gene expression patterns. Front Plant Sci 12:666075. https://doi.org/10.3389/fpls.2021.666075
Casto AL, Mattison AJ, Olson SN, Thakran M, Rooney WL, Mullet JE (2019) Maturity2, a novel regulator of flowering time in Sorghum bicolor, increases expression of SbPRR37 and SbCO in long days delaying flowering. PLoS ONE 14(4):e0212154. https://doi.org/10.1371/journal.pone.0212154
Childs KL, Miller FR, Cordonnier-Pratt M-M, Pratt LH, Morgan PW, Mullet JE (1997) The sorghum photoperiod sensitivity gene, Ma3, encodes a phytochrome B. Plant Physiol 113(2):611–619. https://doi.org/10.1104/pp.113.2.611
Cooper EA, Brenton ZW, Flinn BS, Jenkins J, Shu S, Flowers D, Luo F, Wang Y, Xia P, Barry K, Daum C, Lipzen A, Yoshinaga Y, Schmutz J, Saski C, Vermerris W, Kresovich S (2019) A new reference genome for Sorghum bicolor reveals high levels of sequence similarity between sweet and grain genotypes: implications for the genetics of sugar metabolism. BMC Genomics 20(1):420. https://doi.org/10.1186/s12864-019-5734-x
De Wet JMJ, Harlan JR (1971) The origin and domestication of sorghum bicolor. Econ Bot 25:128–135. https://doi.org/10.1007/bf02860074
De Wet JMJ, Huckabay JP (1967) The origin of sorghum bicolor. II. distribution and domestication. Evolution 21(4):787–802. https://doi.org/10.1111/j.1558-5646.1967.tb03434.x
De Wet JMJ, Shechter Y (1977) Evolutionary dynamics of sorghum domestication. In: Seigler DS (ed) Crop resources. Academic Press, New York, pp 179–191
De Wet JMJ, Harlan JR, Kurmarohita B (1972) Origin and evolution of guinea sorghums. East African Agricultural and Forestry Journal 38(2):114–119. https://doi.org/10.1080/00128325.1972.11662567
Deschamps S, Zhang Y, Llaca V, Ye L, Sanyal A, King M, May G, Lin H (2018) A chromosome-scale assembly of the sorghum genome using nanopore sequencing and optical mapping. Nat Commun 9(1):4844. https://doi.org/10.1038/s41467-018-07271-1
Doebley J, Stec A, Hubbard L (1997) The evolution of apical dominance in maize. Nature 386(6624):485–488. https://doi.org/10.1038/386485a0
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127(7):1309–1321. https://doi.org/10.1016/j.cell.2006.12.006
Du D, Zhang D, Yuan J, Feng M, Li Z, Wang Z, Zhang Z, Li X, Ke W, Li R, Chen Z, Chai L, Hu Z, Guo W, Xing J, Su Z, Peng H, Xin M, Yao Y, Sun Q, Liu J, Ni Z (2021) FRIZZY PANICLE defines a regulatory hub for simultaneously controlling spikelet formation and awn elongation in bread wheat. New Phytol 231(2):814–833. https://doi.org/10.1111/nph.17388
Elbaum R, Zaltzman L, Burgert I, Fratzl P (2007) The role of wheat awns in the seed dispersal unit. Science 316(5826):884–886. https://doi.org/10.1126/science.1140097
Fernie AR, Yan J (2019) De novo domestication: an alternative route toward new crops for the future. Mol Plant 12(5):615–631. https://doi.org/10.1016/j.molp.2019.03.016
Fujimoto M, Sazuka T, Oda Y, Kawahigashi H, Wu J, Takanashi H, Ohnishi T, Yoneda JI, Ishimori M, Kajiya-Kanegae H, Hibara KI, Ishizuna F, Ebine K, Ueda T, Tokunaga T, Iwata H, Matsumoto T, Kasuga S, Yonemaru JI, Tsutsumi N (2018) Transcriptional switch for programmed cell death in pith parenchyma of sorghum stems. Proc Natl Acad Sci U S A 115(37):E8783–E8792. https://doi.org/10.1073/pnas.1807501115
Fuller DQ, Stevens CJ (2018) Sorghum domestication and diversification: a current archaeobotanical perspective. In: Mercuri A, D'Andrea A, Fornaciari R, Höhn A (eds) Plants and people in the African past. Springer Cham, Switzerland, pp 427–452
Gaut BS, Seymour DK, Liu Q, Zhou Y (2018) Demography and its effects on genomic variation in crop domestication. Nat Plants 4(8):512–520. https://doi.org/10.1038/s41477-018-0210-1
Gepts P (2014) Domestication of plants. In: Van Alfen NK (ed) Encyclopedia of agriculture and food systems. Academic Press, New York, pp 474–486
Girma G, Nida H, Seyoum A, Mekonen M, Nega A, Lule D, Dessalegn K, Bekele A, Gebreyohannes A, Adeyanju A, Tirfessa A, Ayana G, Taddese T, Mekbib F, Belete K, Tesso T, Ejeta G, Mengiste T (2019) A large-scale genome-wide association analyses of Ethiopian sorghum landrace collection reveal loci associated with important traits. Front Plant Sci 10:691. https://doi.org/10.3389/fpls.2019.00691
Groppi A, Liu S, Cornille A, Decroocq S, Bui QT, Tricon D, Cruaud C, Arribat S, Belser C, Marande W, Salse J, Huneau C, Rodde N, Rhalloussi W, Cauet S, Istace B, Denis E, Carrere S, Audergon JM, Roch G, Lambert P, Zhebentyayeva T, Liu WS, Bouchez O, Lopez-Roques C, Serre RF, Debuchy R, Tran J, Wincker P, Chen X, Petriacq P, Barre A, Nikolski M, Aury JM, Abbott AG, Giraud T, Decroocq V (2021) Population genomics of apricots unravels domestication history and adaptive events. Nat Commun 12(1):3956. https://doi.org/10.1038/s41467-021-24283-6
Gu B, Zhou T, Luo J, Liu H, Wang Y, Shangguan Y, Zhu J, Li Y, Sang T, Wang Z, Han B (2015) An-2 encodes a cytokinin synthesis enzyme that regulates awn length and grain production in rice. Mol Plant 8(11):1635–1650. https://doi.org/10.1016/j.molp.2015.08.001
Han Y, Lv P, Hou S, Li S, Ji G, Ma X, Du R, Liu G (2015) Combining next generation sequencing with bulked segregant analysis to fine map a stem moisture locus in sorghum (Sorghum bicolor L. Moench). PLoS ONE 10(5):0127065. https://doi.org/10.1371/journal.pone.0127065
Hao H, Li Z, Leng C, Lu C, Luo H, Liu Y, Wu X, Liu Z, Shang L, Jing HC (2021) Sorghum breeding in the genomic era: opportunities and challenges. Theor Appl Genet 134(7):1899–1924. https://doi.org/10.1007/s00122-021-03789-z
Harlan JR, de Wet JMJ (1972) A simplified classification of cultivated sorghum. Crop Sci 12(2):172–176. https://doi.org/10.2135/cropsci1972.0011183x001200020005x
Harlan JR, Stemler ABL (1976) The races of sorghum in Africa. In: Harlan JR, De Wet JMJ, Stemler ABL (eds) Origins of African plant domestication. De Gruyter Mouton, New York, pp 465–478
Hart GE, Schertz KF, Peng Y, Syed NH (2001) Genetic mapping of Sorghum bicolor (L.) Moench QTLs that control variation in tillering and other morphological characters. Theor Appl Genet 103:1232–1242. https://doi.org/10.1007/s001220100582
Hilley J, Truong S, Olson S, Morishige D, Mullet J (2016) Identification of Dw1, a regulator of sorghum stem internode length. PLoS ONE 11(3):e0151271. https://doi.org/10.1371/journal.pone.0151271
Hilley JL, Weers BD, Truong SK, McCormick RF, Mattison AJ, McKinley BA, Morishige DT, Mullet JE (2017) Sorghum Dw2 encodes a protein kinase regulator of stem internode length. Sci Rep 7(1):4616. https://doi.org/10.1038/s41598-017-04609-5
Hua L, Wang DR, Tan L, Fu Y, Liu F, Xiao L, Zhu Z, Fu Q, Sun X, Gu P, Cai H, McCouch SR, Sun C (2015) LABA1, a domestication gene associated with long, barbed awns in wild rice. Plant Cell 27(7):1875–1888. https://doi.org/10.1105/tpc.15.00260
Huang X, Huang S, Han B, Li J (2022) The integrated genomics of crop domestication and breeding. Cell 185(15):2828-2839. https://doi.org/10.1016/j.cell.2022.04.036
Jagathesan D, Bhatia C, Swaminathan MS (1961) Effect of induced awn mutations on yield in wheat. Nature 190(477):468. https://doi.org/10.1038/190468a0
Kebrom TH, Burson BL, Finlayson SA (2006) Phytochrome B represses Teosinte Branched1 expression and induces sorghum axillary bud outgrowth in response to light signals. Plant Physiol 140(3):1109–1117. https://doi.org/10.1104/pp.105.074856
Klein RR, Miller FR, Dugas DV, Brown PJ, Burrell AM, Klein PE (2015) Allelic variants in the PRR37 gene and the human-mediated dispersal and diversification of sorghum. Theor Appl Genet 128(9):1669–1683. https://doi.org/10.1007/s00122-015-2523-z
Larson G, Piperno DR, Allaby RG, Purugganan MD, Andersson L, Arroyo-Kalin M, Barton L, Climer Vigueira C, Denham T, Dobney K, Doust AN, Gepts P, Gilbert MT, Gremillion KJ, Lucas L, Lukens L, Marshall FB, Olsen KM, Pires JC, Richerson PJ, Rubio de Casas R, Sanjur OI, Thomas MG, Fuller DQ (2014) Current perspectives and the future of domestication studies. Proc Natl Acad Sci U S A 111(17):6139–6146. https://doi.org/10.1073/pnas.1323964111
Lenser T, Theissen G (2013) Molecular mechanisms involved in convergent crop domestication. Trends Plant Sci 18(12):704–714. https://doi.org/10.1016/j.tplants.2013.08.007
Li X, Yu J (2022) Unraveling the sorghum domestication. Mol Plant 15(5):791–792. https://doi.org/10.1016/j.molp.2022.03.006
Li C, Zhou A, Sang T (2006) Rice domestication by reducing shattering. Science 311(5769):1936–1939. https://doi.org/10.1126/science.1123604
Li T, Yang X, Yu Y, Si X, Zhai X, Zhang H, Dong W, Gao C, Xu C (2018) Domestication of wild tomato is accelerated by genome editing. Nat Biotechnol 36:1160–1163. https://doi.org/10.1038/nbt.4273
Lin Z, Li X, Shannon LM, Yeh CT, Wang ML, Bai G, Peng Z, Li J, Trick HN, Clemente TE, Doebley J, Schnable PS, Tuinstra MR, Tesso TT, White F, Yu J (2012) Parallel domestication of the Shattering1 genes in cereals. Nat Genet 44(6):720–724. https://doi.org/10.1038/ng.2281
Lin T, Zhu G, Zhang J, Xu X, Yu Q, Zheng Z, Zhang Z, Lun Y, Li S, Wang X, Huang Z, Li J, Zhang C, Wang T, Zhang Y, Wang A, Zhang Y, Lin K, Li C, Xiong G, Xue Y, Mazzucato A, Causse M, Fei Z, Giovannoni JJ, Chetelat RT, Zamir D, Stadler T, Li J, Ye Z, Du Y, Huang S (2014) Genomic analyses provide insights into the history of tomato breeding. Nat Genet 46(11):1220–1226. https://doi.org/10.1038/ng.3117
Lu S, Dong L, Fang C, Liu S, Kong L, Cheng Q, Chen L, Su T, Nan H, Zhang D, Zhang L, Wang Z, Yang Y, Yu D, Liu X, Yang Q, Lin X, Tang Y, Zhao X, Yang X, Tian C, Xie Q, Li X, Yuan X, Tian Z, Liu B, Weller JL, Kong F (2020) Stepwise selection on homeologous PRR genes controlling flowering and maturity during soybean domestication. Nat Genet 52(4):428–436. https://doi.org/10.1038/s41588-020-0604-7
Lyzenga WJ, Pozniak CJ, Kagale S (2021) Advanced domestication: harnessing the precision of gene editing in crop breeding. Plant Biotechnol J 19(4):660–670. https://doi.org/10.1111/pbi.13576
Ma S, Lv L, Meng C, Zhang C, Li Y (2020) Integrative analysis of the metabolome and transcriptome of sorghum bicolor reveals dynamic changes in flavonoids accumulation under saline-alkali stress. J Agric Food Chem 68(50):14781–14789. https://doi.org/10.1021/acs.jafc.0c06249
McCormick RF, Truong SK, Sreedasyam A, Jenkins J, Shu S, Sims D, Kennedy M, Amirebrahimi M, Weers BD, McKinley B, Mattison A, Morishige DT, Grimwood J, Schmutz J, Mullet JE (2018) The sorghum bicolor reference genome: improved assembly, gene annotations, a transcriptome atlas, and signatures of genome organization. Plant J 93(2):338–354. https://doi.org/10.1111/tpj.13781
Multani DS, Briggs SP, Chamberlin MA, Blakeslee JJ, Murphy AS, Johal GS (2003) Loss of an MDR transporter in compact stalks of maize br2 and sorghum dw3 mutants. Science 302(5642):81–84. https://doi.org/10.1126/science.1086072
Murphy RL, Klein RR, Morishige DT, Brady JA, Rooney WL, Miller FR, Dugas DV, Klein PE, Mullet JE (2011) Coincident light and clock regulation of pseudoresponse regulator protein 37 (PRR37) controls photoperiodic flowering in sorghum. Proc Natl Acad Sci U S A 108(39):16469–16474. https://doi.org/10.1073/pnas.1106212108
Murphy RL, Morishige DT, Brady JA, Rooney WL, Yang S, Klein PE, Mullet JE (2014) Ghd7 (Ma6) represses sorghum flowering in long days: Ghd7 alleles enhance biomass accumulation and grain production. Plant Gen 7(2). https://doi.org/10.3835/plantgenome2013.11.0040
Nesi N, Debeaujon I, Jond C, Pelletier G, Caboche M, Lepiniec L (2000) The TT8 gene encodes a basic helix-loop-helix domain protein required for expression of DFR and BAN genes in Arabidopsis siliques. Plant Cell 12(10):1863–1878. https://doi.org/10.1105/tpc.12.10.1863
Ohadi S, Hodnett G, Rooney W, Bagavathiannan M (2018) Gene flow and its consequences in Sorghum spp. Crit Rev Plant Sci 36(5–6):367–385. https://doi.org/10.1080/07352689.2018.1446813
Oliver J, Fan M, McKinley B, Zemelis-Durfee S, Brandizzi F, Wilkerson C, Mullet JE (2021) The AGCVIII kinase Dw2 modulates cell proliferation, endomembrane trafficking, and MLG/xylan cell wall localization in elongating stem internodes of Sorghum bicolor. Plant J 105(4):1053–1071. https://doi.org/10.1111/tpj.15086
Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, Gundlach H, Haberer G, Hellsten U, Mitros T, Poliakov A, Schmutz J, Spannagl M, Tang H, Wang X, Wicker T, Bharti AK, Chapman J, Feltus FA, Gowik U, Grigoriev IV, Lyons E, Maher CA, Martis M, Narechania A, Otillar RP, Penning BW, Salamov AA, Wang Y, Zhang L, Carpita NC, Freeling M, Gingle AR, Hash CT, Keller B, Klein P, Kresovich S, McCann MC, Ming R, Peterson DG, Mehboob Ur R, Ware D, Westhoff P, Mayer KF, Messing J, Rokhsar DS (2009) The sorghum bicolor genome and the diversification of grasses. Nature 457(7229):551–556. https://doi.org/10.1038/nature07723
Prasad VBR, Govindaraj M, Djanaguiraman M, Djalovic I, Shailani A, Rawat N, Singla-Pareek SL, Pareek A, Prasad PVV (2021) Drought and high temperature stress in sorghum: physiological, genetic, and molecular insights and breeding approaches. Int J Mol Sci 22(18):9826. https://doi.org/10.3390/ijms22189826
Purugganan MD (2019) Evolutionary insights into the nature of plant domestication. Curr Biol 29(14):R705–R714. https://doi.org/10.1016/j.cub.2019.05.053
Purugganan MD, Fuller DQ (2009) The nature of selection during plant domestication. Nature 457(7231):843–848. https://doi.org/10.1038/nature07895
Qin P, Lu H, Du H, Wang H, Chen W, Chen Z, He Q, Ou S, Zhang H, Li X, Li X, Li Y, Liao Y, Gao Q, Tu B, Yuan H, Ma B, Wang Y, Qian Y, Fan S, Li W, Wang J, He M, Yin J, Li T, Jiang N, Chen X, Liang C, Li S (2021) Pan-genome analysis of 33 genetically diverse rice accessions reveals hidden genomic variations. Cell 184(13):3542–3558. https://doi.org/10.1016/j.cell.2021.04.046
Quinby JR (1966) Fourth maturity gene locus in sorghum. Crop Sci 6(6):516–518. https://doi.org/10.2135/cropsci1966.0011183X000600060005x
Quinby JR (1967) The maturity genes of sorghum. Adv Agron 19:267–305. https://doi.org/10.1016/S0065-2113(08)60737-3
Quinby J, Karper R (1945) The inheritance of three genes that influence time of floral initiation and maturity date in Milo1. Agron J 37(11):916–936. https://doi.org/10.2134/agronj1945.00021962003700110006x
Quinby JR, Martin JH (1954) Sorghum improvement. In: Norman AG (ed) Advances in agronomy. Academic Press, New York, pp 305–359
Ramsay NA, Glover BJ (2005) MYB–bHLH–WD40 protein complex and the evolution of cellular diversity. Trends Plant Sci 10(2):63–70. https://doi.org/10.1016/j.tplants.2004.12.011
Rooney WL, Aydin S (1999) Genetic control of a photoperiod-sensitive response in sorghum bicolor (L.) Moench. Crop Sci 39(2):397–400. https://doi.org/10.2135/cropsci1999.0011183X0039000200016x
Ross-Ibarra J, Morrell PL, Gaut BS (2007) Plant domestication, a unique opportunity to identify the genetic basis of adaptation. Proc Natl Acad Sci U S A 104(Suppl 1):8641–8648. https://doi.org/10.1073/pnas.0700643104
Ruperao P, Thirunavukkarasu N, Gandham P, Selvanayagam S, Govindaraj M, Nebie B, Manyasa E, Gupta R, Das RR, Odeny DA, Gandhi H, Edwards D, Deshpande SP, Rathore A (2021) Sorghum pan-genome explores the functional utility for genomic-assisted breeding to accelerate the genetic gain. Front Plant Sci 12:666342. https://doi.org/10.3389/fpls.2021.666342
Shechter Y, de Wet JMJ (1975) Comparative electrophoresis and isozyme analysis of seed proteins from cultivated races of sorghum. Am J Bot 62(3):254–261. https://doi.org/10.2307/2441859
Silva TN, Thomas JB, Dahlberg J, Rhee SY, Mortimer JC (2021) Progress and challenges in sorghum biotechnology, a multi-purpose feedstock for the bioeconomy. J Exp Bot 73(3):646–664. https://doi.org/10.1093/jxb/erab450
Smith CW, Frederiksen RA (2000) Sorghum: Origin, History, Technology, and Production. Wiley, New York
Smith O, Nicholson WV, Kistler L, Mace E, Clapham A, Rose P, Stevens C, Ware R, Samavedam S, Barker G, Jordan D, Fuller DQ, Allaby RG (2019) A domestication history of dynamic adaptation and genomic deterioration in sorghum. Nat Plants 5(4):369–379. https://doi.org/10.1038/s41477-019-0397-9
Stetter MG, Gates DJ, Mei W, Ross-Ibarra J (2017) How to make a domesticate. Curr Biol 27(17):R896–R900. https://doi.org/10.1016/j.cub.2017.06.048
Swanson AF, Parker JH (1931) Inheritance of smut resistance and juiciness of stalk: in the sorghum cross. Red Amber X Feterita J Hered 22(2):51–56. https://doi.org/10.1093/oxfordjournals.jhered.a103429
Tang H, Cuevas HE, Das S, Sezen UU, Zhou C, Guo H, Goff VH, Ge Z, Clemente TE, Paterson AH (2013) Seed shattering in a wild sorghum is conferred by a locus unrelated to domestication. Proc Natl Acad Sci U S A 110(39):15824–15829. https://doi.org/10.1073/pnas.1305213110
Tao Y, Luo H, Xu J, Cruickshank A, Zhao X, Teng F, Hathorn A, Wu X, Liu Y, Shatte T, Jordan D, Jing H, Mace E (2021) Extensive variation within the pan-genome of cultivated and wild sorghum. Nat Plants 7(6):766–773. https://doi.org/10.1038/s41477-021-00925-x
Turck F, Fornara F, Coupland G (2008) Regulation and identity of florigen: FLOWERING LOCUS T moves center stage. Annu Rev Plant Biol 59:573–594. https://doi.org/10.1146/annurev.arplant.59.032607.092755
Upadhyaya HD, Wang YH, Gowda CL, Sharma S (2013) Association mapping of maturity and plant height using SNP markers with the sorghum mini core collection. Theor Appl Genet 126(8):2003–2015. https://doi.org/10.1007/s00122-013-2113-x
Upadhyaya HD, Wang L, Prakash CS, Liu Y, Gao L, Meng R, Seetharam K, Gowda CLL, Ganesamurthy K, Singh SK, Kumar R, Li J, Wang YH (2022) Genome-wide association mapping identifies an SNF4 ortholog that impacts biomass and sugar yield in sorghum and sugarcane. J Exp Bot 73(11):3584–3596. https://doi.org/10.1093/jxb/erac110
Varoquaux N, Cole B, Gao C, Pierroz G, Baker CR, Patel D, Madera M, Jeffers T, Hollingsworth J, Sievert J, Yoshinaga Y, Owiti JA, Singan VR, DeGraaf S, Xu L, Blow MJ, Harrison MJ, Visel A, Jansson C, Niyogi KK, Hutmacher R, Coleman-Derr D, O’Malley RC, Taylor JW, Dahlberg J, Vogel JP, Lemaux PG, Purdom E (2019) Transcriptomic analysis of field-droughted sorghum from seedling to maturity reveals biotic and metabolic responses. Proc Natl Acad Sci U S A 116(52):27124–27132. https://doi.org/10.1073/pnas.1907500116
Venkateswaran K, Elangovan M, Sivaraj N (2019) Origin, domestication and diffusion of Sorghum bicolor. In: Aruna C, Visarada KBRS, Bhat BV, Tonapi VA (eds) Breeding sorghum for diverse end uses. Woodhead Publishing, Duxford, pp 15–31
Wallace JG, Rodgers-Melnick E, Buckler ES (2018) On the road to breeding 4.0: unraveling the good, the bad, and the boring of crop quantitative genomics. Annu Rev Genet 52:421–444. https://doi.org/10.1146/annurev-genet-120116-024846
Wang F, Yoshida H, Matsuoka M (2021) Making the “Green Revolution” truly green: improving crop nitrogen use efficiency. Plant Cell Physiol 62(6):942–947. https://doi.org/10.1093/pcp/pcab051
Wei T, van Treuren R, Liu X, Zhang Z, Chen J, Liu Y, Dong S, Sun P, Yang T, Lan T, Wang X, Xiong Z, Liu Y, Wei J, Lu H, Han S, Chen JC, Ni X, Wang J, Yang H, Xu X, Kuang H, van Hintum T, Liu X, Liu H (2021) Whole-genome resequencing of 445 Lactuca accessions reveals the domestication history of cultivated lettuce. Nat Genet 53(5):752–760. https://doi.org/10.1038/s41588-021-00831-0
Winchell F, Stevens CJ, Murphy C, Champion L, Fuller D (2017) Evidence for sorghum domestication in fourth millennium BC eastern Sudan: spikelet morphology from ceramic impressions of the Butana group. Curr Anthropol 58(5):673–683. https://doi.org/10.1086/693898
Winchell F, Brass M, Manzo A, Beldados A, Perna V, Murphy C, Stevens C, Fuller DQ (2018) On the origins and dissemination of domesticated sorghum and pearl millet across Africa and into India: a view from the Butana group of the far eastern Sahel. Afr Archaeol Rev 35(4):483–505. https://doi.org/10.1007/s10437-018-9314-2
Wu Y, Li X, Xiang W, Zhu C, Lin Z, Wu Y, Li J, Pandravada S, Ridder DD, Bai G, Wang ML, Trick HN, Bean SR, Tuinstra MR, Tesso TT, Yu J (2012) Presence of tannins in sorghum grains is conditioned by different natural alleles of Tannin1. Proc Natl Acad Sci U S A 109(26):10281–10286. https://doi.org/10.1073/pnas.1201700109
Wu Y, Guo T, Mu Q, Wang J, Li X, Wu Y, Tian B, Wang ML, Bai G, Perumal R, Trick HN, Bean SR, Dweikat IM, Tuinstra MR, Morris G, Tesso TT, Yu J, Li X (2019) Allelochemicals targeted to balance competing selections in African agroecosystems. Nat Plants 5(12):1229–1236. https://doi.org/10.1038/s41477-019-0563-0
Wu D, Lao S, Fan L (2021) De-domestication: an extension of crop evolution. Trends Plant Sci 26(6):560–574. https://doi.org/10.1016/j.tplants.2021.02.003
Wu X, Liu Y, Luo H, Shang L, Leng C, Liu Z, Li Z, Lu X, Cai H, Hao H, Jing HC (2022) Genomic footprints of sorghum domestication and breeding selection for multiple end uses. Mol Plant 15(3):537–551. https://doi.org/10.1016/j.molp.2022.01.002
Xie Q, Xu Z (2019) Sustainable agriculture: from sweet sorghum planting and ensiling to ruminant feeding. Mol Plant 12(5):603–606. https://doi.org/10.1016/j.molp.2019.04.001
Xie P, Shi J, Tang S, Chen C, Khan A, Zhang F, Xiong Y, Li C, He W, Wang G, Lei F, Wu Y, Xie Q (2019) Control of bird feeding behavior by Tannin1 through modulating the biosynthesis of polyphenols and fatty acid-derived volatiles in sorghum. Mol Plant 12(10):1315–1324. https://doi.org/10.1016/j.molp.2019.08.004
Xie P, Tang S, Chen C, Zhang H, Yu F, Li C, Wei H, Sui Y, Wu C, Diao X, Wu Y, Xie Q (2022) Natural variation in Glume Coverage 1 causes naked grains in sorghum. Nat Commun 13(1):1068. https://doi.org/10.1038/s41467-022-28680-3
Xin Z, Wang M, Cuevas HE, Chen J, Harrison M, Pugh NA, Morris G (2021) Sorghum genetic, genomic, and breeding resources. Planta 254(6):114. https://doi.org/10.1007/s00425-021-03742-w
Yamaguchi M, Fujimoto H, Hirano K, Araki-Nakamura S, Ohmae-Shinohara K, Fujii A, Tsunashima M, Song XJ, Ito Y, Nagae R, Wu J, Mizuno H, Yonemaru J, Matsumoto T, Kitano H, Matsuoka M, Kasuga S, Sazuka T (2016) Sorghum Dw1, an agronomically important gene for lodging resistance, encodes a novel protein involved in cell proliferation. Sci Rep 6:28366. https://doi.org/10.1038/srep28366
Yang S, Murphy RL, Morishige DT, Klein PE, Rooney WL, Mullet JE (2014a) Sorghum phytochrome B inhibits flowering in long days by activating expression of SbPRR37 and SbGHD7, repressors of SbEHD1, SbCN8 and SbCN12. PLoS ONE 9(8):e105352. https://doi.org/10.1371/journal.pone.0105352
Yang Z, Chi X, Guo F, Jin X, Luo H, Hawar A, Chen Y, Feng K, Wang B, Qi J, Yang Y, Sun B (2020) SbWRKY30 enhances the drought tolerance of plants and regulates a drought stress-responsive gene, SbRD19, in sorghum. J Plant Physiol 246–247:153142 https://doi.org/10.1016/j.jplph.2020.153142
Yang S, Weers BD, Morishige DT, Mullet JE (2014b) CONSTAN is a photoperiod regulated activator of flowering in sorghum. BMC Plant Biol 148(2014). https://doi.org/10.1186/1471-2229-14-148
Yu H, Lin T, Meng X, Du H, Zhang J, Liu G, Chen M, Jing Y, Kou L, Li X, Gao Q, Liang Y, Liu X, Fan Z, Liang Y, Cheng Z, Chen M, Tian Z, Wang Y, Chu C, Zuo J, Wan J, Qian Q, Han B, Zuccolo A, Wing RA, Gao C, Liang C, Li J (2021) A route to de novo domestication of wild allotetraploid rice. Cell 184(5):1156–1170. https://doi.org/10.1016/j.cell.2021.01.013
Zhai G, Zou G, Yan S, Wang H, Shao J, Tao Y (2014) Identification and fine mapping of the gene associated with moisture content of stem in sorghum[Sorghum bicolor (L.) Moench]. Acta Agriculturae Zhejiangensis 26(4):856–861
Zhang F, Ping J (2022) Origin, domestication and diffusion of sorghum. Shaanxi Journal of Agricultural Sciences 68(04):82–87
Zhang LM, Leng CY, Luo H, Wu XY, Liu ZQ, Zhang YM, Zhang H, Xia Y, Shang L, Liu CM, Hao DY, Zhou YH, Chu CC, Cai HW, Jing HC (2018) Sweet sorghum originated through selection of Dry, a plant-specific NAC transcription factor gene. Plant Cell 30(10):2286–2307. https://doi.org/10.1105/tpc.18.00313
Zhao Y, Feng M, Paudel D, Islam T, Momotaz A, Luo Z, Zhao Z, Wei N, Li S, Xia Q, Kuang B, Yang X, Wang J (2021) Advances in genomics approaches shed light on crop domestication. Plants (basel) 10(8):1571. https://doi.org/10.3390/plants10081571
Zhou L, Zhu C, Fang X, Liu H, Zhong S, Li Y, Liu J, Song Y, Jian X, Lin Z (2021) Gene duplication drove the loss of awn in sorghum. Mol Plant 14(11):1831–1845. https://doi.org/10.1016/j.molp.2021.07.005
Funding
This review was funded by the National Key R&D Program of China (2019YFD1002701), the National Natural Science Foundation of China (U1906204), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDA24010306), the National Science Foundation for Young Scientists of China (32201780) and the Agricultural Breeding Program in NingXia Province (2019NYYZ04 and 2019BBF02022-05).
Author information
Authors and Affiliations
Contributions
FG wrote this manuscript. The conceptualization and editing of this manuscript were done by PX, YW and QX.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest. Author Qi Xie was not involved in the journal’s review of the manuscript.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ge, F., Xie, P., Wu, Y. et al. Genetic architecture and molecular regulation of sorghum domestication. aBIOTECH 4, 57–71 (2023). https://doi.org/10.1007/s42994-022-00089-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42994-022-00089-y