Abstract
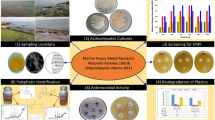
The objective is to develop low cost wastewater treatment systems for the efficient removal of toxic heavy metal ions including arsenic (As). For this, two bacterial strains, one gram negative and other gram positive dubbed as IT6 and S12, were isolated from arsenic contaminated wastewater and soil samples from Sheikhupura, Pakistan. The bacterial isolates were checked for their ability to resist various metal ions and antibiotics and were identified on the basis of 16S rRNA ribotyping. The strains were also checked for their potential to decontaminate arsenite at lab scale. Both strains were identified as Pseudomonas monteilii (IT6) and Bacillus infantis (S12) with minimum inhibitory concentration of 26.5 and 33 mM arsenite, respectively. Apart from arsenite, both bacterial strains showed fair resistance against other metal ions including chromium, lead, cobalt, selenium, zinc, cadmium, and mercury. Both IT6 and S12 showed high potential of arsenite oxidation of 92% and 96% at 37 °C and pH of 7 with 100 µg/mL arsenite after 96 h of incubation. The strains have also shown strong resistance against commonly used antibiotics including amikacin, imipenem, and ciprofloxacin. These bacterial strains are potential candidates to exterminate toxic metal ions from the wastewater for green chemistry due to presence of multiple metal resistance and efficient arsenite oxidizing potential.





Similar content being viewed by others
Data Availability
The data used to support the findings of this study are available from the corresponding author upon request.
References
Aguilar NC, Faria MCS, Pedron T, Batista BL, Mesquita JP, Bomfeti CA, Rodrigues JL (2020) Isolation and characterization of bacteria from a brazilian gold mining area with a capacity of arsenic bioaccumulation. Chemosphere 240:124871
Altowayti WAH, Almoalemi H, Shahir S, Othman N (2020) Comparison of culture-independent and dependent approaches for identification of native arsenic-resistant bacteria and their potential use for arsenic bioremediation. Ecotoxicol Environ Saf 250:111267
Aslam F, Yasmin A, Thomas T (2018) Essential gene clusters identified in Stenotrophomonas MB339 for multiple metal/antibiotic resistance and xenobiotic degradation. Curr Microbiol 75:1484–1492
Bahar MM, Megharaj M, Naidu R (2012) Arsenic bioremediation potential of a new arsenite-oxidizing bacterium Stenotrophomonas sp. MM-7 isolated from soil. Biodegradation 23:803–812
Battaglia-Brunet F, Dictor MC, Garrido F, Crouzet C, Morin D, Dekeyser K, Baranger P (2002) An arsenic (III) oxidizing bacterial population: selection, characterization, and performance in reactors. J Appl Microbiol 93:656–667
Battaglia-Brunet F, Itard Y, Garrido F, Delorme F, Crouzet C, Greffié C, Joulian C (2006a) A simple biogeochemical process removing arsenic from a mine drainage water. Geomicrobiol J 23:201–211
Battaglia-Brunet F, Joulian C, Garrido F (2006b) Oxidation of arsenite by Thiomonas strains and characterization of Thiomonas arsenivorans sp. nov. Antonie Van Leeuwenhoek 89:99–108
Bhakat K, Chakraborty A, Islam E (2019) Characterization of arsenic oxidation and uranium bioremediation potential of arsenic resistant bacteria isolated from uranium ore. Environ Sci Pollut Res 26:12907–12919 ()
Cai L, Liu G, Rensing C, Wang G (2009) Genes involved in arsenic transformation and resistance associated with different levels of arsenic-contaminated soils. BMC Microbiol 9:4–13
Cappucino JG, Sherman N (2001) Microbiology: a laboratory manual, 6th edn. Pearson education, San Francisco
Dey U, Das K, Roy P, Chatterjee SN, Mondal NK (2015) Searching of microbial agent for bioremediation of arsenic. Int J Extentsive Res 5:60–64
Dey U, Chatterjee S, Mondal NK (2016) Isolation and characterization of arsenic-resistant bacteria and possible application in bioremediation. Biotechnol Rep 10:1–7
Dey U, Chatterjee S, Mondal NK (2017) Investigation of bioremediation of arsenic by bacteria isolated from an arsenic contaminated area. Environ Process 4:183–199
Donahoe-Christiansen J, D’Imperio S, Jackson CR, Inskeep WP, McDermott TR (2004) Arsenite-oxidizing Hydrogenobaculum strain isolated from an acid-sulfate-chloride Geothermal spring in Yellowstone National Park. Appl Environ Microbiol 70:1865–1868
Edwards KJ, Bond PL, Gihring TM, Banfield JF (2000) An archaeal iron-oxidizing extreme acidophile important in acid mine drainage. Science 287:1796–1799
Elahi A, Rehman A (2019a) Multiple metal resistance and Cr6+ reduction by bacterium, Staphylococcus sciuri A-HS1, isolated from untreated tannery effluent. J King Saud Uni Sci 31:1005–1013
Finney LA, O’halloran TV (2003) Transition metal speciation in the cell: insights from the chemistry of metal ion receptors. Science 300:931–936
Gihring TM, Banfield JF (2001) Arsenite oxidation and arsenate respiration by a new Thermus isolate. FEMS Microbiol Lett 204:335–340
Green HH (1918) Isolation and description of a bacterium causing oxidation of arsenite to arsenate in cattle dipping baths. Union S. Africa Dept. Agrie. 5th and 6th Repts. Direc. Vet. Research, pp 595–610
Guarino F, Miranda A, Castiglione S, Cicatelli A (2020) Arsenic phytovolatilization and epigenetic modifications in Arundo donax L. assisted by a PGPR consortium. Chemosphere 251:126310
Irem S, Islam E, Maathuis FJM, Niazi NK, Li T (2019) Assessment of potential dietary toxicity and arsenic accumulation in two contrasting rice genotypes: Effect of soil amendments. Chemosphere 225:104–114
Ji PY, Li ZY, Wang H, Dong JT, Li XJ, Yi HI (2019) Arsenic and sulfur dioxide co-exposure induce renal injury via activation of the NF-κB and caspase signaling pathway. Chemosphere 224:280–288
Ke C, Zhao C, Rensing C, Yang S, Zhang Y (2018) Characterization of recombinant E. coli expressing arsR from Rhodopseudomonas palustris CGA009 that displays highly selective arsenic adsorption. Appl Microbiol Biotechnol 102:6247–6255
Lett MC, Muller D, Lièvremont D, Silver S, Santini J (2012) Unified nomenclature for genes involved in prokaryotic aerobic arsenite oxidation. J Bacteriol 194:207–208
Lin YF, Walmsley AR, Rosen BP (2006) An arsenic metallochaperone for an arsenic detoxification pump. Proceed Natl Acad Sci USA 103:15617–15622
Moens M, Branco R, Morais PV (2020) Arsenic accumulation by a rhizosphere bacterial strain Ochrobactrum tritici reduces rice plant arsenic levels. World J Microbiol Biotechnol 36:23
Mujawar SY, Shamim K, Vaigankar DC, Dubey SK (2019) Arsenite biotransformation and bioaccumulation by Klebsiella pneumoniae strain SSSW7 possessing arsenite oxidase (aioA) gene. BioMetals 32:65–76
Mukhopadhyay R, Rosen BP, Phung LT, Silver S (2002) Microbial arsenic: from geocycles to genes and enzymes. FEMS Microbiol Rev 26:311–325
Naidu R, Bhattacharya P (2009) Arsenic in the environment-risks and management strategies. Environ Geochem Health 31:1–8
Naureen A, Rehman A (2016) Arsenite oxidizing multiple metal resistant bacteria isolated from industrial effluent: their potential use in wastewater treatment. World J Microbiol Biotechnol 32:133–144
Pasha C, Narayana B (2008) Determination of arsenic in environmental and biological samples using toluidine blue or safranine O by simple spectrophotometric method. Bull Environ Contam Toxicol 81:47–51
Pattanapipitpaisal P, Brown NL, Macaskie LE (2001) Chromate reduction and 16S rRNA identification of bacteria isolated from a Cr(VI)-contaminated site. Appl Microbiol Biotechnol 57:257–261
Qin J, Rosen BP, Zhang Y, Wang G, Franke S, Rensing C (2006) Arsenic detoxification and evolution of trimethylarsine gas by a microbial arsenite S-adenosylmethionine methyltransferase. Proceed Natl Acad Sci USA 103:2075–2080
Rahman A, Nahar N, Nawani NN, Jass J, Desale P, Kapadnis BP, Hossain K, Saha AK, Ghosh S, Olsson B, Mandal A (2014) Isolation and characterization of Lysinibacillus strain B1-CDA showing potential for bioremediation of arsenics from contaminated water. J Environ Sci Hlth Part A 49:1349–1360
Rehman A, Ali A, Muneer B, Shakoori AR (2007) Resistance and biosorption of mercury by bacteria isolated from industrial effluents. Pakistan J Zool 39:137–146
Romero-Isart N, Vašák M (2002) Advances in the structure and chemistry of metallothioneins. J Inorg Biochem 88:388–396
Sambrook J, MacCallum P, Russell D (2001) Molecular cloning: a laboratory manual. 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Santini JM, Sly LI, Schnagl RD, Macy JM (2000) A new chemolithoautotrophic arsenite oxidizing bacterium isolated from a gold mine: phylogenetic, physiological, and preliminary biochemical studies. Appl Environ Microbiol 66:92–97
Santini JM, Sly LI, Wen A, Comrie D, Wulf-Durand PD, Macy JM (2002) New arsenite oxidizing bacteria isolated from Australian Gold Mining Environments-Phylogenetic relationships. Geomicrobiol J 19:67–76
Santini JM, Kappler U, Ward SA, Honeychurch MJ, vanden RN, Bernhardt PV (2007) The NT-26 cytochrome c 552 and its role in arsenite oxidation. BBA Bioenergetics 1767:189–196
Sehlin HM, Lindström EB (1992) Oxidation and reduction of arsenic by Sulfolobus acidocaldarius strain BC. FEMS Microbiol Lett 93:87–92
Sher S, Rehman A (2019) Use of heavy metals resistant bacteria-a strategy for arsenic bioremediation. Appl Microbial Biotechnol 103:6007–6021
Sher S, Rehman A, Hansen LH, Nielsen TK (2019) Complete genome sequences of highly arsenite-resistant bacteria Brevibacterium sp. strain CS2 and Micrococcus luteus AS2. Microbiol Resour Announc 31:e00531–e00519. https://doi.org/10.1128/MRA.00531-19
Sher S, Hussain SZ, Rehman A (2020a) Phenotypic and genomic analysis of multiple heavy metal–resistant Micrococcus luteus strain AS2 isolated from industrial waste water and its potential use in arsenic bioremediation. Appl Microbial Biotechnol 104:2243–2254
Sher S, Hussain SZ, Rehman A (2020b) Multiple resistance mechanisms in Staphylococcus sp. strain AS6 under arsenite stress and its potential use in amelioration of wastewater. J King Saud Uni Sci 32:3050–3058
Shi K, Cao M, Li C, Huang J, Zheng S, Wang (2019) Efflux proteins MacAB confer resistance to arsenite and penicillin/macrolide-type antibiotics in Agrobacterium tumefaciens 5A. World J Microbiol Biotechnol 35:115
Silver S, Phung LT (2005) Genes and enzymes involved in bacterial oxidation and reduction of inorganic arsenic. Appl Environ Microbiol 71:599–608
Simeonova DD, Lièvremont D, Lagarde F, Muller DA, Groudeva VI, Lett M-C (2004) Microplate screening assay for the detection of arsenite-oxidizing and arsenate-reducing bacteria. FEMS Microbiol Lett 237:249–253
Solioz M, Stoyanov JV (2003) Copper homeostasis in Enterococcus hirae. FEMS Microbiol Rev 27:183–195
ThankGod CE, Michelangeli F, Otitoloju AA (2019) In vitro cyto-toxic assessment of heavy metals and their binary mixtures on mast cell-like, rat basophilic leukemia (RBL-2H3) cells. Chemosphere 223:686–693
Vahter M (2000) Genetic polymorphism in the biotransformation of inorganic arsenic and its role in toxicity. Toxicol Lett 112:209–217
Wu D, Zhang Z, Gao Q, Ma Y (2018) Isolation and characterization of aerobic, culturable, arsenic-tolerant bacteria from lead-zinc mine tailing in southern China. World J Microbiol Biotechnol 34(12):177
Yan Z, Li M, Wang J, Pan J (2019) Genome analysis revealing the potential mechanisms for the heavy metal resistance of Pseudomonas sp. P11, isolated from industrial wastewater sediment. Curr Microbiol 76:1361–1368
Author information
Authors and Affiliations
Contributions
SS1 performed experiments. AG helped in data analysis. SS2 and AR conceived and designed the study. The first draft of the manuscript was written by Shahid Sher and all authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sher, S., Ghani, A., Sultan, S. et al. Bacterial Strains Isolated from Heavy Metals Contaminated Soil and Wastewater with Potential to Oxidize Arsenite. Environ. Process. 8, 333–347 (2021). https://doi.org/10.1007/s40710-020-00488-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40710-020-00488-7