Abstract
To obtain more semantic information with small samples for medical image segmentation, this paper proposes a simple and efficient dual-rotation network (DR-Net) that strengthens the quality of both local and global feature maps. The key steps of the DR-Net algorithm are as follows (as shown in Fig. 1). First, the number of channels in each layer is divided into four equal portions. Then, different rotation strategies are used to obtain a rotation feature map in multiple directions for each subimage. Then, the multiscale volume product and dilated convolution are used to learn the local and global features of feature maps. Finally, the residual strategy and integration strategy are used to fuse the generated feature maps. Experimental results demonstrate that the DR-Net method can obtain higher segmentation accuracy on both the CHAOS and BraTS data sets compared to the state-of-the-art methods.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Computer vision is widely used in medical tasks: medical image semantic segmentation [1,2,3], medical image classification [4,5,6], bioengineering recognition [7,8,9], three-dimensional reconstruction [10,11,12] and others. Although deep learning has achieved great success in the medical field due to the complex structure of medical images, the pathological tissue being similar to the surrounding normal tissue, and the number of data samples being small, it is difficult to obtain deeper semantic information. To further enhance a model's ability to learn features of medical images, researchers build various model strategies to mine deep features and obtain deep semantic information. Ni et al. [3] proposed a global context attention module to obtain the contextual semantic information of medical images and integrate the multiscale receptive field information generated by the SEPP model, thereby providing more learning information for segmentation tasks. The model achieved good segmentation results on three public data sets and one local data set; to better learn the subtle differences in the intensity, location, shape, and size of the lesions in medical images, Chen et al. [13] constructed a new convolutional neural network structure named DRINet. The network integrates multiple feature maps generated by dense network blocks [14], residual network blocks and inception network blocks [15] and acquires a variety of semantic information. Finally, the results show that DRINet is better than UNet in three challenging applications; Alom et al. [16] improved U-Net [17] into a joint model of a recursive U-Net and a residual U-Net. The model realizes the feature fusion of deep features and shallow features through the residual block and realizes the accumulation of the semantic information of the residual convolutional layer through the loop module. The algorithm obtained the best segmentation results on three data sets: retinal images, skin cancer segmentation, and lung lesion segmentation; Gu et al. [18] proposed a context encoder network (CE-Net). The network uses ResNet [19] to achieve feature extraction and uses dense convolution blocks and multilayer pooling blocks [20] to obtain richer image contextual semantic information. The results on 5 data sets prove the feasibility of the model (Fig. 1).
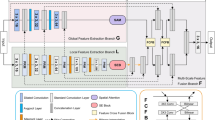
Outer rotation and inner rotation of DR-Net. The reds line represent internal rotation, and the blue lines represent the external rotation. The blue squares mean that the same feature maps are used, where the numbers indicate the direction of rotation. The green squares indicate that the feature map is rotated, and the local semantic features and global semantic features are obtained using the FPM and DPFM (a negative sign indicates reverse rotation)
The above research results show that different model strategies can obtain different semantic information to generate different feature maps (for example, shallow feature maps, deep feature maps, local feature maps and global feature maps). The above strategies can only generate one or two feature maps, which are insufficiently comprehensive to obtain the semantic information of medical images. Although more semantic information can be obtained through the feature maps generated after the fusion of multiple strategies, the total number of parameters will also increase, which greatly enhances the computational complexity of the algorithm. Furthermore, the fusion of the feature maps of each strategy is only established between the deep feature maps of each strategy, making it impossible to obtain other semantic information in the process of feature mining from shallow to deep; and the feature maps generated are insufficiently rich.
The emergence of the group convolution allows researchers to obtain richer semantic information without increasing network parameters. The ResNeXt algorithm proposed by Xie et al. [21] converts high-dimensional feature maps into multiple low-dimensional feature maps and then obtains deep features by learning the low-dimensional feature maps. The advantage of converting high-dimensional feature maps to low-dimensional feature maps is that the number of parameter calculations can be reduced. Their algorithm obtained good results on the ImageNet-1 k data set, thus proving that group convolution has good feature learning capabilities. Romero et al. [22] introduced an attention strategy based on the group convolution, and the constructed attention group convolution can enhance the feature maps. The above research finds that the group convolution can improve a network’s feature learning ability. In the previous work, the researchers increased the number of training samples by preprocessing the images [23, 24], thereby improving the testing accuracy on the data set.
To obtain richer semantic information without increasing the complexity of the algorithm and reducing the preprocessing of the data, this paper proposes a faithful deep learning algorithm: the double rotation network (DR-Net).
The main contributions of this paper are as follows:
-
1.
We divide the generated feature maps of the previous layer into four equal channel parts according to the total number of channels. Then, each part is rotated according to different rotation angles (the rotation angles are 0°, 90°, 180°, and 270°) to achieve the internal rotation of the traditional convolution. Then, different local semantic information is obtained from the rotated feature maps through different sized convolution kernels so as to generate deeper local feature maps. Here, we call these feature maps partial feature maps (PFMs).
-
2.
We also divide the generated feature maps of the previous layer into four parts with equal numbers of channels according to the total number of channels. Subsequently, we also rotated each part in four directions. The feature mining method that is different from the previous method is that we use the dilated convolution to obtain different receptive domains by setting different expansion values so as to obtain more global semantic information. The size of the convolution kernel of all our dilated convolutions is set to 3. Here, we call these feature maps dilated partial feature maps (DPFMs).
-
3.
To further obtain richer semantic information, we fuse the features of the PFM and DPFM in the same rotation direction (through the feature map addition strategy) and finally obtain 16 sets of richer feature maps in each layer of the DR-Net algorithm (these feature maps constitute the outer rotation).
-
4.
To obtain the fusion feature maps of the shallow feature maps and the deep feature maps and simultaneously obtain more semantic information, the shallow feature maps of the previous submodule are obtained. We use the transfer invariance of maximum pooling to compress the feature maps. Finally, the three strategies of the rotation angle, multiscale convolution and different dilation step sizes are used to generate feature maps that describe the semantic information of shallow features, the semantic information of the deep features, the semantic information of the local features and the semantic information of the global features.
The main content of the remainder of this paper is summarized as follows. In the second section, we mainly introduce the background and significance of the algorithm of this paper. In the third section, we introduce the overall flow of the DR-Net algorithm and the functions of each module in detail. In the fourth section, we verify the performance of the DR-Net algorithm through related experiments. In the fifth section, we give a summary of this paper and further research goals.
Related work
Feature maps are defined by their width, height, and number of channels. In recent years, researchers have used the number of channels as a benchmark to enhance the relationship between feature maps. For example, Xie et al. [21] et al. used the group convolution, Chollet [25] used the separable convolution, and You et al. [26] proposed using part of the feature maps. Although the names of these methods are different, they are all based on addressing the number of channels to enhance the quality of feature maps or reduce the number of parameter calculations. Next, we will analyze the pros and cons of the above three sets of strategies in detail.
Group convolution (GC)
The principle of the group convolution is to use a 1 × 1 convolution kernel to compress the number of features from the original feature maps (256 channels) and generate 32 sets of new feature maps with 4 channels. The channel compression strategy has also been verified in previous work [27,28,29,30]. The purpose of this method is to reduce the redundant information of deep feature maps and reduce the number of parameter calculations. Subsequently, the previous 32 sets of feature maps are mined by convolution with a convolution kernel of 3, which reduces the computational complexity of features. Then, the compressed feature maps are restored by convolution with a convolution kernel of 1 (the feature maps with 4 channels are restored to feature maps with the original number of channels of 256). Finally, feature maps are fused through the aggregate residual exchange strategy. The subsequent Res2NeXt network proposed by Gao et al. [31] also proved the efficiency of the group convolution.
Deep separable convolution (DSC)
The principle of the depth separable convolution is to divide the original feature map into N subchannels and implement feature mining through N convolution kernels to generate intermediate feature maps. The number of feature maps with the original number of feature channels is restored by splicing the intermediate feature maps. Subsequently, the spliced intermediate feature maps are compressed by a 1 × 1 convolution to compress the feature maps, and N 1 × 1 convolutions are simultaneously constructed to restore the feature maps.
Multidirectional integrated convolution (MDIC)
The core idea of the algorithm is to divide the original feature maps into four groups of subfeature maps, flip each subfeature map in different directions, and extract the feature maps after flipping through multiscale features. The algorithm obtains a variety of semantic information through the above strategies to enhance the feature maps. As a result, even with a small number of fitters, higher-precision segmentation results can be obtained. In the end, the total number of parameters of 5.2 million is much smaller than those of other models.
Our strategy
Our proposed model obtains various local semantic information through the PFM and simultaneously obtains various global semantic information through the DPFM. The acquired semantic information is richer than that of the abovementioned algorithms. For deep feature maps and shallow feature maps, this paper introduces a new residual strategy. This strategy compresses and expands feature maps. The semantic information of the feature maps of the upper layer is merged in the encoder. The decoder not only obtains the feature maps of the corresponding encoder but also the feature maps of the upper layer decoder. The entire submodule obtains richer semantic information.
Based on previous research, the main purpose of DR-Net algorithm is to enhance features while reducing the number of parameter calculations. Next, we use Table 1 to directly compare the number of parameters of each group convolution module and the type of feature maps obtained. "Shallow" represents shallow feature maps, "Deep" represents deep feature maps, "Local" represents local feature maps, "Global" represents the global feature maps, and "C" represents the traditional convolution.
All blocks are implemented on a convolution. "M" represents the size of the feature maps. The size of the convolution kernel is uniformly set to 3, and the sizes of the input and output fitters are set to 256. Table 1 shows that the total number of parameters of the PFM module we proposed is the lowest. The DPFM module has a similar number of parameters as the GC module and DSC module. Due to the addition strategy of the PFM and DPFM, the number of calculations of all parameters in this part is minimal.
Methodology
In this section, we will introduce our proposed DR-Net algorithm in detail through four subsections. The next section describes the framework of the entire model, the following section describes the details of the PFM and DPFM, the next section describes the different strategies used in the encoder and decoder, and the following section introduces the environmental details of the DR-Net algorithm (Fig. 2).
Illustration of the DR-Net model. Both the encoder and the decoder are composed of 4 sets of fusion submodules (PFM and DPFM) and a set of dual convolutional layers with a convolution kernel equal to 5. The orange lines represent the feature maps generated by the PFM; the blue lines represent the feature maps generated by the DPFM; the numbers in the circle represent different parts of the feature maps and also include the addition strategy; the symbol  denotes the feature map concatenation operation; \(\oplus\) represents the fusion strategy of different submodule feature maps; \(I\) and \(O\) represent the numbers of fitters in each layer, and the numbers of fitters in different layers are different. In both PFM and DPFM modules, the feature maps are rotated; and the rotation angles are 0, 90, 180, 270 in turn. The number of feature map fitters of the layer 1–5 submodules in the encoder are respectively [24, 48, 96, 192, 192], and each layer of the decoder corresponds to the fitters of the encoder
denotes the feature map concatenation operation; \(\oplus\) represents the fusion strategy of different submodule feature maps; \(I\) and \(O\) represent the numbers of fitters in each layer, and the numbers of fitters in different layers are different. In both PFM and DPFM modules, the feature maps are rotated; and the rotation angles are 0, 90, 180, 270 in turn. The number of feature map fitters of the layer 1–5 submodules in the encoder are respectively [24, 48, 96, 192, 192], and each layer of the decoder corresponds to the fitters of the encoder
DR-Net
Figure 1 shows the overall structure of our proposed DR-Net algorithm. The DR-Net algorithm is composed of two inner rotation convolutions (traditional convolution and dilated convolution) from the beginning of the second-layer encoder submodule to the end of the penultimate layer decoder submodule. Let
and
generate 32 sets of new feature maps through multidirection and multiscale feature mining and use these 32 sets of new feature maps to form 16 sets of deep feature maps containing global semantic information and local language information. Here, \(n\) represents the different parts, \(d\) represents the rotation angle, \(k\) represents the size of the convolution kernel, and \(D\) represents the dilation values. Its purpose is to reduce the number of feature maps while achieving strengthening the feature maps. In the encoder part, to further improve the quality of the feature maps \(Y(x)\), the feature maps generated by the current submodule layer \(H(x)\) and the multirotation direction feature maps \(Md(x)\) generated by the submodule of the previous layer are feature fused by the residual operation.
where \({\text{Conv}}1\) represents that the size of the convolution kernel is equal to 1, \({\text{Maxp}}\) represents maximum pooling, and \({\text{Sin}} p\{ 1,1\}\) represents the input and direction of the first set of features of the previous layer of feature maps.
In the decoder, the feature maps \(F(x)\) generated by the submodule of the current layer not only obtain the prior information of the feature maps \(Y(x)\) corresponding to the encoder but also incorporate the feature maps generated by the submodule of the previous layer. Through the above two feature fusion strategies of the encoder and decoder, the feature maps of each layer contain more semantic information of medical images, and finally the multiclassification task is completed through the softmax function.
PFMs and DPFMs
Partial feature maps (PFMs)
In this internal rotation, we divided the four groups of PFMs according to the total number of channels and rotated each group of PFMs in different directions. Even when using the same convolution kernel, different directions of feature maps can be obtained semantic information [26]. To further obtain more semantic information, in this part, we introduced the multiscale convolution in PFMs and finally generated 16 sets of feature maps with different semantic information. Through the above multidirection and multiscale convolution, a large number of rich local feature maps is obtained. Although we divide the total number of channels into multiple groups of PFMs to reduce the number of parameter calculations, to further reduce the number of parameter calculations, we have performed feature map compression on the feature maps. The specific total number of calculations of the local feature map parameters is as follows:
where \(N_{{{\text{PFM}}}}\) represents the total number of parameters, \(M\) represents the scale of the feature map, \(k\) represents the size of the convolution kernel, \(I\) represents the total number of channels of the input feature map, and \(O\) represents the total number of channels of the output feature map.
Dilated partial feature maps (DPFMs)
The DPFM module is similar in structure to the PFM module. The difference is that we replace the traditional convolution with the dilated convolution. The purpose is to obtain different receptive fields through different expansion coefficients and finally obtain more global semantic information. We use different dilated convolution structures in the last two sets of submodules of the encoder and the first two sets of submodules in the decoder. The purpose is to obtain a more valuable feature map through the double-layer dilated convolution as the scale of the feature map decreases. The structure of the two groups of different cavity convolution modules is shown in Fig. 3. The total number of computations with global feature map parameters is as follows:
The network structure of the different submodules of the DPFM. a The structure of the first two layers of the DPFM, and b the structure of the latter two layers of the DPFM. The dotted lines represent different directions in the DPFM module, but the network structure is the same. (Each dimension in the convolution represents the following: the number of input feature maps, the size of the convolution kernel, the number of output feature maps, and the scale of dilation)
Encoder and decoder
The encoder and decoder of DR-Net are composed of 5 submodules. To further fully obtain the feature maps of different semantic information, we combine the second submodule with the fifth submodule in the encoder and the first submodule with the fourth submodule in the decoder to be composed of the PFM and DPFM strategies. Different from the U-Net modes [17, 32, 33], in the encoder part, only the features of the current layer are mined. Furthermore, only the prior feature map of the corresponding encoder is included in the decoder part. In addition, different from a network model such as MC-Net [20], the semantic information of the encoder is increased by constructing multiple scales, and more information is obtained by fusing the different scales of semantic information. Our proposed DR-Net not only uses multidirection and multiscale strategies to obtain more feature maps. In the encoder part, the method also integrates the local feature maps generated by the PFM module and the global feature maps generated by the DPFM module to expand the experience of each feature point. The input feature maps use the feature invariance of the maximum pooling layer to introduce more semantic information of the submodules of the upper layer to the current layer. In the decoder part, in the current previous layer, we not only introduce the feature maps output by the corresponding layer encoder but also reference the feature maps input by the upper layer of the decoder so that more semantic information can be obtained. The different structure models of the encoder and decoder are shown in Fig. 4.
Model diagram of the DR-Net encoder and decoder. a The model structure of the second to fifth submodules of the encoder, and b The model structure of the first to the fourth submodules of the decoder. Here, "submodule 1" represents the structural diagram of the submodule of the current layer, "submodule 2" represents the structural diagram of the submodule of the upper layer, \(\oplus\) represents the residual operation of multiple types of feature maps, \(\otimes\) represents the fusion of 16 groups of residual feature maps, "interpolate" represents that the feature map size is magnified by 2 times using a linear strategy, and d represents the expansion coefficient of the dilated convolution
Figure 3 shows that we divide the feature map generated by the previous layer into 4 groups with equal numbers of channels. In Fig. 4a, the network structure in the blue dashed line implements PFMs with different scales, the network structure in the red dashed line implements DPFMs with different scales, and the other 14 network structures are the same as the first and second groups. Therefore, we obtained 16 sets of new feature maps from the previous layer of feature maps through convolution kernels with different directions and different scales. Figure 4a above shows that in the encoder part, we calculate the residuals between the feature maps generated by the current layer convolution and dilated convolution and the feature maps generated by the previous layer. Figure 4b shows that the biggest difference between the encoder and the decoder is that we have introduced the prior information of the encoder. Through the above feature fusion, each submodule of the DR-Net model can obtain richer feature maps.
Implementation details
All our experiments are performed on 3 sets of Tesla V100 (16 GB) GPUs. The two data sets only adjusted the size of the medical images and retain the scales of their length and width to 256 and 256, respectively. In addition, no preprocessing was performed on the images. The optimization function we chose is Adam, the learning rate is 0.001, the range of betas is 0.9–0.999, the loss function is the cross entropy loss, and all the comparison models are iterated 300 times during the entire training process. We ran each set of experiments 5 times and selected the set of results with the highest comprehensive evaluation.
Experiments
Data sets
In this paper, we validated the proposed model DR-Net on two multiclassification data sets: CHAOS and BraTS.
CHAOS dataset [34]: this data set consists of 4 foreground classes (liver, left kidney, right kidney and spleen) and a background class. The scale of all its medical images is 256 × 256. We selected 647 MRI images with ground truth labels for verification. The ratio of the number of samples in the training set to the test set is 3:2. The evaluation criteria of this data set come from the article by You et al. [26].
BraTS dataset [35, 36]: this data set consists of 3 foreground classes (oedema, nonenhancing solid core, and enhancing core) and a background class. The scale of all its medical images is 240 × 240, so we add zeros to each medical image. We selected the two sample images with the largest lesion area in each case in the 285 groups of cases. Furthermore, we also divide the samples into the training set and the validation set at a ratio of 3:2. The evaluation criteria of this data set come from the article by Bakas et al. [35] WT includes all three tumor structures, ET includes all tumor structures except "edema", and TC only contains the "enhancing core" structures that are unique to high-grade cases.
Comparison with the state-of-the-art models
In this section, we compare the classic medical segmentation model U-Net and compare some recent algorithms that have achieved good results in the field of medical image segmentation, such as CE-Net [18], DenseASPP [37], and MS-Dual [38]. The performance of the DR-Net model proposed in this article is verified by comparing its results with the results of other models (Tables 2, 3).
The experimental results of each model on the CHAOS data set show that although the result for Liver for CE-Net is 1% better than that of our method, in the Kidney L class, our method is 1.38% points better than the second highest method CE-Net. In the Kidney R category, our method is 2.12% points better than the second highest method MS-Dual. In the Spleen class, our method is 1.86% better than the second highest CE-Net. Furthermore, on the other three comprehensive indicators, our method obtained higher accuracy, which proved the feasibility of our method.
To further evaluate the generalization ability of our proposed method, we further verified each model on the BraTS dataset. Our method is better than other models on all WT values, indicating that our model has better generalization ability in multiclassification. The results of the three values of Dice+ clearly show that the DR-Net algorithm far outperforms other algorithms. The TC values in each test clearly show that our proposed model is 4.28%, 5.38%, and 0.1% better than the second-place algorithm in each test, which further proves that DR-Net is better at learning high-level cases. The experimental results on the above two data sets clearly show that the method proposed in this paper is effective. We visualized the segmentation results of each model in Fig. 4.
Figure 5 shows the segmentation results of each model on different data sets. The segmentation results of the first group, the fourth group and the sixth group show that the DR-Net algorithm has a better segmentation effect on the background class, thereby generating less noise. The segmentation results of other groups also show that DR-Net also has a good predictive ability in edge segmentation and small shapes. This further proves that our proposed method obtains deep feature maps that contain more semantic information by strengthening multiple feature maps. Even on small data sets and when there is a small number of feature maps, this method can learn more meaningful deep features.
Comparison of the total number of parameters in the two data sets
The total number of parameters directly affects the learning efficiency of a model, and the number of feature maps directly affects the total number of parameters. In this paper, our proposed method still obtains good results on most indicators on the two data sets while reducing the feature maps. Table 4 details the total number of parameters of each model.
Table 4 shows that among the five groups of models compared, our method uses only 2,700,581 parameters on the CHAOS data set and only 2,704,156 parameters on the BraTS data set, which are far lower than those of other models. Furthermore, there is no difference between the total number of parameters of the model on single-input data (CHAOS data set) and multiple-input data (BraTS data set).
The impact of different strategies on the experiment
To further analyze the feasibility of each strategy proposed in our method, we have verified each strategy. Here, "1" includes only the PFM module, "2" includes only the DPFM module, "3" incorporates only the prior semantic information of the previous layer, and "4" incorporates only the prior semantic information of the symmetric encoder submodule in the decoder submodule. To verify the semantic information, neither the PFM nor DPFM in "5" uses a rotation strategy. To use the feature fusion strategy in strategy "1" and strategy "2", we changed strategy "1" into two sets of DPFM modules and strategy "2" into two sets of PFM modules. We describe the learning process of each model through the line chart in Fig. 5.
The experimental results in Fig. 6 show that each strategy plays an active role in learning the semantic information of medical images. The structure of the PFM on the CHAOS data set has less of an impact. The reason may be that the distribution of various organs is not concentrated, so the global characteristics have greater impacts. In the BraTS data set, the PFM has a greater impact and has great fluctuations. The reason may be that the focus of the data set is relatively concentrated, and local features have become particularly important. Our method has the best experimental effect after fusing various feature maps, which proves the feasibility of our proposed algorithm.
Application of our proposed module in other models
To further prove the feasibility of the PFM and DPFM in our proposed DR-Net model, we introduced the PFM modules into U-Net. We made improvements in both the total number of parameters and the learning results and compared the results with those of the original U-Net. The results are shown in Table 5 below. The higher the value of ↑, the better the performance of the model.
We replaced the original convolution with the PFM module, except for the first layer of convolution. The experimental results on the CHAOS data set show that my improved U-Net has a 3.38% higher mAverage than the traditional U-Net mAverage, and the experimental results on the BraTS data set show that my improved U-Net has a 3.58% better mAverage than the traditional U-Net. The results of the above algorithm clearly illustrate the feasibility of our proposed strategy. The total number of parameters has also considerably decreased.
The influence of rotation angle and different convolution strategies on DR-Net algorithm
To further verify that the rotation strategy and different convolutional strategies can be used at the same time to obtain more semantic information, in this section, we use specific experimental results to analyze the importance of each strategy on the two data sets. DR-Net (0°) means that the feature map is not rotated at all, DR-Net (90°) means that the feature map is rotated by 90°, DR-Net (180°) means that the feature map is rotated by 180°, DR-Net (270°) means that the feature map is rotated by 270°, DR-Net(a) means that only the traditional convolution is used for feature learning, and DR-Net(b) means that only the dilated volumes of product feature learning are used. The concrete results on the two data sets are shown in Table 6.
In Table 6, we find that the rotation strategy has a greater effect on the CHAOS dataset than the convolution strategy. Different rotation strategies obtain different features, which leads to different learning points for each class. When the feature map is rotated by 90°, Liver obtains the highest value of 90.53%; when the feature map is rotated by 180°, Spleen achieved an accuracy of 87.34%; when the feature map was rotated by 270°, Kidney L achieved an accuracy of 81.56%; and when the feature map was not rotated, the various results were relatively balanced. The best prediction results for the various types in the CHAOS data set were obtained when using the 4 types of rotations with DR-Net. In Table 7, on the multimodal BraTS dataset, we find that both the rotation strategy and the convolution strategy play positive roles. If only the traditional convolution is used in the DR-Net algorithm, the results for the learned features are lower than when using only the dilated convolution, which shows that the global features obtained by the dilated convolution on multimodal tasks are more important. Tables 6 and 7 show that when the DR-Net algorithm uses the rotation strategy and multiple convolution strategies at the same time, the best prediction results are obtained, which further proves the rationality of the DR-Net algorithm.
Conclusion
This paper proposes a dual-rotation network, which fully learns and integrates global features, local features, shallow features and deep features to mine more semantic information. Furthermore, to further integrate more semantic information in the encoder and decoder, we further strengthen the feature map through three strategies of rotation, multiscaling, and different dilation step sizes of the feature maps. We fuse the original feature map of the previous submodule through the latter submodule to provide more semantic information for the encoder; furthermore, in order for the decoder to obtain more semantic information, we compress and expand the features. The graph realizes the fusion of multiple types of feature graphs. Finally, the method proposed in this paper obtained good segmentation results on two multiclass medical image data sets.
References
Taghanaki SA, Abhishek K, Cohen JP et al (2021) Deep semantic segmentation of natural and medical images: a review. Artif Intell Rev 54(1):137–178
Wang EK, Chen CM, Hassan MM et al (2020) A deep learning based medical image segmentation technique in internet-of-medical-things domain. Futur Gener Comput Syst 108:135–144
Ni J, Wu J, Tong J et al (2020) GC-Net: global context network for medical image segmentation. Comput Methods Programs Biomed 190:105121
Liu Q, Yu L, Luo L et al (2020) Semi-supervised medical image classification with relation-driven self-ensembling model. IEEE Trans Medical Imaging 39(11):3429–3440
Huang Z, Zhu X, Ding M et al (2020) Medical image classification using a light-weighted hybrid neural network based on PCANet and DenseNet. IEEE Access 8:24697–24712
Zhang Q et al (2020) A GPU-based residual network for medical image classification in smart medicine. Inf Sci 536:91–100
Eastman AJ, Noble KN, Pensabene V et al (2020) Leveraging bioengineering to assess cellular functions and communication within human fetal membranes. J Matern Fetal Neonatal Med 1–13
Sadak F, Saadat M, Hajiyavand AM (2020) Real-time deep learning-based image recognition for applications in automated positioning and injection of biological cells. Comput Biol Med 125:103976
Juneja K, Rana C (2021) Compression-robust and fuzzy-based feature-fusion model for optimizing the iris recognition. Wirel Pers Commun 116(1):267–300
Feng J, Teng Q, Li B et al (2020) An end-to-end three-dimensional reconstruction framework of porous media from a single two-dimensional image based on deep learning. Comput Methods Appl Mech Eng 368:113043
Hu J, Peng A, Deng K et al (2020) Value of CT and three-dimensional reconstruction revealing specific radiological signs for screening causative high jugular bulb in patients with Meniere’s disease. BMC Med Imaging 20(1):1–10
Wang J, Huang Z, Yang X et al (2020) Three-dimensional reconstruction of jaw and dentition cbct images based on improved marching cubes algorithm. Proc CIRP 89:239–244
Chen L, Bentley P, Mori K et al (2018) DRINet for medical image segmentation. IEEE Trans Med Imaging 37(11):2453–2462
Huang G, Liu Z, Van Der Maaten L et al (2017) Densely connected convolutional networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 4700–4708
Szegedy C, Vanhoucke V, Ioffe S et al (2016) Rethinking the inception architecture for computer vision. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 2818–2826
Alom MZ, Yakopcic C, Hasan M et al (2019) Recurrent residual U-Net for medical image segmentation. J Med Imaging 6(1):014006
Ronneberger O, Fischer P, Brox T (2015) U-net: convolutional networks for biomedical image segmentation. In: International conference on medical image computing and computer-assisted intervention. Springer, Cham, pp 234–241
Gu Z, Cheng J, Fu H et al (2019) Ce-net: Context encoder network for 2d medical image segmentation. IEEE Trans Med Imaging 38(10):2281–2292
He K, Zhang X, Ren S et al (2016) Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 770–778
You H, Tian S, Yu L et al (2020) A new multiple max-pooling integration module and cross multiscale deconvolution network based on image semantic segmentation. arXiv:2003.11213
Xie S, Girshick R, Dollár P et al (2017) Aggregated residual transformations for deep neural networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 1492–1500
Romero D, Bekkers E, Tomczak J et al (2020) Attentive group equivariant convolutional networks. In: International conference on machine learning. In: PMLR, pp 8188–8199
Moradmand H, Aghamiri SMR, Ghaderi R (2020) Impact of image preprocessing methods on reproducibility of radiomic features in multimodal magnetic resonance imaging in glioblastoma. J Appl Clin Med Phys 21(1):179–190
Heidari M, Mirniaharikandehei S, Khuzani AZ et al (2020) Improving the performance of CNN to predict the likelihood of COVID-19 using chest X-ray images with preprocessing algorithms. Int J Med Inform 144:104284
Chollet F (2017) Xception: deep learning with depthwise separable convolutions. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 1251–1258
You H, Tian S, Yu L et al (2020) DT-Net: a novel network based on multi-directional integrated convolution and threshold convolution. arXiv:2009.12569v1
Denton E, Zaremba W, Bruna J, LeCun Y, Fergus R (2014) Exploiting linear structure within convolutional networks for efficient evaluation. In: NIPS
Kim Y-D, Park E, Yoo S, Choi T, Yang L, Shin D (2016) Compression of deep convolutional neural networks for fast and low power mobile applications. In: ICLR
Ioannou Y, Robertson D, Cipolla R, Criminisi A (2016) Deep roots: improving CNN efficiency with hierarchical filter groups. arXiv:1605.06489
Jaderberg M, Vedaldi A, Zisserman A (2014) Speeding up convolutional neural networks with low rank expansions. In: BMVC
Gao S, Cheng M M, Zhao K et al (2019) Res2net: a new multi-scale backbone architecture. In: IEEE transactions on pattern analysis and machine intelligence
Oktay O, Schlemper J, Folgoc LL et al (2018) Attention u-net: learning where to look for the pancreas. arXiv:1804.03999
Wang Y, He Z, Xie P et al (2020) Segment medical image using U-Net combining recurrent residuals and attention. In: International conference on medical imaging and computer-aided diagnosis. Springer, Singapore, pp 77–86
Selvi E, Selver MA, Kavur AE, Guzelis C, Dicle O (2015) Segmentation of abdominal organs from MR images using multi-level hierarchical classification. J Fac Eng Architect Gazi Univ 30:533–546
Bakas S (2017) Advancing The Cancer Genome Atlas glioma MRI collections with expert segmentation labels and radiomic features. Nat Sci Data 4:170117. https://doi.org/10.1038/sdata.2017.117
Menze BH (2015) The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans Med Imag 34(10):1993–2024. https://doi.org/10.1109/TMI.2014.2377694
Yang M, Yu K, Zhang C et al (2018) Denseaspp for semantic segmentation in street scenes. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 3684–3692.
Sinha A, Dolz J (2019) Multi-scale guided attention for medical image segmentation. arXiv:1906.02849
Funding
This research is partially supported by Science and Technology Department of Xinjiang Uyghur Autonomous Region (2020E0234). Xinjiang Autonomous Region key research and development project (2021B03001-4).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest in the publication of this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
You, H., Yu, L., Tian, S. et al. DR-Net: dual-rotation network with feature map enhancement for medical image segmentation. Complex Intell. Syst. 8, 611–623 (2022). https://doi.org/10.1007/s40747-021-00525-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40747-021-00525-4